

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144721.14 - phase: 0
(234 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
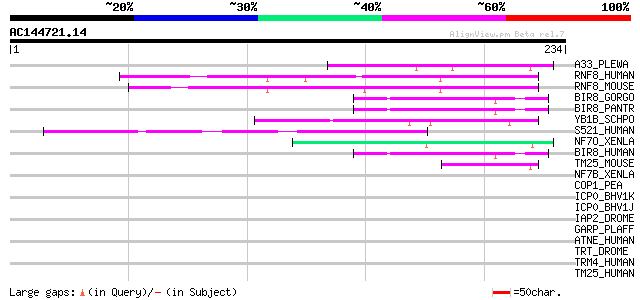
Score E
Sequences producing significant alignments: (bits) Value
A33_PLEWA (Q02084) Zinc-binding protein A33 49 1e-05
RNF8_HUMAN (O76064) Ubiquitin ligase protein RNF8 (EC 6.3.2.-) (... 48 2e-05
RNF8_MOUSE (Q8VC56) Ubiquitin ligase protein RNF8 (EC 6.3.2.-) (... 47 4e-05
BIR8_GORGO (Q95M71) Baculoviral IAP repeat-containing protein 8 ... 47 5e-05
BIR8_PANTR (Q95M72) Baculoviral IAP repeat-containing protein 8 ... 44 3e-04
YB1B_SCHPO (P87176) Hypothetical RING finger protein C3D6.11c in... 44 4e-04
S521_HUMAN (Q96MU7) Putative splicing factor YT521 44 4e-04
NF7O_XENLA (Q91431) Nuclear factor 7, ovary (xnf7-O) 43 6e-04
BIR8_HUMAN (Q96P09) Baculoviral IAP repeat-containing protein 8 ... 43 6e-04
TM25_MOUSE (Q61510) Tripartite motif-containing protein 25 (Zinc... 43 7e-04
NF7B_XENLA (Q92021) Nuclear factor 7, brain (xnf7) (xnf7-B) 42 0.001
COP1_PEA (P93471) Ubiquitin ligase protein COP1 (EC 6.3.2.-) (Co... 42 0.001
ICP0_BHV1K (P29836) Trans-acting transcriptional protein ICP0 (P... 42 0.001
ICP0_BHV1J (P29128) Trans-acting transcriptional protein ICP0 (P... 42 0.001
IAP2_DROME (Q24307) Apoptosis 2 inhibitor (Inhibitor of apoptosi... 42 0.001
GARP_PLAFF (P13816) Glutamic acid-rich protein precursor 42 0.001
ATNE_HUMAN (Q9UN42) X/potassium-transporting ATPase beta-m chain... 42 0.001
TRT_DROME (P19351) Troponin T, skeletal muscle (Upheld protein) ... 42 0.002
TRM4_HUMAN (Q9C037) Tripartite motif-containing protein 4 42 0.002
TM25_HUMAN (Q14258) Tripartite motif-containing protein 25 (Zinc... 42 0.002
>A33_PLEWA (Q02084) Zinc-binding protein A33
Length = 624
Score = 48.9 bits (115), Expect = 1e-05
Identities = 27/108 (25%), Positives = 49/108 (45%), Gaps = 13/108 (12%)
Query: 135 LLEESEIELERISDVVDEGDDVEENNEKREEEEERE--EEGEGEGSNVKVEHN------- 185
+L+ +I+ E +++ DE + V + +E + + E+G+G+ KV+
Sbjct: 100 VLQAKQIKTEELNNTEDETNGVSDQSEGKAARSNKRKIEDGDGDQKKRKVDDEEDDFTED 159
Query: 186 --CCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRG--NCPLCNHFILE 229
C +C K + CGH FC+ C + W S +CP C + E
Sbjct: 160 LTCPLCRSLFKEPVILECGHNFCKHCIDKSWESASAFSCPECKEVLTE 207
>RNF8_HUMAN (O76064) Ubiquitin ligase protein RNF8 (EC 6.3.2.-)
(RING finger protein 8)
Length = 485
Score = 47.8 bits (112), Expect = 2e-05
Identities = 43/189 (22%), Positives = 80/189 (41%), Gaps = 21/189 (11%)
Query: 47 IRENEPNNVVVNVSVTNSSSSSNNNVVKDRKSWKAFKELLLFKRRTSDESTHQQQQNEDI 106
I+ E + V+NV +S V ++ EL + + E QQ + E +
Sbjct: 261 IQMQEKHEAVMNVKKQTQKGNSKKVVQMEQ-------ELQDLQSQLCAEQAQQQARVEQL 313
Query: 107 P-TNSDPSEFLPGGEFSD----------ETAQVSMSLMDLLEESEIELERISDVVDEGDD 155
T + + L G E + + Q +LM+ L S+ + E I + + +
Sbjct: 314 EKTFQEEEQHLQGLEIAQGEKDLKQQLAQALQEHWALMEELNRSKKDFEAI--IQAKNKE 371
Query: 156 VEENNEKREEEEEREEEGEGEGSNV-KVEHNCCVCMVRDKGAAFIPCGHTFCRMCSRELW 214
+E+ E++E+ + ++EE ++V + E C +C A + C H+FC C E
Sbjct: 372 LEQTKEEKEKMQAQKEEVLSHMNDVLENELQCIICSEYFIEAVTLNCAHSFCSYCINEWM 431
Query: 215 VSRGNCPLC 223
+ CP+C
Sbjct: 432 KRKIECPIC 440
>RNF8_MOUSE (Q8VC56) Ubiquitin ligase protein RNF8 (EC 6.3.2.-)
(RING finger protein 8) (AIP37) (ActA-interacting
protein 37) (LaXp180)
Length = 488
Score = 47.0 bits (110), Expect = 4e-05
Identities = 39/183 (21%), Positives = 77/183 (41%), Gaps = 17/183 (9%)
Query: 51 EPNNVVVNVSVTNSSSSSNNNVVKDRKSWKAFKELLLFKRRTSDESTHQQQQNEDIP-TN 109
E V+NV SS +K + KEL + + E QQ + E + T
Sbjct: 268 EKQIAVLNVKRQTRKGSS-------KKIVRMEKELRNLQSQLYAEQAQQQARVEQLEKTF 320
Query: 110 SDPSEFLPGGEFSDETAQVSMSLMDLLEESEIELERISD--------VVDEGDDVEENNE 161
+ + +L G E + L+ L+E + +E ++ + + ++E+ E
Sbjct: 321 QEEAHYLQGLEKEQGECDLKQQLVQALQEHQALMEELNCSKKDFEKIIQAKNKELEQTKE 380
Query: 162 KREEEEEREEEGEGEGSNV-KVEHNCCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRGNC 220
++++ + ++EE +++ + E C +C A + C H+FC C E + C
Sbjct: 381 EKDKVQAQKEEVLSHMNDLLENELQCIICSEYFIEAVTLNCAHSFCSFCINEWMKRKVEC 440
Query: 221 PLC 223
P+C
Sbjct: 441 PIC 443
>BIR8_GORGO (Q95M71) Baculoviral IAP repeat-containing protein 8
(Inhibitor of apoptosis-like protein 2) (IAP-like
protein 2) (ILP-2)
Length = 236
Score = 46.6 bits (109), Expect = 5e-05
Identities = 27/83 (32%), Positives = 39/83 (46%), Gaps = 6/83 (7%)
Query: 146 ISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGH-T 204
++D+V D EN E + +RE E ++ E C +CM R FIPCGH
Sbjct: 150 VADLVSAQKDTTEN-ESNQTSLQREISPEEPLRRLQEEKLCKICMDRHIAVVFIPCGHLV 208
Query: 205 FCRMCSRELWVSRGNCPLCNHFI 227
C+ C+ + CP+CN I
Sbjct: 209 TCKQCAEAV----DRCPMCNAVI 227
>BIR8_PANTR (Q95M72) Baculoviral IAP repeat-containing protein 8
(Inhibitor of apoptosis-like protein 2) (IAP-like
protein 2) (ILP-2)
Length = 236
Score = 44.3 bits (103), Expect = 3e-04
Identities = 26/83 (31%), Positives = 39/83 (46%), Gaps = 6/83 (7%)
Query: 146 ISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGH-T 204
++D+V D EN E + +RE E ++ E C +CM R FIPCGH
Sbjct: 150 VADLVSAQKDTTEN-ESNQTSLQREISPEEPLRRLQDEKLCKICMDRHIAVVFIPCGHLV 208
Query: 205 FCRMCSRELWVSRGNCPLCNHFI 227
C+ C+ + CP+C+ I
Sbjct: 209 TCKQCAEAV----DRCPMCSAVI 227
>YB1B_SCHPO (P87176) Hypothetical RING finger protein C3D6.11c in
chromosome II
Length = 269
Score = 43.5 bits (101), Expect = 4e-04
Identities = 31/137 (22%), Positives = 58/137 (41%), Gaps = 18/137 (13%)
Query: 104 EDIPTNSDPSEFLPGGEFSDETAQVSMSLMDLLEESEIELERISDVVDEGDDVEENNEKR 163
EDI +D + +++ A V SL ++ + ISD++D D+ + K+
Sbjct: 111 EDINFQNDADDINQRFTYNNHPASVENSLTNV-NSIHAQPTTISDMIDLTDETSYDPRKQ 169
Query: 164 EEEE-----------EREEEGEGE---GSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMC 209
+ E+ E+EE + + S ++ C +C+ + + PCGH FC C
Sbjct: 170 KFEQGKNPSTTNAEIEKEEPSKKQVVPSSQRLADYKCVICLDSPENLSCTPCGHIFCNFC 229
Query: 210 ---SRELWVSRGNCPLC 223
+ + CP+C
Sbjct: 230 ILSALGTTAATQKCPVC 246
>S521_HUMAN (Q96MU7) Putative splicing factor YT521
Length = 727
Score = 43.5 bits (101), Expect = 4e-04
Identities = 39/162 (24%), Positives = 67/162 (41%), Gaps = 19/162 (11%)
Query: 15 STSNTRSSLASLIQNDAVLSNQKNTNQTLLDIIRENEPNNVVVNVSVTNSSSSSNNNVVK 74
S N R I+ + S + NQ +R+ +P S +
Sbjct: 103 SERNKRLDADRKIRLSSSASREPYKNQPEKTCVRKRDPER---RAKSPTPDGSERIGLEV 159
Query: 75 DRKSWKAFKELLLFKRRTSDESTHQQQQNEDIPTNSDPSEFLPGGEFSDETAQVSMSLMD 134
DR++ ++ ++S E + ++ D T S S SDE + + +
Sbjct: 160 DRRASRS--------SQSSKEEVNSEEYGSDHETGSSGS--------SDEQGNNTENEEE 203
Query: 135 LLEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGE 176
+EE E E + + +E ++V+E+ E+ EEEEE EEE E E
Sbjct: 204 GVEEDVEEDEEVEEDAEEDEEVDEDGEEEEEEEEEEEEEEEE 245
>NF7O_XENLA (Q91431) Nuclear factor 7, ovary (xnf7-O)
Length = 610
Score = 43.1 bits (100), Expect = 6e-04
Identities = 27/125 (21%), Positives = 50/125 (39%), Gaps = 15/125 (12%)
Query: 120 EFSDETAQVSMSLMDLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGE----- 174
E+ D++ V +E + + E +++ ++ D K EE E ++ +
Sbjct: 67 EWVDKSRLVLTKPPKEVETNGTDQEEMTEPTEQPDSKTPQKRKLEEPEPEPKKAKVEDKD 126
Query: 175 --------GEGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRGN--CPLCN 224
G + E C +C+ K + CGH FCR C ++W + + CP C
Sbjct: 127 ASKTAASLGAAGDFAEELTCPLCVELFKDPVMVACGHNFCRSCIDKVWEGQSSFACPECK 186
Query: 225 HFILE 229
I +
Sbjct: 187 ESITD 191
>BIR8_HUMAN (Q96P09) Baculoviral IAP repeat-containing protein 8
(Inhibitor of apoptosis-like protein 2) (IAP-like
protein 2) (ILP-2) (Testis-specific inhibitor of
apoptosis)
Length = 236
Score = 43.1 bits (100), Expect = 6e-04
Identities = 26/83 (31%), Positives = 39/83 (46%), Gaps = 6/83 (7%)
Query: 146 ISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGH-T 204
++D+V D EN E + +RE E ++ E C +CM R FIPCGH
Sbjct: 150 VADLVSAQKDTTEN-ELNQTSLQREISPEEPLRRLQEEKLCKICMDRYIAVVFIPCGHLV 208
Query: 205 FCRMCSRELWVSRGNCPLCNHFI 227
C+ C+ + CP+C+ I
Sbjct: 209 TCKQCAEAV----DRCPMCSAVI 227
>TM25_MOUSE (Q61510) Tripartite motif-containing protein 25 (Zinc
finger protein 147) (Estrogen responsive finger protein)
(Efp)
Length = 634
Score = 42.7 bits (99), Expect = 7e-04
Identities = 19/44 (43%), Positives = 22/44 (49%), Gaps = 3/44 (6%)
Query: 183 EHNCCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRG---NCPLC 223
E +C VC+ K PCGH FC C E WV +G CP C
Sbjct: 10 ELSCSVCLELFKEPVTTPCGHNFCTSCLDETWVVQGPPYRCPQC 53
>NF7B_XENLA (Q92021) Nuclear factor 7, brain (xnf7) (xnf7-B)
Length = 609
Score = 42.4 bits (98), Expect = 0.001
Identities = 19/82 (23%), Positives = 35/82 (42%), Gaps = 2/82 (2%)
Query: 150 VDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMC 209
++E + + + E++ + G + E C +C+ K + CGH FCR C
Sbjct: 109 IEEPEPEPKKAKVEEKDASKNASSLGAAGDFAEELTCPLCVELFKDPVMVACGHNFCRSC 168
Query: 210 SRELWVSRGN--CPLCNHFILE 229
+ W + + CP C I +
Sbjct: 169 IDKAWEGQSSFACPECRESITD 190
>COP1_PEA (P93471) Ubiquitin ligase protein COP1 (EC 6.3.2.-)
(Constitutive photomorphogenesis protein 1)
Length = 672
Score = 42.4 bits (98), Expect = 0.001
Identities = 17/50 (34%), Positives = 27/50 (54%)
Query: 178 SNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRGNCPLCNHFI 227
S++ + C +CM K A CGH+FC MC ++ +CP C H++
Sbjct: 39 SDLDKDFLCPICMQIIKDAFLTACGHSFCYMCIITHLRNKSDCPCCGHYL 88
>ICP0_BHV1K (P29836) Trans-acting transcriptional protein ICP0 (P135
protein) (IER 2.9/ER2.6)
Length = 676
Score = 42.0 bits (97), Expect = 0.001
Identities = 18/40 (45%), Positives = 22/40 (55%), Gaps = 1/40 (2%)
Query: 185 NCCVCMVRDKGAA-FIPCGHTFCRMCSRELWVSRGNCPLC 223
+CC+C+ GAA +PC H FC C R R CPLC
Sbjct: 12 SCCICLDAITGAARALPCLHAFCLACIRRWLEGRPTCPLC 51
>ICP0_BHV1J (P29128) Trans-acting transcriptional protein ICP0 (P135
protein) (IER 2.9/ER2.6)
Length = 676
Score = 42.0 bits (97), Expect = 0.001
Identities = 18/40 (45%), Positives = 22/40 (55%), Gaps = 1/40 (2%)
Query: 185 NCCVCMVRDKGAA-FIPCGHTFCRMCSRELWVSRGNCPLC 223
+CC+C+ GAA +PC H FC C R R CPLC
Sbjct: 12 SCCICLDAITGAARALPCLHAFCLACIRRWLEGRPTCPLC 51
>IAP2_DROME (Q24307) Apoptosis 2 inhibitor (Inhibitor of apoptosis
2) (dIAP2) (DIAP) (IAP homolog A) (IAP-like protein)
(DILP)
Length = 498
Score = 42.0 bits (97), Expect = 0.001
Identities = 25/87 (28%), Positives = 38/87 (42%), Gaps = 14/87 (16%)
Query: 138 ESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAA 197
E+ + +I+D + + N EEE R+ +K C VC+ + G
Sbjct: 412 EAVANISKITDEIQKMSVATPNGNLSLEEENRQ---------LKDARLCKVCLDEEVGVV 462
Query: 198 FIPCGH-TFCRMCSRELWVSRGNCPLC 223
F+PCGH C C+ S NCP+C
Sbjct: 463 FLPCGHLATCNQCA----PSVANCPMC 485
>GARP_PLAFF (P13816) Glutamic acid-rich protein precursor
Length = 678
Score = 42.0 bits (97), Expect = 0.001
Identities = 27/128 (21%), Positives = 55/128 (42%)
Query: 58 NVSVTNSSSSSNNNVVKDRKSWKAFKELLLFKRRTSDESTHQQQQNEDIPTNSDPSEFLP 117
+V + V ++ K + +E + +E ++++ E+ + E
Sbjct: 544 HVDKEEDKKEESKEVQEESKEVQEDEEEVEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEE 603
Query: 118 GGEFSDETAQVSMSLMDLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGEG 177
+ +E + D EE E + E D D+ ++ ++ +E +EE+E EEE E E
Sbjct: 604 EEDEDEEDEDDAEEDEDDAEEDEDDAEEDDDEEDDDEEDDDEDEDEDEEDEEEEEEEEEE 663
Query: 178 SNVKVEHN 185
S K++ N
Sbjct: 664 SEKKIKRN 671
Score = 35.0 bits (79), Expect = 0.15
Identities = 23/101 (22%), Positives = 48/101 (46%), Gaps = 12/101 (11%)
Query: 76 RKSWKAFKELLLFKRRTSDESTHQQQQNEDIPTNSDPSEFLPGGEFSDETAQVSMSLMDL 135
+K +E L K++ D+ ++++++++ E S E + + +
Sbjct: 528 KKKMAKIEEAELQKQKHVDKEEDKKEESKEVQ------------EESKEVQEDEEEVEED 575
Query: 136 LEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGE 176
EE E E E + +E ++ EE E+ EE+E+ E+E + E
Sbjct: 576 EEEEEEEEEEEEEEEEEEEEEEEEEEEEEEDEDEEDEDDAE 616
Score = 30.8 bits (68), Expect = 2.9
Identities = 19/89 (21%), Positives = 39/89 (43%), Gaps = 7/89 (7%)
Query: 94 DESTHQQQQNEDIPTNSDPSEFLPGGEFSDETAQVSMSLMDLLEESEIELERISDVVDEG 153
+E +++++ED D E E ++ A+ ++ E + E D ++
Sbjct: 597 EEEEEEEEEDEDEEDEDDAEEDEDDAEEDEDDAEED-------DDEEDDDEEDDDEDEDE 649
Query: 154 DDVEENNEKREEEEEREEEGEGEGSNVKV 182
D+ +E E+ EEEE ++ N K+
Sbjct: 650 DEEDEEEEEEEEEESEKKIKRNLRKNAKI 678
>ATNE_HUMAN (Q9UN42) X/potassium-transporting ATPase beta-m chain
(X,K-ATPase beta-m subunit)
Length = 357
Score = 42.0 bits (97), Expect = 0.001
Identities = 21/41 (51%), Positives = 26/41 (63%)
Query: 137 EESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGEG 177
EE E E E VV + ++ EE EK EEEEE +EE EG+G
Sbjct: 35 EEEEAEEEARVTVVPKSEEEEEEEEKEEEEEEEKEEEEGQG 75
>TRT_DROME (P19351) Troponin T, skeletal muscle (Upheld protein)
(Intended thorax protein)
Length = 396
Score = 41.6 bits (96), Expect = 0.002
Identities = 23/54 (42%), Positives = 31/54 (56%)
Query: 123 DETAQVSMSLMDLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGE 176
+E A+ +++ EE E E+ D DE D+ EE E+ EEEEE EEE E E
Sbjct: 335 EEDAKADEDIVEDDEEVEEEVVEEEDEEDEEDEEEEEEEEEEEEEEEEEEEEEE 388
Score = 38.9 bits (89), Expect = 0.011
Identities = 20/41 (48%), Positives = 25/41 (60%)
Query: 136 LEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGE 176
+EE +E E D DE ++ EE E+ EEEEE EEE E E
Sbjct: 351 VEEEVVEEEDEEDEEDEEEEEEEEEEEEEEEEEEEEEEEEE 391
Score = 36.6 bits (83), Expect = 0.052
Identities = 18/43 (41%), Positives = 27/43 (61%)
Query: 134 DLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGE 176
+++EE + E E + +E ++ EE E+ EEEEE EEE E E
Sbjct: 354 EVVEEEDEEDEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 396
Score = 33.1 bits (74), Expect = 0.58
Identities = 16/50 (32%), Positives = 29/50 (58%)
Query: 123 DETAQVSMSLMDLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEE 172
++ +V +++ +E + E E + +E ++ EE E+ EEEEE EEE
Sbjct: 346 EDDEEVEEEVVEEEDEEDEEDEEEEEEEEEEEEEEEEEEEEEEEEEEEEE 395
Score = 32.3 bits (72), Expect = 0.99
Identities = 14/39 (35%), Positives = 25/39 (63%)
Query: 138 ESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGE 176
E E + + D+V++ ++VEE + E+EE+ E+E E E
Sbjct: 333 EGEEDAKADEDIVEDDEEVEEEVVEEEDEEDEEDEEEEE 371
>TRM4_HUMAN (Q9C037) Tripartite motif-containing protein 4
Length = 500
Score = 41.6 bits (96), Expect = 0.002
Identities = 18/53 (33%), Positives = 24/53 (44%), Gaps = 3/53 (5%)
Query: 176 EGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRGN---CPLCNH 225
E +++ E C +C+ + I CGH FCR C W G CP C H
Sbjct: 2 EAEDIQEELTCPICLDYFQDPVSIECGHNFCRGCLHRNWAPGGGPFPCPECRH 54
>TM25_HUMAN (Q14258) Tripartite motif-containing protein 25 (Zinc
finger protein 147) (Estrogen responsive finger protein)
(Efp)
Length = 630
Score = 41.6 bits (96), Expect = 0.002
Identities = 17/44 (38%), Positives = 22/44 (49%), Gaps = 3/44 (6%)
Query: 183 EHNCCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRGN---CPLC 223
E +C +C+ K PCGH FC C E W +G+ CP C
Sbjct: 10 ELSCSICLEPFKEPVTTPCGHNFCGSCLNETWAVQGSPYLCPQC 53
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.311 0.128 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,611,835
Number of Sequences: 164201
Number of extensions: 1220622
Number of successful extensions: 15416
Number of sequences better than 10.0: 757
Number of HSP's better than 10.0 without gapping: 407
Number of HSP's successfully gapped in prelim test: 357
Number of HSP's that attempted gapping in prelim test: 9938
Number of HSP's gapped (non-prelim): 2602
length of query: 234
length of database: 59,974,054
effective HSP length: 107
effective length of query: 127
effective length of database: 42,404,547
effective search space: 5385377469
effective search space used: 5385377469
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC144721.14


