

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144617.6 - phase: 0
(178 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
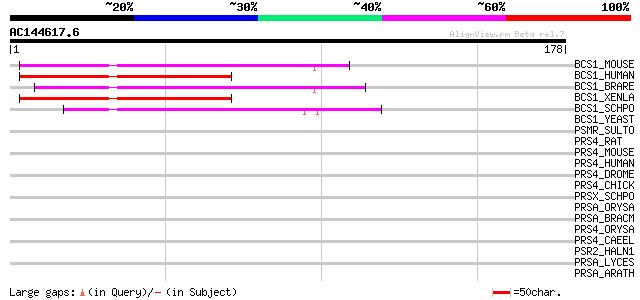
Score E
Sequences producing significant alignments: (bits) Value
BCS1_MOUSE (Q9CZP5) Mitochondrial chaperone BCS1 (BCS1-like prot... 55 7e-08
BCS1_HUMAN (Q9Y276) Mitochondrial chaperone BCS1 (BCS1-like prot... 54 2e-07
BCS1_BRARE (Q7ZV60) Mitochondrial chaperone BCS1 (BCS1-like prot... 52 9e-07
BCS1_XENLA (Q7ZTL7) Mitochondrial chaperone BCS1 (BCS1-like prot... 51 2e-06
BCS1_SCHPO (Q9P6Q3) Probable mitochondrial chaperone BCS1 (BCS1-... 47 2e-05
BCS1_YEAST (P32839) Mitochondrial chaperone BCS1 42 0.001
PSMR_SULTO (Q975U2) Proteasome-activating nucleotidase (Proteaso... 38 0.011
PRS4_RAT (P62193) 26S protease regulatory subunit 4 (P26s4) (Pro... 37 0.018
PRS4_MOUSE (P62192) 26S protease regulatory subunit 4 (P26s4) (P... 37 0.018
PRS4_HUMAN (P62191) 26S protease regulatory subunit 4 (P26s4) (P... 37 0.018
PRS4_DROME (P48601) 26S protease regulatory subunit 4 (P26s4) 37 0.018
PRS4_CHICK (Q90732) 26S protease regulatory subunit 4 (P26s4) (P... 37 0.018
PRSX_SCHPO (O74445) Probable 26S protease subunit rpt4 37 0.024
PRSA_ORYSA (P46465) 26S protease regulatory subunit 6A homolog (... 37 0.024
PRSA_BRACM (O23894) 26S protease regulatory subunit 6A homolog (... 37 0.024
PRS4_ORYSA (P46466) 26S protease regulatory subunit 4 homolog (T... 37 0.024
PRS4_CAEEL (O16368) Probable 26S protease regulatory subunit 4 37 0.024
PSR2_HALN1 (Q9HRW6) Proteasome-activating nucleotidase 2 (Protea... 36 0.054
PRSA_LYCES (P54776) 26S protease regulatory subunit 6A homolog (... 36 0.054
PRSA_ARATH (O04019) 26S protease regulatory subunit 6A homolog (... 36 0.054
>BCS1_MOUSE (Q9CZP5) Mitochondrial chaperone BCS1 (BCS1-like
protein)
Length = 418
Score = 55.5 bits (132), Expect = 7e-08
Identities = 34/108 (31%), Positives = 57/108 (52%), Gaps = 4/108 (3%)
Query: 4 KESQAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMD 63
++ EN K + ++T GLLN +DG+ AST R++ TTNY ++LD ALI GR+D
Sbjct: 291 RDLAVENPIKYQGLGRLTFSGLLNALDGV--ASTEARIVFMTTNYIDRLDPALIRPGRVD 348
Query: 64 MLIELPYCCFDGFKMLATKYLSLESHFLFDKIA--CLLVETNMTPADV 109
+ + YC + ++ ++ L + A L + ++PA V
Sbjct: 349 LKEYVGYCSHWQLTQMFQRFYPGQAPSLAENFAEHVLKATSEISPAQV 396
>BCS1_HUMAN (Q9Y276) Mitochondrial chaperone BCS1 (BCS1-like
protein) (H-BCS1)
Length = 419
Score = 53.9 bits (128), Expect = 2e-07
Identities = 27/68 (39%), Positives = 42/68 (61%), Gaps = 2/68 (2%)
Query: 4 KESQAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMD 63
++ EN K + ++T GLLN +DG+ AST R++ TTN+ ++LD ALI GR+D
Sbjct: 291 RDLAVENPVKYQGLGRLTFSGLLNALDGV--ASTEARIVFMTTNHVDRLDPALIRPGRVD 348
Query: 64 MLIELPYC 71
+ + YC
Sbjct: 349 LKEYVGYC 356
>BCS1_BRARE (Q7ZV60) Mitochondrial chaperone BCS1 (BCS1-like
protein)
Length = 420
Score = 51.6 bits (122), Expect = 9e-07
Identities = 33/108 (30%), Positives = 55/108 (50%), Gaps = 4/108 (3%)
Query: 9 ENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIEL 68
EN M ++T GLLN +DG+ AS+ R++ TTN+ E+LD AL+ GR+D+ +
Sbjct: 297 ENPLAYQGMGRLTFSGLLNALDGV--ASSEARIVFMTTNFIERLDPALVRPGRVDLKQYV 354
Query: 69 PYCCFDGFKMLATKYLSLESHFLFDKIA--CLLVETNMTPADVAENLM 114
+C + ++ ES D + L T+++ A V + M
Sbjct: 355 GHCSHWQLTQMFRRFYPQESAAEADHFSEQALAAHTDLSAAQVQGHFM 402
>BCS1_XENLA (Q7ZTL7) Mitochondrial chaperone BCS1 (BCS1-like
protein)
Length = 419
Score = 50.8 bits (120), Expect = 2e-06
Identities = 26/68 (38%), Positives = 42/68 (61%), Gaps = 2/68 (2%)
Query: 4 KESQAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMD 63
++ +N T M ++T GLLN +DG+ AST R++ TTN+ ++LD ALI GR+D
Sbjct: 291 RDLNKQNPTAYQGMGRLTFSGLLNALDGV--ASTEARIVFMTTNHIDRLDPALIRPGRVD 348
Query: 64 MLIELPYC 71
+ + +C
Sbjct: 349 VKQYVGHC 356
>BCS1_SCHPO (Q9P6Q3) Probable mitochondrial chaperone BCS1
(BCS1-like protein)
Length = 449
Score = 47.4 bits (111), Expect = 2e-05
Identities = 35/115 (30%), Positives = 52/115 (44%), Gaps = 15/115 (13%)
Query: 18 SQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELPYCCFDGFK 77
+ +T GLLN +DG+ S+ ER+I TTN+ EKLD AL+ GR+D+ L + +
Sbjct: 320 ANVTFSGLLNALDGV--TSSDERIIFMTTNHPEKLDPALVRPGRVDVKAYLGNATPEQVR 377
Query: 78 MLATKYLSLESHFLFD--KIAC-----------LLVETNMTPADVAENLMPKVDN 119
+ T++ D I C L V +PAD + DN
Sbjct: 378 EMFTRFYGHSPEMADDLSDIVCPKNTSMASLQGLFVMNKSSPADAVDMAKELPDN 432
>BCS1_YEAST (P32839) Mitochondrial chaperone BCS1
Length = 456
Score = 41.6 bits (96), Expect = 0.001
Identities = 21/46 (45%), Positives = 30/46 (64%), Gaps = 2/46 (4%)
Query: 18 SQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMD 63
S +T GLLN +DG+ S+ E + TTN+ EKLD A++ GR+D
Sbjct: 338 SSVTFSGLLNALDGVTSSE--ETITFMTTNHPEKLDAAIMRPGRID 381
>PSMR_SULTO (Q975U2) Proteasome-activating nucleotidase (Proteasome
regulatory subunit)
Length = 392
Score = 38.1 bits (87), Expect = 0.011
Identities = 29/78 (37%), Positives = 38/78 (48%), Gaps = 2/78 (2%)
Query: 7 QAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLI 66
+ + T + Q TL LL IDG + II TN + LD AL+ GR D LI
Sbjct: 243 RVDMGTSGEREIQRTLMQLLAEIDGFKPLDNVK--IIAATNRLDILDPALLRPGRFDRLI 300
Query: 67 ELPYCCFDGFKMLATKYL 84
E+P F+G K + YL
Sbjct: 301 EVPLPNFEGRKEIFRIYL 318
>PRS4_RAT (P62193) 26S protease regulatory subunit 4 (P26s4)
(Proteasome 26S subunit ATPase 1)
Length = 440
Score = 37.4 bits (85), Expect = 0.018
Identities = 24/51 (47%), Positives = 29/51 (56%), Gaps = 2/51 (3%)
Query: 19 QITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELP 69
Q T+ LLN +DG S G+ +I TN E LD ALI GR+D IE P
Sbjct: 306 QRTMLELLNQLDGF--DSRGDVKVIMATNRIETLDPALIRPGRIDRKIEFP 354
>PRS4_MOUSE (P62192) 26S protease regulatory subunit 4 (P26s4)
(Proteasome 26S subunit ATPase 1)
Length = 440
Score = 37.4 bits (85), Expect = 0.018
Identities = 24/51 (47%), Positives = 29/51 (56%), Gaps = 2/51 (3%)
Query: 19 QITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELP 69
Q T+ LLN +DG S G+ +I TN E LD ALI GR+D IE P
Sbjct: 306 QRTMLELLNQLDGF--DSRGDVKVIMATNRIETLDPALIRPGRIDRKIEFP 354
>PRS4_HUMAN (P62191) 26S protease regulatory subunit 4 (P26s4)
(Proteasome 26S subunit ATPase 1)
Length = 440
Score = 37.4 bits (85), Expect = 0.018
Identities = 24/51 (47%), Positives = 29/51 (56%), Gaps = 2/51 (3%)
Query: 19 QITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELP 69
Q T+ LLN +DG S G+ +I TN E LD ALI GR+D IE P
Sbjct: 306 QRTMLELLNQLDGF--DSRGDVKVIMATNRIETLDPALIRPGRIDRKIEFP 354
>PRS4_DROME (P48601) 26S protease regulatory subunit 4 (P26s4)
Length = 439
Score = 37.4 bits (85), Expect = 0.018
Identities = 24/51 (47%), Positives = 29/51 (56%), Gaps = 2/51 (3%)
Query: 19 QITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELP 69
Q T+ LLN +DG S G+ +I TN E LD ALI GR+D IE P
Sbjct: 305 QRTMLELLNQLDGF--DSRGDVKVIMATNRIETLDPALIRPGRIDRKIEFP 353
>PRS4_CHICK (Q90732) 26S protease regulatory subunit 4 (P26s4)
(Proteasome 26S subunit ATPase 1)
Length = 440
Score = 37.4 bits (85), Expect = 0.018
Identities = 24/51 (47%), Positives = 29/51 (56%), Gaps = 2/51 (3%)
Query: 19 QITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELP 69
Q T+ LLN +DG S G+ +I TN E LD ALI GR+D IE P
Sbjct: 306 QRTMLELLNQLDGF--DSRGDVKVIMATNRIETLDPALIRPGRIDRKIEFP 354
>PRSX_SCHPO (O74445) Probable 26S protease subunit rpt4
Length = 388
Score = 37.0 bits (84), Expect = 0.024
Identities = 24/58 (41%), Positives = 32/58 (54%), Gaps = 2/58 (3%)
Query: 12 TKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELP 69
T ++ Q TL LLN +DG G+ II TN + LD AL+ GR+D IE+P
Sbjct: 246 TSADREIQRTLMELLNQMDGF--DYLGQTKIIMATNRPDTLDPALLRPGRLDRKIEIP 301
>PRSA_ORYSA (P46465) 26S protease regulatory subunit 6A homolog
(TAT-binding protein homolog 1) (TBP-1)
Length = 429
Score = 37.0 bits (84), Expect = 0.024
Identities = 24/63 (38%), Positives = 37/63 (58%), Gaps = 4/63 (6%)
Query: 9 ENATKNNKMSQITLPGLLNFIDGIWSASTGERL-IIFTTNYAEKLDHALICRGRMDMLIE 67
++ ++ Q T+ LLN +DG S+ ER+ +I TN A+ LD AL+ GR+D IE
Sbjct: 287 DSEVSGDREVQRTMLELLNQLDGF---SSDERIKVIAATNRADILDPALMRSGRLDRKIE 343
Query: 68 LPY 70
P+
Sbjct: 344 FPH 346
>PRSA_BRACM (O23894) 26S protease regulatory subunit 6A homolog
(TAT-binding protein homolog 1) (TBP-1)
Length = 424
Score = 37.0 bits (84), Expect = 0.024
Identities = 24/63 (38%), Positives = 37/63 (58%), Gaps = 4/63 (6%)
Query: 9 ENATKNNKMSQITLPGLLNFIDGIWSASTGERL-IIFTTNYAEKLDHALICRGRMDMLIE 67
++ ++ Q T+ LLN +DG S+ ER+ +I TN A+ LD AL+ GR+D IE
Sbjct: 282 DSEVSGDREVQRTMLELLNQLDGF---SSDERIKVIAATNRADILDPALMRSGRLDRKIE 338
Query: 68 LPY 70
P+
Sbjct: 339 FPH 341
>PRS4_ORYSA (P46466) 26S protease regulatory subunit 4 homolog
(TAT-binding protein homolog 2)
Length = 448
Score = 37.0 bits (84), Expect = 0.024
Identities = 23/51 (45%), Positives = 29/51 (56%), Gaps = 2/51 (3%)
Query: 19 QITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELP 69
Q T+ LLN +DG S G+ +I TN E LD AL+ GR+D IE P
Sbjct: 314 QRTMLELLNQLDGF--DSRGDVKVILATNRIESLDPALLRPGRIDRKIEFP 362
>PRS4_CAEEL (O16368) Probable 26S protease regulatory subunit 4
Length = 443
Score = 37.0 bits (84), Expect = 0.024
Identities = 23/51 (45%), Positives = 29/51 (56%), Gaps = 2/51 (3%)
Query: 19 QITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLIELP 69
Q T+ LLN +DG S G+ ++ TN E LD ALI GR+D IE P
Sbjct: 309 QRTMLELLNQLDGF--DSRGDVKVLMATNRIESLDPALIRPGRIDRKIEFP 357
>PSR2_HALN1 (Q9HRW6) Proteasome-activating nucleotidase 2
(Proteasome regulatory subunit 2)
Length = 407
Score = 35.8 bits (81), Expect = 0.054
Identities = 22/64 (34%), Positives = 35/64 (54%), Gaps = 2/64 (3%)
Query: 7 QAENATKNNKMSQITLPGLLNFIDGIWSASTGERLIIFTTNYAEKLDHALICRGRMDMLI 66
+ ++ T + Q T+ LL+ +DG G+ II TN + LD A++ GR D LI
Sbjct: 258 RTDSKTSGDAEVQRTMMQLLSEMDGF--DERGDIRIIAATNRFDMLDRAILRPGRFDRLI 315
Query: 67 ELPY 70
E+P+
Sbjct: 316 EVPH 319
>PRSA_LYCES (P54776) 26S protease regulatory subunit 6A homolog
(TAT-binding protein homolog 1) (TBP-1)
(Mg(2+)-dependent ATPase 1) (LEMA-1)
Length = 423
Score = 35.8 bits (81), Expect = 0.054
Identities = 23/63 (36%), Positives = 37/63 (58%), Gaps = 4/63 (6%)
Query: 9 ENATKNNKMSQITLPGLLNFIDGIWSASTGERL-IIFTTNYAEKLDHALICRGRMDMLIE 67
++ ++ Q T+ LLN +DG S+ +R+ +I TN A+ LD AL+ GR+D IE
Sbjct: 281 DSEVSGDREVQRTMLELLNQLDGF---SSDDRIKVIAATNRADILDPALMRSGRLDRKIE 337
Query: 68 LPY 70
P+
Sbjct: 338 FPH 340
>PRSA_ARATH (O04019) 26S protease regulatory subunit 6A homolog
(TAT-binding protein homolog 1) (TBP-1)
Length = 419
Score = 35.8 bits (81), Expect = 0.054
Identities = 23/63 (36%), Positives = 37/63 (58%), Gaps = 4/63 (6%)
Query: 9 ENATKNNKMSQITLPGLLNFIDGIWSASTGERL-IIFTTNYAEKLDHALICRGRMDMLIE 67
++ ++ Q T+ LLN +DG S+ +R+ +I TN A+ LD AL+ GR+D IE
Sbjct: 277 DSEVSGDREVQRTMLELLNQLDGF---SSDDRIKVIAATNRADILDPALMRSGRLDRKIE 333
Query: 68 LPY 70
P+
Sbjct: 334 FPH 336
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.130 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,193,286
Number of Sequences: 164201
Number of extensions: 752249
Number of successful extensions: 4981
Number of sequences better than 10.0: 192
Number of HSP's better than 10.0 without gapping: 93
Number of HSP's successfully gapped in prelim test: 101
Number of HSP's that attempted gapping in prelim test: 4438
Number of HSP's gapped (non-prelim): 512
length of query: 178
length of database: 59,974,054
effective HSP length: 103
effective length of query: 75
effective length of database: 43,061,351
effective search space: 3229601325
effective search space used: 3229601325
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC144617.6


