

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144726.3 - phase: 0
(151 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
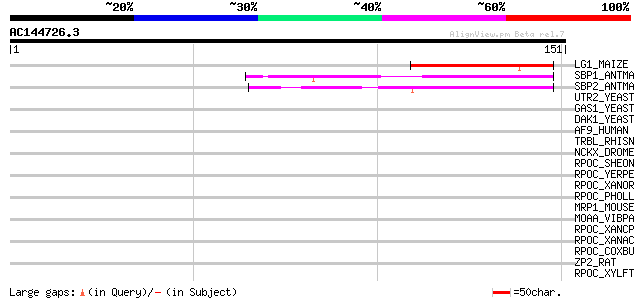
LG1_MAIZE (O04003) LIGULELESS1 protein 49 3e-06
SBP1_ANTMA (Q38741) Squamosa-promoter binding protein 1 47 1e-05
SBP2_ANTMA (Q38740) Squamosa-promoter binding protein 2 45 5e-05
UTR2_YEAST (P32623) UTR2 protein (Unknown transcript 2 protein) 35 0.048
GAS1_YEAST (P22146) Glycolipid anchored surface protein 1 precur... 32 0.41
DAK1_YEAST (P54838) Dihydroxyacetone kinase 1 (EC 2.7.1.29) (Gly... 32 0.70
AF9_HUMAN (P42568) AF-9 protein 32 0.70
TRBL_RHISN (P55402) Probable conjugal transfer protein trbL 31 1.2
NCKX_DROME (Q9U6A0) Sodium/potassium/calcium exchanger (Na(+)/K(... 31 1.2
RPOC_SHEON (Q8EK73) DNA-directed RNA polymerase beta' chain (EC ... 30 2.0
RPOC_YERPE (Q8D1H3) DNA-directed RNA polymerase beta' chain (EC ... 30 2.6
RPOC_XANOR (Q8KTH8) DNA-directed RNA polymerase beta' chain (EC ... 30 2.6
RPOC_PHOLL (Q7N9A3) DNA-directed RNA polymerase beta' chain (EC ... 30 2.6
MRP1_MOUSE (O35379) Multidrug resistance-associated protein 1 (A... 30 2.6
MOAA_VIBPA (Q87MY0) Molybdenum cofactor biosynthesis protein A 30 2.6
RPOC_XANCP (Q8PC55) DNA-directed RNA polymerase beta' chain (EC ... 29 3.5
RPOC_XANAC (Q8PNS9) DNA-directed RNA polymerase beta' chain (EC ... 29 3.5
RPOC_COXBU (Q83ET0) DNA-directed RNA polymerase beta' chain (EC ... 29 4.5
ZP2_RAT (O54767) Zona pellucida sperm-binding protein 2 precurso... 28 5.9
RPOC_XYLFT (Q87A33) DNA-directed RNA polymerase beta' chain (EC ... 28 5.9
>LG1_MAIZE (O04003) LIGULELESS1 protein
Length = 399
Score = 49.3 bits (116), Expect = 3e-06
Identities = 24/40 (60%), Positives = 29/40 (72%), Gaps = 1/40 (2%)
Query: 110 ADGCQADLSNYRGYHMRHKVCKLHSKTS-QVTLGGHKQRF 148
A+GC+ADLS+ + YH RHKVC+ HSK VT GG QRF
Sbjct: 187 AEGCKADLSSAKRYHRRHKVCEHHSKAPVVVTAGGLHQRF 226
>SBP1_ANTMA (Q38741) Squamosa-promoter binding protein 1
Length = 131
Score = 47.4 bits (111), Expect = 1e-05
Identities = 29/88 (32%), Positives = 45/88 (50%), Gaps = 16/88 (18%)
Query: 65 SAESGLTKSKDVGGFTK----ITSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSNY 120
S E G + D+G +K +T S SGS++++ + N C A+++N
Sbjct: 17 SQEQG-EEEDDIGEDSKKTRALTPSGKRASGSTQRSCQVEN-----------CAAEMTNA 64
Query: 121 RGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
+ YH RHKVC+ H+K V G +QRF
Sbjct: 65 KPYHRRHKVCEFHAKAPVVLHSGLQQRF 92
>SBP2_ANTMA (Q38740) Squamosa-promoter binding protein 2
Length = 171
Score = 45.4 bits (106), Expect = 5e-05
Identities = 30/86 (34%), Positives = 43/86 (49%), Gaps = 12/86 (13%)
Query: 66 AESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSS---LADGCQADLSNYRG 122
AES L K K + S SG A +G +V+ L + C ADL N +
Sbjct: 49 AESQLKKKK-----LNLGEGSGGKSGEKHTA----SGGGVVAQPCCLVENCGADLRNCKK 99
Query: 123 YHMRHKVCKLHSKTSQVTLGGHKQRF 148
Y+ RH+VC++H+K V++ G QRF
Sbjct: 100 YYQRHRVCEVHAKAPVVSVEGLMQRF 125
>UTR2_YEAST (P32623) UTR2 protein (Unknown transcript 2 protein)
Length = 347
Score = 35.4 bits (80), Expect = 0.048
Identities = 20/47 (42%), Positives = 27/47 (56%)
Query: 64 SSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLA 110
SS S TK T +SSSSS+S SS + A NG ++VSS++
Sbjct: 251 SSDHSSSTKKSSKTSSTASSSSSSSSSSSSSSSTATKNGDKVVSSVS 297
>GAS1_YEAST (P22146) Glycolipid anchored surface protein 1 precursor
(Glycoprotein GP115)
Length = 559
Score = 32.3 bits (72), Expect = 0.41
Identities = 19/62 (30%), Positives = 36/62 (57%), Gaps = 4/62 (6%)
Query: 57 KLGHVGNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGCQAD 116
++G +G +SA S D+G T+ +++SS+ SGSS K+ + ++G+ SS + +
Sbjct: 468 EIGSMGTNSASG----SVDLGSGTESSTASSNASGSSSKSNSGSSGSSSSSSSSSASSSS 523
Query: 117 LS 118
S
Sbjct: 524 SS 525
Score = 29.3 bits (64), Expect = 3.5
Identities = 16/38 (42%), Positives = 22/38 (57%)
Query: 63 NSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAIN 100
N+S S + S G + +SSS+S+S SSKK A N
Sbjct: 495 NASGSSSKSNSGSSGSSSSSSSSSASSSSSSKKNAATN 532
>DAK1_YEAST (P54838) Dihydroxyacetone kinase 1 (EC 2.7.1.29)
(Glycerone kinase 1) (DHA kinase 1)
Length = 584
Score = 31.6 bits (70), Expect = 0.70
Identities = 22/90 (24%), Positives = 38/90 (41%), Gaps = 8/90 (8%)
Query: 57 KLGHVGNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGCQAD 116
K G VG + + K VG F + SS G++K A+ IN+ + S D C+
Sbjct: 141 KGGMVGRRALAGTVLVHKIVGAFAEEYSSKYGLDGTAKVAKIINDNLVTIGSSLDHCKVP 200
Query: 117 LSNYRGYHMRHKVCKLHSKTSQVTLGGHKQ 146
+ +L+ K ++ +G H +
Sbjct: 201 GRKFES--------ELNEKQMELGMGIHNE 222
>AF9_HUMAN (P42568) AF-9 protein
Length = 568
Score = 31.6 bits (70), Expect = 0.70
Identities = 19/82 (23%), Positives = 38/82 (46%)
Query: 67 ESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSNYRGYHMR 126
++G ++ + + +SSSSS+S SS + + ++ + SS + + S+ +
Sbjct: 137 KAGGDPNRSIHTSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSSTSFSKP 196
Query: 127 HKVCKLHSKTSQVTLGGHKQRF 148
HK+ K H + HK F
Sbjct: 197 HKLMKEHKEKPSKDSREHKSAF 218
>TRBL_RHISN (P55402) Probable conjugal transfer protein trbL
Length = 391
Score = 30.8 bits (68), Expect = 1.2
Identities = 18/51 (35%), Positives = 24/51 (46%)
Query: 78 GFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSNYRGYHMRHK 128
GF S+SS +GS+ K +AI + S+ A L RG H HK
Sbjct: 334 GFGAGIGSASSAAGSAAKEKAIGSPGAYAGSILGLANAKLDESRGGHSGHK 384
>NCKX_DROME (Q9U6A0) Sodium/potassium/calcium exchanger
(Na(+)/K(+)/Ca(2+)-exchange protein)
Length = 856
Score = 30.8 bits (68), Expect = 1.2
Identities = 28/116 (24%), Positives = 46/116 (39%), Gaps = 21/116 (18%)
Query: 7 CLFNVCSILYIVLFLNNDEVFWRPWHDSI-WNILYGFVCF-------SMCANEFFVDLKL 58
C F S+L ++ F ++ +FW W I + I G+V F C + K+
Sbjct: 440 CSFYSISLLVLIYFFRDNRIFW--WEALILFTIYIGYVAFMKWNVQVETCVKKMITKNKV 497
Query: 59 GHV---------GNSSAESGLTKSKDVGGFTKITSSSSSTSG--SSKKARAINNGT 103
V GN++ S + + GG ++S + SG S A A N +
Sbjct: 498 TRVRSTDQLMPAGNAANSSETSMATQPGGSVTSRAASETRSGPPGSSNAGATGNSS 553
>RPOC_SHEON (Q8EK73) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta'
chain) (RNA polymerase beta' subunit)
Length = 1405
Score = 30.0 bits (66), Expect = 2.0
Identities = 17/58 (29%), Positives = 30/58 (51%), Gaps = 1/58 (1%)
Query: 45 FSMCANEFFVDLKLGHVGNSSAESGLTKSKDVG-GFTKITSSSSSTSGSSKKARAINN 101
F +CA + DL GH+ N G+ ++ +G T++T + G++ +A A NN
Sbjct: 892 FGVCAACYGRDLARGHIINHGEAIGVVAAQSIGEPGTQLTMRTFHIGGAASRASAENN 949
>RPOC_YERPE (Q8D1H3) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta'
chain) (RNA polymerase beta' subunit)
Length = 1406
Score = 29.6 bits (65), Expect = 2.6
Identities = 16/55 (29%), Positives = 29/55 (52%), Gaps = 1/55 (1%)
Query: 45 FSMCANEFFVDLKLGHVGNSSAESGLTKSKDVG-GFTKITSSSSSTSGSSKKARA 98
F +CAN + DL GH+ N G+ ++ +G T++T + G++ +A A
Sbjct: 892 FGVCANCYGRDLARGHIINKGEAVGVIAAQSIGEPGTQLTMRTFHIGGAASRAAA 946
>RPOC_XANOR (Q8KTH8) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta'
chain) (RNA polymerase beta' subunit)
Length = 1405
Score = 29.6 bits (65), Expect = 2.6
Identities = 17/60 (28%), Positives = 31/60 (51%), Gaps = 1/60 (1%)
Query: 45 FSMCANEFFVDLKLGHVGNSSAESGLTKSKDVGG-FTKITSSSSSTSGSSKKARAINNGT 103
F +CA + DL GH N G+ ++ +G T++T + G++ +A A++N T
Sbjct: 894 FGVCARCYGRDLARGHQVNIGEAVGVIAAQSIGDPGTQLTMRTFHIGGAASRAAAVDNIT 953
>RPOC_PHOLL (Q7N9A3) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta'
chain) (RNA polymerase beta' subunit)
Length = 1406
Score = 29.6 bits (65), Expect = 2.6
Identities = 16/55 (29%), Positives = 29/55 (52%), Gaps = 1/55 (1%)
Query: 45 FSMCANEFFVDLKLGHVGNSSAESGLTKSKDVG-GFTKITSSSSSTSGSSKKARA 98
F +CAN + DL GH+ N G+ ++ +G T++T + G++ +A A
Sbjct: 892 FGVCANCYGRDLARGHIINKGEAVGVIAAQSIGEPGTQLTMRTFHIGGAASRAAA 946
>MRP1_MOUSE (O35379) Multidrug resistance-associated protein 1
(ATP-binding cassette, sub-family C, member 1)
Length = 1528
Score = 29.6 bits (65), Expect = 2.6
Identities = 16/65 (24%), Positives = 30/65 (45%), Gaps = 2/65 (3%)
Query: 82 ITSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTL 141
+ S S SGS K+++ + NG + ++ Q LSN + + HS +++
Sbjct: 876 LASEDDSVSGSGKESKPVENGMLVTDTVGKHLQRHLSNSSSH--SGDTSQQHSSIAELQK 933
Query: 142 GGHKQ 146
G K+
Sbjct: 934 AGAKE 938
>MOAA_VIBPA (Q87MY0) Molybdenum cofactor biosynthesis protein A
Length = 334
Score = 29.6 bits (65), Expect = 2.6
Identities = 24/78 (30%), Positives = 38/78 (47%), Gaps = 9/78 (11%)
Query: 50 NEFFVDL-KLGHVGNSSAESGLTKSKDVGG-------FTKITSSSSSTSGSSKKARAINN 101
N F+ L ++ V + A+ G +K + GG FT I S ++T G KK N
Sbjct: 48 NSSFLSLPEIKRVVKAFADCGTSKVRITGGEPSLRKDFTDIIHSVATTPGI-KKVATTTN 106
Query: 102 GTQIVSSLADGCQADLSN 119
G ++ +AD +A L+N
Sbjct: 107 GYRMAKQVADWREAGLTN 124
>RPOC_XANCP (Q8PC55) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta'
chain) (RNA polymerase beta' subunit)
Length = 1405
Score = 29.3 bits (64), Expect = 3.5
Identities = 17/60 (28%), Positives = 31/60 (51%), Gaps = 1/60 (1%)
Query: 45 FSMCANEFFVDLKLGHVGNSSAESGLTKSKDVG-GFTKITSSSSSTSGSSKKARAINNGT 103
F +CA + DL GH N G+ ++ +G T++T + G++ +A A++N T
Sbjct: 894 FGVCARCYGRDLARGHQVNIGEAVGVIAAQSIGEPGTQLTMRTFHIGGAASRAAAVDNIT 953
>RPOC_XANAC (Q8PNS9) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta'
chain) (RNA polymerase beta' subunit)
Length = 1404
Score = 29.3 bits (64), Expect = 3.5
Identities = 17/60 (28%), Positives = 31/60 (51%), Gaps = 1/60 (1%)
Query: 45 FSMCANEFFVDLKLGHVGNSSAESGLTKSKDVG-GFTKITSSSSSTSGSSKKARAINNGT 103
F +CA + DL GH N G+ ++ +G T++T + G++ +A A++N T
Sbjct: 893 FGVCARCYGRDLARGHQVNIGEAVGVIAAQSIGEPGTQLTMRTFHIGGAASRAAAVDNIT 952
>RPOC_COXBU (Q83ET0) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta'
chain) (RNA polymerase beta' subunit)
Length = 1414
Score = 28.9 bits (63), Expect = 4.5
Identities = 16/58 (27%), Positives = 30/58 (51%), Gaps = 1/58 (1%)
Query: 45 FSMCANEFFVDLKLGHVGNSSAESGLTKSKDVG-GFTKITSSSSSTSGSSKKARAINN 101
+ +C+ + DL GHV N G+ ++ +G T++T + G++ +A A NN
Sbjct: 892 YGICSMCYGRDLARGHVVNVGEAIGVVAAQSIGEPGTQLTMRTFHIGGAASRATAANN 949
>ZP2_RAT (O54767) Zona pellucida sperm-binding protein 2 precursor
(Zona pellucida glycoprotein ZP2) (Zona pellucida
protein A)
Length = 695
Score = 28.5 bits (62), Expect = 5.9
Identities = 14/40 (35%), Positives = 18/40 (45%), Gaps = 8/40 (20%)
Query: 109 LADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
+ DGC+ +L NYR H+ S GH QRF
Sbjct: 544 IMDGCEYELDNYR--------TTFHAANSSAAHSGHYQRF 575
>RPOC_XYLFT (Q87A33) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6) (RNAP beta' subunit) (Transcriptase beta'
chain) (RNA polymerase beta' subunit)
Length = 1407
Score = 28.5 bits (62), Expect = 5.9
Identities = 18/60 (30%), Positives = 31/60 (51%), Gaps = 1/60 (1%)
Query: 45 FSMCANEFFVDLKLGHVGNSSAESGLTKSKDVG-GFTKITSSSSSTSGSSKKARAINNGT 103
F +CA + DL GH+ N G+ ++ +G T++T + G++ A AI+N T
Sbjct: 893 FGVCALCYGRDLARGHLVNMGEAVGVIAAQSIGEPGTQLTMRTFHIGGTALSAAAIDNIT 952
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.137 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,128,245
Number of Sequences: 164201
Number of extensions: 617240
Number of successful extensions: 3057
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 10
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 3027
Number of HSP's gapped (non-prelim): 49
length of query: 151
length of database: 59,974,054
effective HSP length: 100
effective length of query: 51
effective length of database: 43,553,954
effective search space: 2221251654
effective search space used: 2221251654
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144726.3


