

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144726.5 - phase: 0 /pseudo
(293 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
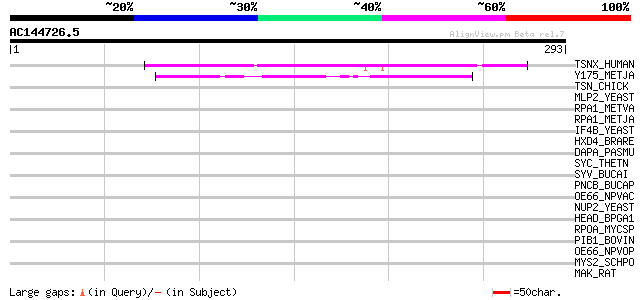
TSNX_HUMAN (Q99598) Translin-associated protein X (Translin-asso... 127 4e-29
Y175_METJA (Q57639) Hypothetical protein MJ0175 54 6e-07
TSN_CHICK (P79769) Translin 39 0.012
MLP2_YEAST (P40457) MLP2 protein (Myosin-like protein 2) 37 0.058
RPA1_METVA (P41556) DNA-directed RNA polymerase subunit A' (EC 2... 35 0.17
RPA1_METJA (Q58445) DNA-directed RNA polymerase subunit A' (EC 2... 35 0.17
IF4B_YEAST (P34167) Eukaryotic translation initiation factor 4B ... 34 0.49
HXD4_BRARE (O57374) Homeobox protein Hox-D4 33 1.1
DAPA_PASMU (Q9CLZ7) Dihydrodipicolinate synthase (EC 4.2.1.52) (... 32 1.4
SYC_THETN (Q8R7T3) Cysteinyl-tRNA synthetase (EC 6.1.1.16) (Cyst... 32 1.9
SYV_BUCAI (P57447) Valyl-tRNA synthetase (EC 6.1.1.9) (Valine--t... 32 2.4
PNCB_BUCAP (Q8K9I6) Nicotinate phosphoribosyltransferase (EC 2.4... 32 2.4
OE66_NPVAC (Q00704) Occlusion-derived virus envelope protein E66... 31 3.2
NUP2_YEAST (P32499) Nucleoporin NUP2 (Nuclear pore protein NUP2)... 31 3.2
HEAD_BPGA1 (Q9FZW7) Major head protein 31 3.2
RPOA_MYCSP (P38018) DNA-directed RNA polymerase alpha chain (EC ... 31 4.1
PIB1_BOVIN (P10894) 1-phosphatidylinositol-4,5-bisphosphate phos... 31 4.1
OE66_NPVOP (O10305) Occlusion-derived virus envelope protein E66... 30 5.4
MYS2_SCHPO (Q9USI6) Myosin type II heavy chain 1 30 5.4
MAK_RAT (P20793) Serine/threonine-protein kinase MAK (EC 2.7.1.3... 30 5.4
>TSNX_HUMAN (Q99598) Translin-associated protein X
(Translin-associated factor X)
Length = 290
Score = 127 bits (318), Expect = 4e-29
Identities = 77/220 (35%), Positives = 127/220 (57%), Gaps = 21/220 (9%)
Query: 72 FTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRM-SKYNKDEVLEKAEKDLAAVT 130
F + + L+ +DK ER+VK SRDIT+ SK+ IF +HR+ S + +++L ++E L V
Sbjct: 39 FKSFQQELDARHDKYERLVKLSRDITVESKRTIFLLHRITSAPDMEDILTESEIKLDGV- 97
Query: 131 NQHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLDEINKTLL---- 186
Q + ++ +EL G D + RA + G+QEYVEA +F F K +L+ +DEINK L+
Sbjct: 98 RQKIFQVAQELSGEDMHQFHRAITTGLQEYVEAVSFQHFIKTRSLISMDEINKQLIFTTE 157
Query: 187 --------PLSDPSLQP-----LQINILDYILGLADLTGELMRLAIGRISDGELEFAEKI 233
P SD + L++ +DY+LG+ADLTGELMR+ I + +G+++ ++
Sbjct: 158 DNGKENKTPSSDAQDKQFGTWRLRVTPVDYLLGVADLTGELMRMCINSVGNGDIDTPFEV 217
Query: 234 CSFARDIYRELTLVVPHMDDSSDMKTKMETMLQSVMKIEN 273
F R +Y + + ++ K+ T+ QS+ K+EN
Sbjct: 218 SQFLRQVYDGFSFI--GNTGPYEVSKKLYTLKQSLAKVEN 255
>Y175_METJA (Q57639) Hypothetical protein MJ0175
Length = 222
Score = 53.5 bits (127), Expect = 6e-07
Identities = 51/169 (30%), Positives = 79/169 (46%), Gaps = 28/169 (16%)
Query: 78 YLNNLNDKRERVVKASRDITMNSKKVIFQVHRMSKYNKDEVLEKAEKDLAAVTNQHVSRL 137
YL N + RE ++K SR+IT + +I ++H+ +KDE +K + +S
Sbjct: 27 YLANKDSVREEILKLSREITRDCAMLIRKIHKSD--DKDEFKDKLNE---------ISEK 75
Query: 138 VKELQG-TDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLDEINKTLLPLSDPSLQPL 196
+K+L F + S QE+VEA + ++K D NK PS + L
Sbjct: 76 IKKLNSLATFPEFVGYLSTPQQEFVEALSLY-------MIKFD--NKI------PSFKEL 120
Query: 197 Q-INILDYILGLADLTGELMRLAIGRISDGELEFAEKICSFARDIYREL 244
I +YILGLAD+ GEL R + + + L E+ F D+Y L
Sbjct: 121 DFIKEENYILGLADVIGELRREVLEAMKNDNLAEVERYFKFMEDLYEFL 169
>TSN_CHICK (P79769) Translin
Length = 229
Score = 39.3 bits (90), Expect = 0.012
Identities = 40/198 (20%), Positives = 89/198 (44%), Gaps = 15/198 (7%)
Query: 84 DKRERVVKASRDITMNSKKVIFQ---VHRMSKYNK-DEVLEKAEKDLAAVTNQHVSRLVK 139
D RE + K + + +++++ VH+ + + + +KA + V Q S L
Sbjct: 19 DIREEIRKVVQALEQTAREMLTLPQGVHQGAGFQDIPKKCQKAREHFGTVRTQMES-LKT 77
Query: 140 ELQGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLDEINKTLLPLSDPSLQPLQIN 199
+ +++ + +Q V A+F + + TL+ + + + +L + + ++
Sbjct: 78 KFPADQYYRFHEHWRFVLQRLVFLASFVVYLETETLVTREAVAE-ILGIEADRERGFHLD 136
Query: 200 ILDYILGLADLTGELMRLAIGRISDGELEFAEKICSFARDI---YRELTLVVPHMDDSSD 256
I DY+ G+ L EL RLA+ ++ G+ +I +F ++ +R L L +
Sbjct: 137 IEDYLSGVLTLASELARLAVNSVTAGDYSRPLRISTFINELDSGFRLLNL------KNDS 190
Query: 257 MKTKMETMLQSVMKIENV 274
++ + + + V KIE V
Sbjct: 191 LRKRYDGLKYDVKKIEEV 208
>MLP2_YEAST (P40457) MLP2 protein (Myosin-like protein 2)
Length = 1679
Score = 37.0 bits (84), Expect = 0.058
Identities = 28/136 (20%), Positives = 58/136 (42%), Gaps = 12/136 (8%)
Query: 19 EKTQRLHQSILLFFDFGFSTVTGTNFQNTSKRPKTMSIATDTATVTDSAMKEPFTKYTEY 78
++TQ L + ++ S V F++ +K ++I + + ++K K E
Sbjct: 1143 QRTQTLSEK-----EYQCSAVIIDEFKDITKEVTQVNILKENNAILQKSLKNVTEKNREI 1197
Query: 79 LNNLNDKRERVVKASRDITMNSKKVIFQVHRMSKYNKDEVLEKAEKDLAAVTNQHVSRLV 138
LND++E + + RD+ ++V +++ Y ++E + Q +S+
Sbjct: 1198 YKQLNDRQEEISRLQRDLIQTKEQVSINSNKILVY-------ESEMEQCKQRYQDLSQQQ 1250
Query: 139 KELQGTDFWKLRRAYS 154
K+ Q D KL S
Sbjct: 1251 KDAQKKDIEKLTNEIS 1266
>RPA1_METVA (P41556) DNA-directed RNA polymerase subunit A' (EC
2.7.7.6)
Length = 889
Score = 35.4 bits (80), Expect = 0.17
Identities = 23/85 (27%), Positives = 43/85 (50%), Gaps = 8/85 (9%)
Query: 195 PLQINILDYILGLADLTGELMRLAIGRISD-----GELEFAEKIC--SFARDIYRELTLV 247
P+ ++D LG+ D G + + GR+ G +E ++ + FA+DIY+ L V
Sbjct: 43 PIDGGLMDTRLGVID-PGLVCKSCSGRVGTCPGHFGHIELSKSVIHIGFAKDIYKLLKAV 101
Query: 248 VPHMDDSSDMKTKMETMLQSVMKIE 272
PH + + K + L+ ++K+E
Sbjct: 102 CPHCGKVTVTEIKRDEYLEKMLKLE 126
>RPA1_METJA (Q58445) DNA-directed RNA polymerase subunit A' (EC
2.7.7.6) [Contains: Mja rpoA1 intein (Mja rpol A'
intein)]
Length = 1341
Score = 35.4 bits (80), Expect = 0.17
Identities = 25/85 (29%), Positives = 44/85 (51%), Gaps = 8/85 (9%)
Query: 195 PLQINILDYILGLADLTGELMRLAIGRISD-----GELEFAEKIC--SFARDIYRELTLV 247
P+ ++D LG+ D G + + GRI + G +E A+ + FA+ IY+ L V
Sbjct: 43 PIDGGLMDTRLGVID-PGLVCKTCGGRIGECPGHFGHIELAKPVIHIGFAKTIYKILKAV 101
Query: 248 VPHMDDSSDMKTKMETMLQSVMKIE 272
PH + +TK + +L+ + K+E
Sbjct: 102 CPHCGRVAISETKRKEILEKMEKLE 126
>IF4B_YEAST (P34167) Eukaryotic translation initiation factor 4B
(eIF-4B)
Length = 436
Score = 33.9 bits (76), Expect = 0.49
Identities = 29/110 (26%), Positives = 47/110 (42%), Gaps = 4/110 (3%)
Query: 33 DFGFSTVTGTNFQNTSKRPKTMSIATDTATVTDSAMKEPFTKYTEYLNNLNDKRERVVKA 92
D +S G+NFQ++S+ P+ + A +A F K + N D R K
Sbjct: 311 DIDWSAARGSNFQSSSRPPRREREKEEPALDWGAARGAQFGKPQQTKNTYKD-RSLTNKK 369
Query: 93 SRDITMNSKKVIFQVHRMSKYNKDEVLEKAEKDLAAVTNQHVSRLVKELQ 142
+ D +K ++ V R ++D E+AEK V V++LQ
Sbjct: 370 TTDEQPKIQKSVYDVLRTEDDDED---EEAEKQNGDAKENKVDAAVEKLQ 416
>HXD4_BRARE (O57374) Homeobox protein Hox-D4
Length = 236
Score = 32.7 bits (73), Expect = 1.1
Identities = 28/106 (26%), Positives = 45/106 (42%), Gaps = 11/106 (10%)
Query: 39 VTGTNFQNTSKRPKTMSIATDTATVTDSAMKEPFTKYTEYLNNLNDKRERVVKASRDITM 98
VT N T PK A V + + F +Y R R ++++ +++
Sbjct: 134 VTTVNPDYTGPEPKRSRTAYTRQQVLELEKEFHFNRYLT--------RRRRIESAHTLSL 185
Query: 99 NSK--KVIFQVHRMSKYNKDEVLEKAEKDLAAVTNQHVSRLVKELQ 142
+ + K+ FQ RM K+ KD L + A+V NQH K+ Q
Sbjct: 186 SERQIKIWFQNRRM-KWKKDHKLPNTKGRSASVGNQHAQHAQKDSQ 230
>DAPA_PASMU (Q9CLZ7) Dihydrodipicolinate synthase (EC 4.2.1.52)
(DHDPS)
Length = 298
Score = 32.3 bits (72), Expect = 1.4
Identities = 33/137 (24%), Positives = 55/137 (40%), Gaps = 27/137 (19%)
Query: 107 VHRMSKYNKDEVLEKAEKDLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEY------ 160
V R++K N +++A DL+ V +KEL G DF L + G++
Sbjct: 154 VARLAKINNIVAIKEATGDLSRVAK------IKELAGEDFIFLSGDDATGLESIKLGGQG 207
Query: 161 ----------VEAATFCSFCKNGTLLKLDEINKTLLPLSD---PSLQPLQINILDYILGL 207
+ A C NG + ++IN+ L+ L P+ + Y LGL
Sbjct: 208 VISVTNNVAAADMAKMCHLALNGQFEEAEQINQRLMALHKNLFVESNPIPVKWAAYRLGL 267
Query: 208 ADLTGELMRLAIGRISD 224
D +RL + +S+
Sbjct: 268 IDT--PTLRLPLTTLSE 282
>SYC_THETN (Q8R7T3) Cysteinyl-tRNA synthetase (EC 6.1.1.16)
(Cysteine--tRNA ligase) (CysRS)
Length = 467
Score = 32.0 bits (71), Expect = 1.9
Identities = 39/182 (21%), Positives = 81/182 (44%), Gaps = 18/182 (9%)
Query: 64 TDSAMKEPFTKYTEYLNNLNDKRERVVKAS------RDITMNSKKVIFQVHRMSKYNKD- 116
+++A +PF+KY ++ LN E++ K+ R++T + ++ + + +
Sbjct: 240 SEAAYDQPFSKYWMHIGYLNVNNEKMSKSKGNFFTVRELTEKYDPEVLRLFMLMAHYRSP 299
Query: 117 -----EVLEKAEKDLAAVTN-----QHVSRLVKELQGTDFWKLRRAYSPGIQEYVEAATF 166
++L++A+ ++N +H++ + K+ + +D K ++ E A
Sbjct: 300 INFSLDLLDQAKSAYERLSNAVVNLKHLANITKDRELSDEEKDLIVKFDEYKKQFEEAMD 359
Query: 167 CSFCKNGTLLKLDEINKTL-LPLSDPSLQPLQINILDYILGLADLTGELMRLAIGRISDG 225
F + L E+ KT LS S + L IL+ L L+D+ G + A I D
Sbjct: 360 DDFNTADAISVLFEMAKTANANLSGNSSKKLVEYILEMFLKLSDVLGLSYKNAETEIEDE 419
Query: 226 EL 227
E+
Sbjct: 420 EI 421
>SYV_BUCAI (P57447) Valyl-tRNA synthetase (EC 6.1.1.9) (Valine--tRNA
ligase) (ValRS)
Length = 955
Score = 31.6 bits (70), Expect = 2.4
Identities = 33/119 (27%), Positives = 58/119 (48%), Gaps = 13/119 (10%)
Query: 78 YLNNLNDKRERVVKASRDITMNSKKVIFQVHRMSKYNKD----EVLEKAEKDLAAV---- 129
+L N+N ++E+V+K + + N + KY+KD E+++ AE + +
Sbjct: 825 FLYNINSEQEKVIKENISLIKNISFLDKITILSEKYDKDLCIKEIIDGAEIIIPILKLVD 884
Query: 130 TNQHVSRLVKELQGTD--FWKLR-RAYSPGIQEYVEAATFCSFCKNGTLLKLDEINKTL 185
T + RLVKE + T+ +K++ + + Y A T + LLKL+EIN L
Sbjct: 885 TKVELKRLVKEQKKTELNIFKIKTKILNKDFLSY--APTKIVTQEKNKLLKLNEINLKL 941
>PNCB_BUCAP (Q8K9I6) Nicotinate phosphoribosyltransferase (EC
2.4.2.11) (NAPRTase)
Length = 398
Score = 31.6 bits (70), Expect = 2.4
Identities = 20/68 (29%), Positives = 35/68 (51%), Gaps = 5/68 (7%)
Query: 79 LNNLNDKRERVVKASRDITMNSKKVI-FQVHRMSKYNKDEVLEKAEKD----LAAVTNQH 133
+ +LN K + K +RD+ ++ K++ F R Y+ + K KD L +N H
Sbjct: 147 VQHLNKKLKEFFKKNRDVDLSRLKIVDFGTRRRFSYDVQYSIIKRLKDTFPFLIGSSNYH 206
Query: 134 VSRLVKEL 141
+SR++K L
Sbjct: 207 ISRILKLL 214
>OE66_NPVAC (Q00704) Occlusion-derived virus envelope protein E66
(ODV-E66)
Length = 704
Score = 31.2 bits (69), Expect = 3.2
Identities = 13/38 (34%), Positives = 23/38 (60%)
Query: 80 NNLNDKRERVVKASRDITMNSKKVIFQVHRMSKYNKDE 117
N+LN+ RV+ SRD ++N+ + F+ R++ N E
Sbjct: 510 NSLNNTNGRVIVLSRDTSVNTNDLSFEAQRINNNNSSE 547
>NUP2_YEAST (P32499) Nucleoporin NUP2 (Nuclear pore protein NUP2)
(p95)
Length = 720
Score = 31.2 bits (69), Expect = 3.2
Identities = 22/100 (22%), Positives = 45/100 (45%), Gaps = 2/100 (2%)
Query: 34 FGFSTVTGTNFQNTSKRPKTMSIATDTATVTDSAMKEPFTKYTEYLNNLNDKRERVV--K 91
F FS+ T T Q SK P +++ AT T +S + FT T++ + + + V +
Sbjct: 247 FSFSSATSTTEQTKSKNPLSLTEATKTNVDNNSKAEASFTFGTKHAADSQNNKPSFVFGQ 306
Query: 92 ASRDITMNSKKVIFQVHRMSKYNKDEVLEKAEKDLAAVTN 131
A+ ++ F + K N + ++ + ++ +N
Sbjct: 307 AAAKPSLEKSSFTFGSTTIEKKNDENSTSNSKPEKSSDSN 346
>HEAD_BPGA1 (Q9FZW7) Major head protein
Length = 472
Score = 31.2 bits (69), Expect = 3.2
Identities = 24/97 (24%), Positives = 45/97 (45%), Gaps = 8/97 (8%)
Query: 188 LSDPSLQPLQINILDYILGLADLTGELMRLAIGRISDGELEFAEKICSFARDIYRELTLV 247
L+D +LQ I+ L +GL + +LM+ + G LE+ KI D+ RE
Sbjct: 52 LADKTLQNDFIHTLVDRIGLVVVHHKLMQNPLKIFKKGTLEYGRKIEEIFTDLTRE---- 107
Query: 248 VPHMDDSSDMKTKMETMLQSVMKIENVFMSEDQSIYH 284
H+ D K + E + + ++ +F D+ +++
Sbjct: 108 --HVYDPE--KAETEVFKREIPNVKTLFHERDRQVFY 140
>RPOA_MYCSP (P38018) DNA-directed RNA polymerase alpha chain (EC
2.7.7.6) (RNAP alpha subunit) (Transcriptase alpha
chain) (RNA polymerase alpha subunit)
Length = 353
Score = 30.8 bits (68), Expect = 4.1
Identities = 22/82 (26%), Positives = 40/82 (47%), Gaps = 2/82 (2%)
Query: 77 EYLNNLNDKRERVVKASRDITMNSKKVIFQ-VHRMSKYNK-DEVLEKAEKDLAAVTNQHV 134
E +K+ER +K++ IT V + R +KYN +EVL ++++L+ + N
Sbjct: 251 ELFEENKEKKEREIKSTTPITKLGLSVRSENALRRAKYNTVEEVLGLSDEELSNIKNLGK 310
Query: 135 SRLVKELQGTDFWKLRRAYSPG 156
+ ++ + WK R Y G
Sbjct: 311 KSIQDIIEKRNEWKERIGYDDG 332
>PIB1_BOVIN (P10894) 1-phosphatidylinositol-4,5-bisphosphate
phosphodiesterase beta 1 (EC 3.1.4.11) (Phosphoinositide
phospholipase C) (PLC-beta-1) (Phospholipase C-beta-1)
(PLC-I) (PLC-154)
Length = 1216
Score = 30.8 bits (68), Expect = 4.1
Identities = 36/129 (27%), Positives = 56/129 (42%), Gaps = 20/129 (15%)
Query: 64 TDSAMKEPFTKYTEYLNNLNDKRERVVKASRDITMNSKKVIFQVHRMSKYNKDEVLEKAE 123
T +KE TKY E N+ +R + K ++ ++KK + + D V E
Sbjct: 941 TTDLIKEHTTKYNEIQNDYLRRRAALEKTAK---KDNKK------KSEPSSPDHVSSTIE 991
Query: 124 KDLAAVTNQHVSRLV--KELQGTDFWKLRRA--YSPGIQ--EYV-----EAATFCSFCKN 172
+DLAA+ + +LV K+ Q LR+ YS Q E++ + C+N
Sbjct: 992 QDLAALDAEMTQKLVDLKDKQQQQLLNLRQEQYYSEKYQKREHIKLLIQKLTDVAEECQN 1051
Query: 173 GTLLKLDEI 181
L KL EI
Sbjct: 1052 NQLKKLKEI 1060
>OE66_NPVOP (O10305) Occlusion-derived virus envelope protein E66
(ODV-E66)
Length = 682
Score = 30.4 bits (67), Expect = 5.4
Identities = 13/38 (34%), Positives = 23/38 (60%)
Query: 80 NNLNDKRERVVKASRDITMNSKKVIFQVHRMSKYNKDE 117
N+LN+ RVV SRD ++N+ + F+ R++ N +
Sbjct: 487 NSLNNVNGRVVVLSRDTSVNTNDLSFEAQRLNNNNSSD 524
>MYS2_SCHPO (Q9USI6) Myosin type II heavy chain 1
Length = 1526
Score = 30.4 bits (67), Expect = 5.4
Identities = 26/125 (20%), Positives = 56/125 (44%), Gaps = 9/125 (7%)
Query: 42 TNFQNTSKRPKTMSIATDTATVTDSAMKEPFTKYTEYLNNLNDKRERVVKASRDITMNS- 100
TNFQ SK+ + ++ ++ ++ KE + + +L++K + K +++ +
Sbjct: 1189 TNFQELSKKHRDLTFNHESLLRQSASYKEKLSLASSENKDLSNKVSSLTKQVNELSPKAS 1248
Query: 101 ------KKVIFQVHRMSKYNKDEVLEKAEKDLAAVTNQHVSRLVKELQGTDFWKLRRAYS 154
+K+ +H S+ K EK + +A+ N+ + L EL+ KL Y
Sbjct: 1249 KVPELERKITNLMHEYSQLGKTFEDEKRKALIASRDNEELRSLKSELESKR--KLEVEYQ 1306
Query: 155 PGIQE 159
++E
Sbjct: 1307 KVLEE 1311
>MAK_RAT (P20793) Serine/threonine-protein kinase MAK (EC 2.7.1.37)
(Male germ cell-associated kinase)
Length = 622
Score = 30.4 bits (67), Expect = 5.4
Identities = 19/69 (27%), Positives = 31/69 (44%)
Query: 32 FDFGFSTVTGTNFQNTSKRPKTMSIATDTATVTDSAMKEPFTKYTEYLNNLNDKRERVVK 91
F F + + +N R S+ +DT T S K+ + K + YL +N K +V
Sbjct: 438 FRFPEAGLPVSNHLKGENRNLHASLKSDTNLSTASTAKQYYLKQSRYLPGVNPKNVSLVA 497
Query: 92 ASRDITMNS 100
+DI +S
Sbjct: 498 GGKDINSHS 506
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.133 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,310,204
Number of Sequences: 164201
Number of extensions: 1178582
Number of successful extensions: 3833
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 18
Number of HSP's that attempted gapping in prelim test: 3814
Number of HSP's gapped (non-prelim): 35
length of query: 293
length of database: 59,974,054
effective HSP length: 109
effective length of query: 184
effective length of database: 42,076,145
effective search space: 7742010680
effective search space used: 7742010680
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC144726.5


