

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
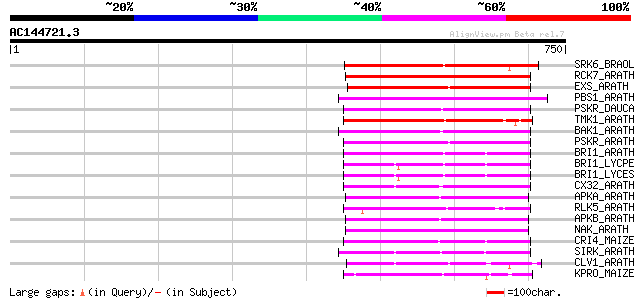
Query= AC144721.3 - phase: 0 /pseudo
(750 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 307 8e-83
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 194 7e-49
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 192 4e-48
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 190 1e-47
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 189 2e-47
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 182 3e-45
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 176 2e-43
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 174 8e-43
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 172 2e-42
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 172 4e-42
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 172 4e-42
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 163 1e-39
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 163 1e-39
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 157 7e-38
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 157 7e-38
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 156 2e-37
CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4 pr... 156 2e-37
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 150 2e-35
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 149 3e-35
KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1 precu... 144 9e-34
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 307 bits (786), Expect = 8e-83
Identities = 151/269 (56%), Positives = 199/269 (73%), Gaps = 8/269 (2%)
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
DG+EIAVKRLSK S QG +EFMNEV +I++LQH NLV++LGCC+E EKML+YE++ N S
Sbjct: 549 DGKEIAVKRLSKTSVQGTDEFMNEVTLIARLQHINLVQVLGCCIEGDEKMLIYEYLENLS 608
Query: 513 LDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
LD++LF ++ KL+W +R +I G+ARG++YLH+DSR +IIHRDLK SNILLD MIPK
Sbjct: 609 LDSYLFGKTRRSKLNWNERFDITNGVARGLLYLHQDSRFRIIHRDLKVSNILLDKNMIPK 668
Query: 573 ISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
ISDFG+ARI + E EANT +VVGTYGYM PEYAM G+FSEKSDV+SFGV++LEIVSG+
Sbjct: 669 ISDFGMARIFERDE-TEANTMKVVGTYGYMSPEYAMYGIFSEKSDVFSFGVIVLEIVSGK 727
Query: 633 RNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFE-------SSMLRCMHIGLL 685
+N FY + L+ + W W E + ++D + D+ +L+C+ IGLL
Sbjct: 728 KNRGFYNLDYENDLLSYVWSRWKEGRALEIVDPVIVDSLSSQPSIFQPQEVLKCIQIGLL 787
Query: 686 CVQELPKERPSISTVVLMLISEITHLPPP 714
CVQEL + RP++S+VV M SE T +P P
Sbjct: 788 CVQELAEHRPAMSSVVWMFGSEATEIPQP 816
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 194 bits (493), Expect = 7e-49
Identities = 109/252 (43%), Positives = 155/252 (61%), Gaps = 3/252 (1%)
Query: 455 QEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSLD 514
Q +A+K+L + QGI EF+ EV+ +S H NLV+L+G C E +++LVYE+MP SL+
Sbjct: 127 QVVAIKQLDRNGVQGIREFVVEVLTLSLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLE 186
Query: 515 AFLFD-PIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKI 573
L P KK LDW R I G ARG+ YLH +I+RDLK SNILL ++ PK+
Sbjct: 187 DHLHVLPSGKKPLDWNTRMKIAAGAARGLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKL 246
Query: 574 SDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRR 633
SDFGLA++ G+ +T RV+GTYGY P+YAM G + KSD+YSFGV+LLE+++GR+
Sbjct: 247 SDFGLAKVGPSGDKTHVST-RVMGTYGYCAPDYAMTGQLTFKSDIYSFGVVLLELITGRK 305
Query: 634 NNSFYQNEDSLSLVGFAWKLWLE-ENTISLIDREVWDASFESSMLRCMHIGLLCVQELPK 692
+ +LVG+A L+ + N ++D + + + + I +CVQE P
Sbjct: 306 AIDNTKTRKDQNLVGWARPLFKDRRNFPKMVDPLLQGQYPVRGLYQALAISAMCVQEQPT 365
Query: 693 ERPSISTVVLML 704
RP +S VVL L
Sbjct: 366 MRPVVSDVVLAL 377
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 192 bits (487), Expect = 4e-48
Identities = 106/250 (42%), Positives = 155/250 (61%), Gaps = 4/250 (1%)
Query: 457 IAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSLDAF 516
+AVK+LS+A QG EFM E+ + K++H NLV LLG C EK+LVYE+M N SLD +
Sbjct: 942 VAVKKLSEAKTQGNREFMAEMETLGKVKHPNLVSLLGYCSFSEEKLLVYEYMVNGSLDHW 1001
Query: 517 LFDPI-QKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISD 575
L + + LDW KR I G ARG+ +LH IIHRD+KASNILLD + PK++D
Sbjct: 1002 LRNQTGMLEVLDWSKRLKIAVGAARGLAFLHHGFIPHIIHRDIKASNILLDGDFEPKVAD 1061
Query: 576 FGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRR-N 634
FGLAR++ E + + GT+GY+PPEY + K DVYSFGV+LLE+V+G+
Sbjct: 1062 FGLARLISACESHVSTV--IAGTFGYIPPEYGQSARATTKGDVYSFGVILLELVTGKEPT 1119
Query: 635 NSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKER 694
++ + +LVG+A + + + +ID + + ++S LR + I +LC+ E P +R
Sbjct: 1120 GPDFKESEGGNLVGWAIQKINQGKAVDVIDPLLVSVALKNSQLRLLQIAMLCLAETPAKR 1179
Query: 695 PSISTVVLML 704
P++ V+ L
Sbjct: 1180 PNMLDVLKAL 1189
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 190 bits (482), Expect = 1e-47
Identities = 114/286 (39%), Positives = 164/286 (56%), Gaps = 5/286 (1%)
Query: 445 GRFWPGIQD--GQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKM 502
GR + G D GQ +AVK+L + QG EF+ EV+++S L H NLV L+G C + +++
Sbjct: 98 GRVYKGRLDSTGQVVAVKQLDRNGLQGNREFLVEVLMLSLLHHPNLVNLIGYCADGDQRL 157
Query: 503 LVYEFMPNKSLDAFLFD-PIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKAS 561
LVYEFMP SL+ L D P K+ LDW R I G A+G+ +LH + +I+RD K+S
Sbjct: 158 LVYEFMPLGSLEDHLHDLPPDKEALDWNMRMKIAAGAAKGLEFLHDKANPPVIYRDFKSS 217
Query: 562 NILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSF 621
NILLD+ PK+SDFGLA++ G+ +T RV+GTYGY PEYAM G + KSDVYSF
Sbjct: 218 NILLDEGFHPKLSDFGLAKLGPTGDKSHVST-RVMGTYGYCAPEYAMTGQLTVKSDVYSF 276
Query: 622 GVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENT-ISLIDREVWDASFESSMLRCM 680
GV+ LE+++GR+ +LV +A L+ + I L D + ++ + +
Sbjct: 277 GVVFLELITGRKAIDSEMPHGEQNLVAWARPLFNDRRKFIKLADPRLKGRFPTRALYQAL 336
Query: 681 HIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSR 726
+ +C+QE RP I+ VV L P K N++ R
Sbjct: 337 AVASMCIQEQAATRPLIADVVTALSYLANQAYDPSKDDSRRNRDER 382
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 189 bits (480), Expect = 2e-47
Identities = 101/255 (39%), Positives = 148/255 (57%), Gaps = 3/255 (1%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG ++A+KRLS +GQ EF EV +S+ QH NLV LLG C + +K+L+Y +M N
Sbjct: 762 LPDGTKVAIKRLSGDTGQMDREFQAEVETLSRAQHPNLVHLLGYCNYKNDKLLIYSYMDN 821
Query: 511 KSLDAFLFDPIQ-KKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEM 569
SLD +L + + LDW+ R I G A G+ YLH+ I+HRD+K+SNILL D
Sbjct: 822 GSLDYWLHEKVDGPPSLDWKTRLRIARGAAEGLAYLHQSCEPHILHRDIKSSNILLSDTF 881
Query: 570 IPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIV 629
+ ++DFGLAR++ D T +VGT GY+PPEY + + K DVYSFGV+LLE++
Sbjct: 882 VAHLADFGLARLIL--PYDTHVTTDLVGTLGYIPPEYGQASVATYKGDVYSFGVVLLELL 939
Query: 630 SGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQE 689
+GRR + S L+ + ++ E+ + D ++D ML + I C+ E
Sbjct: 940 TGRRPMDVCKPRGSRDLISWVLQMKTEKRESEIFDPFIYDKDHAEEMLLVLEIACRCLGE 999
Query: 690 LPKERPSISTVVLML 704
PK RP+ +V L
Sbjct: 1000 NPKTRPTTQQLVSWL 1014
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 182 bits (462), Expect = 3e-45
Identities = 105/267 (39%), Positives = 164/267 (61%), Gaps = 17/267 (6%)
Query: 451 IQDGQEIAVKRLSKA--SGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFM 508
+ DG +IAVKR+ +G+G EF +E+ V++K++HR+LV LLG C++ EK+LVYE+M
Sbjct: 607 LHDGTKIAVKRMENGVIAGKGFAEFKSEIAVLTKVRHRHLVTLLGYCLDGNEKLLVYEYM 666
Query: 509 PNKSLDAFLFDPIQK--KKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLD 566
P +L LF+ ++ K L W++R + +ARG+ YLH + IHRDLK SNILL
Sbjct: 667 PQGTLSRHLFEWSEEGLKPLLWKQRLTLALDVARGVEYLHGLAHQSFIHRDLKPSNILLG 726
Query: 567 DEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLL 626
D+M K++DFGL R+ G+G + R+ GT+GY+ PEYA+ G + K DVYSFGV+L+
Sbjct: 727 DDMRAKVADFGLVRLAPEGKG--SIETRIAGTFGYLAPEYAVTGRVTTKVDVYSFGVILM 784
Query: 627 EIVSGRRNNSFYQNEDSLSLVGFAWKLWL--EENTISLIDREVWDASFESSMLRCMH--- 681
E+++GR++ Q E+S+ LV + ++++ E + ID + + L +H
Sbjct: 785 ELITGRKSLDESQPEESIHLVSWFKRMYINKEASFKKAIDTTI---DLDEETLASVHTVA 841
Query: 682 --IGLLCVQELPKERPSISTVVLMLIS 706
G C +E P +RP + V +L S
Sbjct: 842 ELAGHCCARE-PYQRPDMGHAVNILSS 867
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 176 bits (447), Expect = 2e-43
Identities = 108/265 (40%), Positives = 159/265 (59%), Gaps = 7/265 (2%)
Query: 445 GRFWPG-IQDGQEIAVKRLSKASGQGIE-EFMNEVVVISKLQHRNLVRLLGCCVERGEKM 502
G+ + G + DG +AVKRL + QG E +F EV +IS HRNL+RL G C+ E++
Sbjct: 301 GKVYKGRLADGTLVAVKRLKEERTQGGELQFQTEVEMISMAVHRNLLRLRGFCMTPTERL 360
Query: 503 LVYEFMPNKSLDAFLFD-PIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKAS 561
LVY +M N S+ + L + P + LDW KR I G ARG+ YLH KIIHRD+KA+
Sbjct: 361 LVYPYMANGSVASCLRERPESQPPLDWPKRQRIALGSARGLAYLHDHCDPKIIHRDVKAA 420
Query: 562 NILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSF 621
NILLD+E + DFGLA+++ D T V GT G++ PEY G SEK+DV+ +
Sbjct: 421 NILLDEEFEAVVGDFGLAKLM--DYKDTHVTTAVRGTIGHIAPEYLSTGKSSEKTDVFGY 478
Query: 622 GVLLLEIVSGRR--NNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRC 679
GV+LLE+++G+R + + N+D + L+ + L E+ +L+D ++ + + +
Sbjct: 479 GVMLLELITGQRAFDLARLANDDDVMLLDWVKGLLKEKKLEALVDVDLQGNYKDEEVEQL 538
Query: 680 MHIGLLCVQELPKERPSISTVVLML 704
+ + LLC Q P ERP +S VV ML
Sbjct: 539 IQVALLCTQSSPMERPKMSEVVRML 563
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 174 bits (441), Expect = 8e-43
Identities = 94/255 (36%), Positives = 147/255 (56%), Gaps = 3/255 (1%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG+++A+K+LS GQ EF EV +S+ QH NLV L G C + +++L+Y +M N
Sbjct: 753 LPDGKKVAIKKLSGDCGQIEREFEAEVETLSRAQHPNLVLLRGFCFYKNDRLLIYSYMEN 812
Query: 511 KSLDAFLFDPIQKKKL-DWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEM 569
SLD +L + L W+ R I +G A+G++YLH I+HRD+K+SNILLD+
Sbjct: 813 GSLDYWLHERNDGPALLKWKTRLRIAQGAAKGLLYLHEGCDPHILHRDIKSSNILLDENF 872
Query: 570 IPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIV 629
++DFGLAR++ E + +VGT GY+PPEY + + K DVYSFGV+LLE++
Sbjct: 873 NSHLADFGLARLMSPYETHVSTD--LVGTLGYIPPEYGQASVATYKGDVYSFGVVLLELL 930
Query: 630 SGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQE 689
+ +R + + L+ + K+ E + D ++ + M R + I LC+ E
Sbjct: 931 TDKRPVDMCKPKGCRDLISWVVKMKHESRASEVFDPLIYSKENDKEMFRVLEIACLCLSE 990
Query: 690 LPKERPSISTVVLML 704
PK+RP+ +V L
Sbjct: 991 NPKQRPTTQQLVSWL 1005
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 172 bits (437), Expect = 2e-42
Identities = 100/256 (39%), Positives = 153/256 (59%), Gaps = 6/256 (2%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
++DG +A+K+L SGQG EFM E+ I K++HRNLV LLG C E++LVYEFM
Sbjct: 902 LKDGSAVAIKKLIHVSGQGDREFMAEMETIGKIKHRNLVPLLGYCKVGDERLLVYEFMKY 961
Query: 511 KSLDAFLFDPIQK-KKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEM 569
SL+ L DP + KL+W R I G ARG+ +LH + IIHRD+K+SN+LLD+ +
Sbjct: 962 GSLEDVLHDPKKAGVKLNWSTRRKIAIGSARGLAFLHHNCSPHIIHRDMKSSNVLLDENL 1021
Query: 570 IPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIV 629
++SDFG+AR++ + + + GT GY+PPEY S K DVYS+GV+LLE++
Sbjct: 1022 EARVSDFGMARLMSAMD-THLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELL 1080
Query: 630 SGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVW--DASFESSMLRCMHIGLLCV 687
+G+R D+ +LVG+ K + + D E+ D + E +L+ + + + C+
Sbjct: 1081 TGKRPTDSPDFGDN-NLVGWV-KQHAKLRISDVFDPELMKEDPALEIELLQHLKVAVACL 1138
Query: 688 QELPKERPSISTVVLM 703
+ RP++ V+ M
Sbjct: 1139 DDRAWRRPTMVQVMAM 1154
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 172 bits (435), Expect = 4e-42
Identities = 102/258 (39%), Positives = 156/258 (59%), Gaps = 10/258 (3%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
++DG +A+K+L SGQG EF E+ I K++HRNLV LLG C E++LVYE+M
Sbjct: 907 LKDGSVVAIKKLIHVSGQGDREFTAEMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKY 966
Query: 511 KSLDAFLFDPIQKK---KLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDD 567
SL+ L D +KK KL+W R I G ARG+ +LH + IIHRD+K+SN+LLD+
Sbjct: 967 GSLEDVLHD--RKKTGIKLNWPARRKIAIGAARGLAFLHHNCIPHIIHRDMKSSNVLLDE 1024
Query: 568 EMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLE 627
+ ++SDFG+AR++ + + + GT GY+PPEY S K DVYS+GV+LLE
Sbjct: 1025 NLEARVSDFGMARLMSAMD-THLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLE 1083
Query: 628 IVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVW--DASFESSMLRCMHIGLL 685
+++G++ D+ +LVG+ KL + + DRE+ DAS E +L+ + +
Sbjct: 1084 LLTGKQPTDSADFGDN-NLVGWV-KLHAKGKITDVFDRELLKEDASIEIELLQHLKVACA 1141
Query: 686 CVQELPKERPSISTVVLM 703
C+ + +RP++ V+ M
Sbjct: 1142 CLDDRHWKRPTMIQVMAM 1159
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 172 bits (435), Expect = 4e-42
Identities = 102/258 (39%), Positives = 156/258 (59%), Gaps = 10/258 (3%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
++DG +A+K+L SGQG EF E+ I K++HRNLV LLG C E++LVYE+M
Sbjct: 907 LKDGSVVAIKKLIHVSGQGDREFTAEMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKY 966
Query: 511 KSLDAFLFDPIQKK---KLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDD 567
SL+ L D +KK KL+W R I G ARG+ +LH + IIHRD+K+SN+LLD+
Sbjct: 967 GSLEDVLHD--RKKIGIKLNWPARRKIAIGAARGLAFLHHNCIPHIIHRDMKSSNVLLDE 1024
Query: 568 EMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLE 627
+ ++SDFG+AR++ + + + GT GY+PPEY S K DVYS+GV+LLE
Sbjct: 1025 NLEARVSDFGMARLMSAMD-THLSVSTLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLE 1083
Query: 628 IVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVW--DASFESSMLRCMHIGLL 685
+++G++ D+ +LVG+ KL + + DRE+ DAS E +L+ + +
Sbjct: 1084 LLTGKQPTDSADFGDN-NLVGWV-KLHAKGKITDVFDRELLKEDASIEIELLQHLKVACA 1141
Query: 686 CVQELPKERPSISTVVLM 703
C+ + +RP++ V+ M
Sbjct: 1142 CLDDRHWKRPTMIQVMAM 1159
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 163 bits (413), Expect = 1e-39
Identities = 104/257 (40%), Positives = 146/257 (56%), Gaps = 9/257 (3%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ G +A+KRL+ S QG E+ +EV + L HRNLV+LLG C E E +LVYEFMP
Sbjct: 115 VGSGMIVAIKRLNSESVQGFAEWRSEVNFLGMLSHRNLVKLLGYCREDKELLLVYEFMPK 174
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
SL++ LF + W R IV G ARG+ +LH R ++I+RD KASNILLD
Sbjct: 175 GSLESHLFR--RNDPFPWDLRIKIVIGAARGLAFLHSLQR-EVIYRDFKASNILLDSNYD 231
Query: 571 PKISDFGLARIVKGGEGDEAN--TKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEI 628
K+SDFGLA++ G DE + T R++GTYGY PEY G KSDV++FGV+LLEI
Sbjct: 232 AKLSDFGLAKL---GPADEKSHVTTRIMGTYGYAAPEYMATGHLYVKSDVFAFGVVLLEI 288
Query: 629 VSGRRNNSFYQNEDSLSLVGFAW-KLWLEENTISLIDREVWDASFESSMLRCMHIGLLCV 687
++G ++ + SLV + +L + ++D+ + I L C+
Sbjct: 289 MTGLTAHNTKRPRGQESLVDWLRPELSNKHRVKQIMDKGIKGQYTTKVATEMARITLSCI 348
Query: 688 QELPKERPSISTVVLML 704
+ PK RP + VV +L
Sbjct: 349 EPDPKNRPHMKEVVEVL 365
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 163 bits (413), Expect = 1e-39
Identities = 101/251 (40%), Positives = 150/251 (59%), Gaps = 6/251 (2%)
Query: 454 GQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSL 513
G IAVK+L++ QG +E++ EV + + HR+LV+L+G C+E ++LVYEFMP SL
Sbjct: 100 GLVIAVKKLNQDGWQGHQEWLAEVNYLGQFSHRHLVKLIGYCLEDEHRLLVYEFMPRGSL 159
Query: 514 DAFLF-DPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
+ LF + + L W+ R + G A+G+ +LH S ++I+RD K SNILLD E K
Sbjct: 160 ENHLFRRGLYFQPLSWKLRLKVALGAAKGLAFLH-SSETRVIYRDFKTSNILLDSEYNAK 218
Query: 573 ISDFGLARIVKGGEGDEAN-TKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSG 631
+SDFGLA+ G GD+++ + RV+GT+GY PEY G + KSDVYSFGV+LLE++SG
Sbjct: 219 LSDFGLAK--DGPIGDKSHVSTRVMGTHGYAAPEYLATGHLTTKSDVYSFGVVLLELLSG 276
Query: 632 RRNNSFYQNEDSLSLVGFAWKLWLEENTI-SLIDREVWDASFESSMLRCMHIGLLCVQEL 690
RR + +LV +A + + I +ID + D + + L C+
Sbjct: 277 RRAVDKNRPSGERNLVEWAKPYLVNKRKIFRVIDNRLQDQYSMEEACKVATLSLRCLTTE 336
Query: 691 PKERPSISTVV 701
K RP++S VV
Sbjct: 337 IKLRPNMSEVV 347
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 157 bits (398), Expect = 7e-38
Identities = 103/269 (38%), Positives = 156/269 (57%), Gaps = 21/269 (7%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMN----------EVVVISKLQHRNLVRLLGCCVERGE 500
++ G+ +AVK+L+K+ G +E+ + EV + ++H+++VRL CC
Sbjct: 702 LRGGEVVAVKKLNKSVKGGDDEYSSDSLNRDVFAAEVETLGTIRHKSIVRLWCCCSSGDC 761
Query: 501 KMLVYEFMPNKSL-DAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
K+LVYE+MPN SL D D L W +R I A G+ YLH D I+HRD+K
Sbjct: 762 KLLVYEYMPNGSLADVLHGDRKGGVVLGWPERLRIALDAAEGLSYLHHDCVPPIVHRDVK 821
Query: 560 ASNILLDDEMIPKISDFGLARI--VKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSD 617
+SNILLD + K++DFG+A++ + G + EA + G+ GY+ PEY +EKSD
Sbjct: 822 SSNILLDSDYGAKVADFGIAKVGQMSGSKTPEA-MSGIAGSCGYIAPEYVYTLRVNEKSD 880
Query: 618 VYSFGVLLLEIVSGRR-NNSFYQNEDSLSLVGFAW-KLWLEENTISLIDREVWDASFESS 675
+YSFGV+LLE+V+G++ +S ++D V A K LE +ID ++ D F+
Sbjct: 881 IYSFGVVLLELVTGKQPTDSELGDKDMAKWVCTALDKCGLE----PVIDPKL-DLKFKEE 935
Query: 676 MLRCMHIGLLCVQELPKERPSISTVVLML 704
+ + +HIGLLC LP RPS+ VV+ML
Sbjct: 936 ISKVIHIGLLCTSPLPLNRPSMRKVVIML 964
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 157 bits (398), Expect = 7e-38
Identities = 98/251 (39%), Positives = 146/251 (58%), Gaps = 6/251 (2%)
Query: 454 GQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSL 513
G IAVK+L++ QG +E++ EV + + H NLV+L+G C+E ++LVYEFMP SL
Sbjct: 101 GVVIAVKKLNQDGWQGHQEWLAEVNYLGQFSHPNLVKLIGYCLEDEHRLLVYEFMPRGSL 160
Query: 514 DAFLF-DPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
+ LF + L W R + G A+G+ +LH ++ +I+RD K SNILLD E K
Sbjct: 161 ENHLFRRGSYFQPLSWTLRLKVALGAAKGLAFLH-NAETSVIYRDFKTSNILLDSEYNAK 219
Query: 573 ISDFGLARIVKGGEGDEAN-TKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSG 631
+SDFGLA+ G GD+++ + R++GTYGY PEY G + KSDVYS+GV+LLE++SG
Sbjct: 220 LSDFGLAK--DGPTGDKSHVSTRIMGTYGYAAPEYLATGHLTTKSDVYSYGVVLLEVLSG 277
Query: 632 RRNNSFYQNEDSLSLVGFAWKLWLEENTI-SLIDREVWDASFESSMLRCMHIGLLCVQEL 690
RR + LV +A L + + +ID + D + + L C+
Sbjct: 278 RRAVDKNRPPGEQKLVEWARPLLANKRKLFRVIDNRLQDQYSMEEACKVATLALRCLTFE 337
Query: 691 PKERPSISTVV 701
K RP+++ VV
Sbjct: 338 IKLRPNMNEVV 348
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 156 bits (394), Expect = 2e-37
Identities = 100/250 (40%), Positives = 145/250 (58%), Gaps = 4/250 (1%)
Query: 454 GQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSL 513
G IAVKRL++ QG E++ E+ + +L H NLV+L+G C+E ++LVYEFM SL
Sbjct: 100 GIVIAVKRLNQEGFQGHREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLVYEFMTRGSL 159
Query: 514 DAFLF-DPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPK 572
+ LF + L W R + G ARG+ +LH +++ ++I+RD KASNILLD K
Sbjct: 160 ENHLFRRGTFYQPLSWNTRVRMALGAARGLAFLH-NAQPQVIYRDFKASNILLDSNYNAK 218
Query: 573 ISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
+SDFGLAR G+ +T RV+GT GY PEY G S KSDVYSFGV+LLE++SGR
Sbjct: 219 LSDFGLARDGPMGDNSHVST-RVMGTQGYAAPEYLATGHLSVKSDVYSFGVVLLELLSGR 277
Query: 633 RNNSFYQNEDSLSLVGFAWK-LWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELP 691
R Q +LV +A L + + ++D + + L+ + L C+
Sbjct: 278 RAIDKNQPVGEHNLVDWARPYLTNKRRLLRVMDPRLQGQYSLTRALKIAVLALDCISIDA 337
Query: 692 KERPSISTVV 701
K RP+++ +V
Sbjct: 338 KSRPTMNEIV 347
>CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4
precursor (EC 2.7.1.-)
Length = 901
Score = 156 bits (394), Expect = 2e-37
Identities = 98/258 (37%), Positives = 151/258 (57%), Gaps = 7/258 (2%)
Query: 451 IQDGQEIAVKRLSKASG--QGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFM 508
++DG +AVKR KAS + +EF NE+ ++S+L H +L+ LLG C + E++LVYEFM
Sbjct: 524 LRDGTVVAVKRAIKASDVKKSSKEFHNELDLLSRLNHAHLLNLLGYCEDGSERLLVYEFM 583
Query: 509 PNKSLDAFLF--DPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLD 566
+ SL L DP KK+L+W +R I ARGI YLH + +IHRD+K+SNIL+D
Sbjct: 584 AHGSLYQHLHGKDPNLKKRLNWARRVTIAVQAARGIEYLHGYACPPVIHRDIKSSNILID 643
Query: 567 DEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLL 626
++ +++DFGL+ I+ + ++ GT GY+ PEY + KSDVYSFGV+LL
Sbjct: 644 EDHNARVADFGLS-ILGPADSGTPLSELPAGTLGYLDPEYYRLHYLTTKSDVYSFGVVLL 702
Query: 627 EIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLC 686
EI+SGR+ E ++V +A L + +++D + S ++ + + C
Sbjct: 703 EILSGRKAIDMQFEEG--NIVEWAVPLIKAGDIFAILDPVLSPPSDLEALKKIASVACKC 760
Query: 687 VQELPKERPSISTVVLML 704
V+ K+RPS+ V L
Sbjct: 761 VRMRGKDRPSMDKVTTAL 778
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 150 bits (378), Expect = 2e-35
Identities = 87/260 (33%), Positives = 148/260 (56%), Gaps = 3/260 (1%)
Query: 445 GRFWPGIQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLV 504
G+ + G+ +G+++AVK LS+ S QG +EF EV ++ ++ H NL L+G C E +L+
Sbjct: 586 GKVYHGVINGEQVAVKVLSEESAQGYKEFRAEVDLLMRVHHTNLTSLVGYCNEINHMVLI 645
Query: 505 YEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNIL 564
YE+M N++L +L + L W +R I A+G+ YLH + I+HRD+K +NIL
Sbjct: 646 YEYMANENLGDYLAGK-RSFILSWEERLKISLDAAQGLEYLHNGCKPPIVHRDVKPTNIL 704
Query: 565 LDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVL 624
L++++ K++DFGL+R EG + V G+ GY+ PEY +EKSDVYS GV+
Sbjct: 705 LNEKLQAKMADFGLSRSF-SVEGSGQISTVVAGSIGYLDPEYYSTRQMNEKSDVYSLGVV 763
Query: 625 LLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGL 684
LLE+++G+ + + E + + + + ++D+ + + S + I L
Sbjct: 764 LLEVITGQPAIASSKTE-KVHISDHVRSILANGDIRGIVDQRLRERYDVGSAWKMSEIAL 822
Query: 685 LCVQELPKERPSISTVVLML 704
C + +RP++S VV+ L
Sbjct: 823 ACTEHTSAQRPTMSQVVMEL 842
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 149 bits (376), Expect = 3e-35
Identities = 93/273 (34%), Positives = 146/273 (53%), Gaps = 23/273 (8%)
Query: 456 EIAVKRL-SKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSLD 514
++A+KRL + +G+ F E+ + +++HR++VRLLG + +L+YE+MPN SL
Sbjct: 716 DVAIKRLVGRGTGRSDHGFTAEIQTLGRIRHRHIVRLLGYVANKDTNLLLYEYMPNGSLG 775
Query: 515 AFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKIS 574
L + L W R + A+G+ YLH D I+HRD+K++NILLD + ++
Sbjct: 776 ELLHGS-KGGHLQWETRHRVAVEAAKGLCYLHHDCSPLILHRDVKSNNILLDSDFEAHVA 834
Query: 575 DFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRN 634
DFGLA+ + G E + + G+YGY+ PEYA EKSDVYSFGV+LLE+++G++
Sbjct: 835 DFGLAKFLVDGAASECMSS-IAGSYGYIAPEYAYTLKVDEKSDVYSFGVVLLELIAGKKP 893
Query: 635 -NSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFE--------SSMLRCMHIGLL 685
F + D + W EE D + A + +S++ I ++
Sbjct: 894 VGEFGEGVDIV-----RWVRNTEEEITQPSDAAIVVAIVDPRLTGYPLTSVIHVFKIAMM 948
Query: 686 CVQELPKERPSISTVVLMLISEITHLPPPGKVA 718
CV+E RP++ VV ML + PP VA
Sbjct: 949 CVEEEAAARPTMREVVHMLTN------PPKSVA 975
>KPRO_MAIZE (P17801) Putative receptor protein kinase ZmPK1
precursor (EC 2.7.1.37)
Length = 817
Score = 144 bits (363), Expect = 9e-34
Identities = 93/269 (34%), Positives = 140/269 (51%), Gaps = 20/269 (7%)
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
++D + +AVK+L QG E F E+ VI ++ H NLVR+ G C E ++LV E++ N
Sbjct: 553 LEDDRHVAVKKLENVR-QGKEVFQAELSVIGRINHMNLVRIWGFCSEGSHRLLVSEYVEN 611
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
SL LF LDW R NI G+A+G+ YLH + +IH D+K NILLD
Sbjct: 612 GSLANILFSEGGNILLDWEGRFNIALGVAKGLAYLHHECLEWVIHCDVKPENILLDQAFE 671
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKI+DFGL +++ G G N V GT GY+ PE+ + K DVYS+GV+LLE+++
Sbjct: 672 PKITDFGLVKLLNRG-GSTQNVSHVRGTLGYIAPEWVSSLPITAKVDVYSYGVVLLELLT 730
Query: 631 GRRNNSFYQNED-------------SLSLVGFAWKLWLEENTISLIDREVWDASFESSML 677
G R + D S L G + W++ S ++R V +
Sbjct: 731 GTRVSELVGGTDEVHSMLRKLVRMLSAKLEG-EEQSWIDGYLDSKLNRPVNYVQART--- 786
Query: 678 RCMHIGLLCVQELPKERPSISTVVLMLIS 706
+ + + C++E +RP++ V L+S
Sbjct: 787 -LIKLAVSCLEEDRSKRPTMEHAVQTLLS 814
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.358 0.159 0.601
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 74,508,992
Number of Sequences: 164201
Number of extensions: 2751925
Number of successful extensions: 18950
Number of sequences better than 10.0: 1605
Number of HSP's better than 10.0 without gapping: 1128
Number of HSP's successfully gapped in prelim test: 477
Number of HSP's that attempted gapping in prelim test: 15199
Number of HSP's gapped (non-prelim): 2067
length of query: 750
length of database: 59,974,054
effective HSP length: 118
effective length of query: 632
effective length of database: 40,598,336
effective search space: 25658148352
effective search space used: 25658148352
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 70 (31.6 bits)
Medicago: description of AC144721.3


