

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144592.1 - phase: 0 /pseudo
(486 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
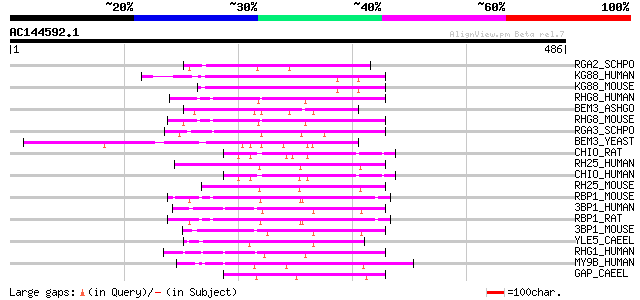
Score E
Sequences producing significant alignments: (bits) Value
RGA2_SCHPO (Q10164) Probable Rho-type GTPase-activating protein 2 77 1e-13
KG88_HUMAN (Q9C0H5) Protein KIAA1688 66 2e-10
KG88_MOUSE (P59281) Protein KIAA1688 homolog (Fragment) 65 3e-10
RHG8_HUMAN (Q9NSG0) Rho-GTPase-activating protein 8 (PP610) 62 3e-09
BEM3_ASHGO (Q74ZH7) GTPase-activating protein BEM3 61 6e-09
RHG8_MOUSE (Q9CXP4) Rho-GTPase-activating protein 8 60 2e-08
RGA3_SCHPO (O14014) Probable Rho-type GTPase-activating protein 3 59 4e-08
BEM3_YEAST (P32873) GTPase-activating protein BEM3 59 4e-08
CHIO_RAT (Q03070) Beta-chimaerin (Beta-chimerin) (Rho-GTPase-act... 57 1e-07
RH25_HUMAN (P42331) Rho-GTPase-activating protein 25 56 2e-07
CHIO_HUMAN (P52757) Beta-chimaerin (Beta-chimerin) (Rho-GTPase-a... 56 2e-07
RH25_MOUSE (Q8BYW1) Rho-GTPase-activating protein 25 54 9e-07
RBP1_MOUSE (Q62172) RalA binding protein 1 (RalBP1) (Ral interac... 54 1e-06
3BP1_HUMAN (Q9Y3L3) SH3-domain binding protein 1 (3BP-1) 54 1e-06
RBP1_RAT (Q62796) RalA binding protein 1 (RalBP1) (Ral interacti... 52 3e-06
3BP1_MOUSE (P55194) SH3-domain binding protein 1 (3BP-1) 52 3e-06
YLE5_CAEEL (P46941) Hypothetical protein C38D4.5 in chromosome III 52 4e-06
RHG1_HUMAN (Q07960) Rho-GTPase-activating protein 1 (GTPase-acti... 51 8e-06
MY9B_HUMAN (Q13459) Myosin IXb (Unconventional myosin-9b) 51 8e-06
GAP_CAEEL (P34288) GTPase-activating protein GAP (CeGAP) 50 1e-05
>RGA2_SCHPO (Q10164) Probable Rho-type GTPase-activating protein 2
Length = 1275
Score = 77.0 bits (188), Expect = 1e-13
Identities = 53/180 (29%), Positives = 90/180 (49%), Gaps = 18/180 (10%)
Query: 153 VFGV---SAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRD 209
+FG+ A ++ ++D G +P ++ R + L S K EGI+R++ S +++
Sbjct: 1062 IFGLPLNEAVNISTQFNDSG--LPIVVYRCIEYLESCRAEKEEGIYRLSGSASTIKHLKE 1119
Query: 210 QLNKGV------VPHGIDVHCLSGLIKAWFRELPTGVLDS-------LTPEQVMQCNTED 256
Q N+GV DVH ++GL+K + R LPT +LD+ L P
Sbjct: 1120 QFNEGVDYDLLSSDEEFDVHVIAGLLKLYLRNLPTNLLDTSMHKLFELLPNVPNDSAALG 1179
Query: 257 DCTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADPLTALI 316
+ +++ LP ALLD ++ + ++ E+ NKMN RNV +VF+P + +D LI
Sbjct: 1180 ELCDVISKLPPENFALLDSLLHHLRRIIAFEKVNKMNIRNVCIVFSPTLNIPSDIFMMLI 1239
>KG88_HUMAN (Q9C0H5) Protein KIAA1688
Length = 1083
Score = 65.9 bits (159), Expect = 2e-10
Identities = 57/220 (25%), Positives = 101/220 (45%), Gaps = 28/220 (12%)
Query: 116 EVRHVSHVTFDRFNGFLGLPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTI 175
E+RH + F P+ SA +V G+ + Y +R +P +
Sbjct: 874 EIRHAKNAVFS----------------PSMFGSALQEVMGMQRER----YPER--QLPWV 911
Query: 176 LLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNKGVVPHGI-DVHCLSGLIKAWFR 234
R+ +++ + G + EGIFR+ + + ++ Q+++ VP G+ D H + L+K W+R
Sbjct: 912 QTRLSEEVLALNGDQTEGIFRVPGDIDEVNALKLQVDQWKVPTGLEDPHVPASLLKLWYR 971
Query: 235 ELPTGVLDSLTPEQ-VMQCNTEDDCTNLVKLLPSTEAALLDWAINLMADVVE--NEQFNK 291
EL ++ EQ + ++ + +V LP +L + I + V+ N K
Sbjct: 972 ELEEPLIPHEFYEQCIAHYDSPEAAVAVVHALPRINRMVLCYLIRFLQVFVQPANVAVTK 1031
Query: 292 MNARNVAMVFAPN--MTQMADPLTALIHAVQVMNFLKTLI 329
M+ N+AMV APN Q DP + + M+FL+ LI
Sbjct: 1032 MDVSNLAMVMAPNCLRCQSDDPRVIFENTRKEMSFLRVLI 1071
>KG88_MOUSE (P59281) Protein KIAA1688 homolog (Fragment)
Length = 1097
Score = 65.5 bits (158), Expect = 3e-10
Identities = 49/171 (28%), Positives = 85/171 (49%), Gaps = 8/171 (4%)
Query: 165 YDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNKGVVPHGI-DVH 223
Y DR +P + R+ +++ + G + EGIFR+ + + ++ Q+++ VP G+ D H
Sbjct: 917 YPDR--QLPWVQTRLSEEVLALNGDQTEGIFRVPGDIDEVNALKLQVDQWKVPTGLEDPH 974
Query: 224 CLSGLIKAWFRELPTGVLDSLTPEQ-VMQCNTEDDCTNLVKLLPSTEAALLDWAINLMAD 282
+ L+K W+REL ++ EQ + + + +V LP +L + I +
Sbjct: 975 VPASLLKLWYRELEEPLIPHEFYEQCIAHYESPEAAVAVVHALPRINRMVLCYLIRFLQV 1034
Query: 283 VVE--NEQFNKMNARNVAMVFAPN--MTQMADPLTALIHAVQVMNFLKTLI 329
V+ N KM+ N+AMV APN Q DP + + M+FL+ LI
Sbjct: 1035 FVQPANVAITKMDVSNLAMVMAPNCLRCQSDDPRVIFENTRKEMSFLRVLI 1085
>RHG8_HUMAN (Q9NSG0) Rho-GTPase-activating protein 8 (PP610)
Length = 718
Score = 62.0 bits (149), Expect = 3e-09
Identities = 51/199 (25%), Positives = 92/199 (45%), Gaps = 14/199 (7%)
Query: 141 EVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAE 200
+ P P + FGVS + ++ ++G +P +L R E GL+ EG+FR +A
Sbjct: 465 KTPPPRPPLPTQQFGVSLQYLKDK--NQGELIPPVL-RFTVTYLREKGLRTEGLFRRSAS 521
Query: 201 NSQEAFVRDQLNKGVV----PHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCNTED 256
++ N+G +G D+H + ++K + RELP +L EQ++ +
Sbjct: 522 VQTVREIQRLYNQGKPVNFDDYG-DIHIPAVILKTFLRELPQPLLTFQAYEQILGITCVE 580
Query: 257 D------CTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMAD 310
C +++ LP +L + + + V FNKMN+ N+A VF N+ +
Sbjct: 581 SSLRVTGCRQILRSLPEHNYVVLRYLMGFLHAVSRESIFNKMNSSNLACVFGLNLIWPSQ 640
Query: 311 PLTALIHAVQVMNFLKTLI 329
+++L V + F + LI
Sbjct: 641 GVSSLSALVPLNMFTELLI 659
>BEM3_ASHGO (Q74ZH7) GTPase-activating protein BEM3
Length = 1013
Score = 61.2 bits (147), Expect = 6e-09
Identities = 48/189 (25%), Positives = 91/189 (47%), Gaps = 38/189 (20%)
Query: 153 VFGVSAKS--MQCSYDDRGN-SVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRD 209
VFG +S S+ +G +P+++ R + LY G++ EGIFR++ +S +++
Sbjct: 790 VFGADLRSCLQLSSHPYQGKYEIPSVVFRTLEFLYKNRGIQEEGIFRLSGSSSLIKSLQE 849
Query: 210 QLNK-------------GVVPHG-------IDVHCLSGLIKAWFRELPTGVLDS---LTP 246
Q +K V P +DV+ +SGL+K + R+LP + +
Sbjct: 850 QFDKEYDVDLCNYNDKVSVTPGNENQGGLYVDVNTVSGLLKLYLRKLPHMIFGDAAYMDF 909
Query: 247 EQVMQCNTEDDCTNLVKL----------LPSTEAALLDWAINLMADVVENEQFNKMNARN 296
+++++ N +D + L+ L + AL+ L+ + EN ++NKMN RN
Sbjct: 910 KRIVERNGDD--SKLIALEFRALVNSGRIAKEYVALMYALFELLVKITENSKYNKMNLRN 967
Query: 297 VAMVFAPNM 305
+ +VF+P +
Sbjct: 968 LCIVFSPTL 976
>RHG8_MOUSE (Q9CXP4) Rho-GTPase-activating protein 8
Length = 425
Score = 59.7 bits (143), Expect = 2e-08
Identities = 54/205 (26%), Positives = 96/205 (46%), Gaps = 18/205 (8%)
Query: 139 QPEVPTRVPSAS----VKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGI 194
QP PT+ P + FGVS + ++ ++G +P +L R E GL+ EG+
Sbjct: 174 QPPPPTKTPPPRPPLPTQQFGVSLQYLRDK--NQGELIPPVL-RWTVTYLREKGLRTEGL 230
Query: 195 FRITAENSQEAFVRDQLNKGVV----PHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVM 250
FR +A V+ ++G +G D+H + ++K + RELP +L EQ++
Sbjct: 231 FRRSASAQTVRQVQRLYDQGKPVNFDDYG-DMHLPAVILKTFLRELPQPLLTFQAYEQIL 289
Query: 251 QCNTEDD------CTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPN 304
+ + C +++ LP A+L + + + +V NKMN+ N+A VF N
Sbjct: 290 GITSVESSLRVTHCRLILRSLPEHNYAVLRYLMGFLHEVSLESISNKMNSSNLACVFGLN 349
Query: 305 MTQMADPLTALIHAVQVMNFLKTLI 329
+ + + +L V + F + LI
Sbjct: 350 LIWPSQGVASLSALVPLNLFTELLI 374
>RGA3_SCHPO (O14014) Probable Rho-type GTPase-activating protein 3
Length = 969
Score = 58.5 bits (140), Expect = 4e-08
Identities = 50/206 (24%), Positives = 98/206 (47%), Gaps = 16/206 (7%)
Query: 136 SEFQPEVPTRVP--SASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEG 193
S++ PE+PTR+P S +FG +S++ G+ +P ++ M GL+ EG
Sbjct: 758 SKWMPEMPTRMPPPGPSPTMFG---RSLENQLKIEGSVLPQVIA-MCVSCVDAHGLEVEG 813
Query: 194 IFRITAENSQEAFVRDQLNKGVVPHG---IDVHCLSGLIKAWFRELPTGVLDSLTPEQVM 250
I+RI+ SQ + D+ G + D+ + ++K + LP V+ E+++
Sbjct: 814 IYRISGSASQVRVLVDEFENGSIRMEHLTSDLFACTSVLKTYLHRLPEPVIPGTQYEELL 873
Query: 251 QCNT----EDDCTNLVKLLPSTEAALLD---WAINLMADVVENEQFNKMNARNVAMVFAP 303
+ E+ +V+++ + A L + I + V ++ + N MN++NV+ VFAP
Sbjct: 874 EAEKIEKEEEKIERVVEVMKTLHPAHLSVFRFLIAHLGRVCKHAEKNLMNSKNVSTVFAP 933
Query: 304 NMTQMADPLTALIHAVQVMNFLKTLI 329
+ + L HA + L+ ++
Sbjct: 934 TLMRDKVNRFDLQHATKKSTALQFML 959
>BEM3_YEAST (P32873) GTPase-activating protein BEM3
Length = 1128
Score = 58.5 bits (140), Expect = 4e-08
Identities = 74/337 (21%), Positives = 142/337 (41%), Gaps = 56/337 (16%)
Query: 13 AGFSVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTK 72
AG S+F A + +E + + + N+ + D I S+ ++ F
Sbjct: 765 AGTHHSKFGNATISATDTPSYVTDLTQEYNNNNNISNSSNNIANSDGIDSNPSSHSNFLA 824
Query: 73 GSNNNHQNQ-------------FAILDIVMAALKKSIVTCSVEREDVSSLDISWPTEVRH 119
S+ N + + F L +A+ +T S + + +S D + +
Sbjct: 825 SSSGNAEEEKDSRRAKMRSLFPFKKLTGPASAMNHIGITISNDSDSPTSPDSIIKSPSKK 884
Query: 120 VSHVTFDRFNGFLGLPSEFQPEVPTRVPSASVKV-FGVSAKSMQCSYDDRGNSVPTILLR 178
+ V+ S P V T + +S++ +S+ Q YD +P+++ R
Sbjct: 885 LMEVSSSS-------NSSTGPHVSTAIFGSSLETCLRLSSHKYQNVYD-----LPSVVYR 932
Query: 179 MQKQLYSEGGLKAEGIFRITAENS-----QEAFVRD------QLNKGVVPHG-------- 219
+ LY G++ EGIFR++ ++ QE F ++ + N+ +
Sbjct: 933 CLEYLYKNRGIQEEGIFRLSGSSTVIKTLQERFDKEYDVDLCRYNESIEAKDDEASPSLY 992
Query: 220 IDVHCLSGLIKAWFRELPT---GVLDSLTPEQVMQCNTEDDCT------NLVK--LLPST 268
I V+ +SGL+K + R+LP G L+ ++V+ N + L++ L+P
Sbjct: 993 IGVNTVSGLLKLYLRKLPHLLFGDEQFLSFKRVVDENHNNPVQISLGFKELIESGLVPHA 1052
Query: 269 EAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNM 305
+L+ L+ + EN +FNKMN RN+ +VF+P +
Sbjct: 1053 NLSLMYALFELLVRINENSKFNKMNLRNLCIVFSPTL 1089
>CHIO_RAT (Q03070) Beta-chimaerin (Beta-chimerin)
(Rho-GTPase-activating protein 3)
Length = 295
Score = 56.6 bits (135), Expect = 1e-07
Identities = 48/167 (28%), Positives = 86/167 (50%), Gaps = 22/167 (13%)
Query: 188 GLKAEGIFRITA-----ENSQEAFVRD----QLNKGVVPHGIDVHCLSGLIKAWFRELPT 238
GLK+EG++R++ E+ + AF RD ++ + P D++ ++G +K +FR+LP
Sbjct: 132 GLKSEGLYRVSGFTEHIEDVKMAFDRDGEKADISANIYP---DINIITGALKLYFRDLPI 188
Query: 239 GVL--DSLTP--EQVMQCNTEDDCT---NLVKLLPSTEAALLDWAINLMADVVENEQFNK 291
++ D+ T E N ++ ++ LLP L + + + V NE+ N
Sbjct: 189 PIITYDTYTKFIEAAKISNADERLEAVHEVLMLLPPAHYETLRYLMIHLKKVTMNEKDNL 248
Query: 292 MNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKMLREREE 338
MNA N+ +VF P T M P + + + M + K LI+++L E E+
Sbjct: 249 MNAENLGIVFGP--TLMRPPEDSTLTTLHDMRYQK-LIVQILIENED 292
>RH25_HUMAN (P42331) Rho-GTPase-activating protein 25
Length = 638
Score = 56.2 bits (134), Expect = 2e-07
Identities = 46/199 (23%), Positives = 81/199 (40%), Gaps = 14/199 (7%)
Query: 145 RVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQE 204
RV VFG G + IL+ + E G EGIFR+ +++
Sbjct: 141 RVAGTPCGVFGQRLDETVAYEQKFGPHLVPILVEKCAEFILEHGRNEEGIFRLPGQDNLV 200
Query: 205 AFVRDQLNKGVVP---HGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCN-------- 253
+RD + G P DVH ++ L+K + R+LP V+ E + C
Sbjct: 201 KQLRDAFDAGERPSFDRDTDVHTVASLLKLYLRDLPEPVVPWSQYEGFLLCGQLTNADEA 260
Query: 254 -TEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNM--TQMAD 310
+ + + +LP +LL + + ++ N NKM+ N+A V N+ +++ D
Sbjct: 261 KAQQELMKQLSILPRDNYSLLSYICRFLHEIQLNCAVNKMSVDNLATVIGVNLIRSKVED 320
Query: 311 PLTALIHAVQVMNFLKTLI 329
P + Q+ + +I
Sbjct: 321 PAVIMRGTPQIQRVMTMMI 339
>CHIO_HUMAN (P52757) Beta-chimaerin (Beta-chimerin)
(Rho-GTPase-activating protein 3)
Length = 468
Score = 56.2 bits (134), Expect = 2e-07
Identities = 48/167 (28%), Positives = 85/167 (50%), Gaps = 22/167 (13%)
Query: 188 GLKAEGIFRITA-----ENSQEAFVRD----QLNKGVVPHGIDVHCLSGLIKAWFRELPT 238
GLK+EG++R++ E+ + AF RD ++ V P D++ ++G +K +FR+LP
Sbjct: 305 GLKSEGLYRVSGFTEHIEDVKMAFDRDGEKADISANVYP---DINIITGALKLYFRDLPI 361
Query: 239 GVLDSLTPEQVMQC----NTEDDCT---NLVKLLPSTEAALLDWAINLMADVVENEQFNK 291
V+ T + + N ++ ++ LLP L + + + V NE+ N
Sbjct: 362 PVITYDTYSKFIDAAKISNADERLEAVHEVLMLLPPAHYETLRYLMIHLKKVTMNEKDNF 421
Query: 292 MNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTLILKMLREREE 338
MNA N+ +VF P T M P + + + M + K LI+++L E E+
Sbjct: 422 MNAENLGIVFGP--TLMRPPEDSTLTTLHDMRYQK-LIVQILIENED 465
>RH25_MOUSE (Q8BYW1) Rho-GTPase-activating protein 25
Length = 648
Score = 53.9 bits (128), Expect = 9e-07
Identities = 41/175 (23%), Positives = 75/175 (42%), Gaps = 14/175 (8%)
Query: 169 GNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQLNKGVVP---HGIDVHCL 225
G + IL+ + E G+ EGIFR+ +++ +RD + G P DVH +
Sbjct: 173 GPHLVPILVEKCAEFILEHGVSEEGIFRLPGQDNLVKQLRDAFDAGERPSFDRDTDVHTV 232
Query: 226 SGLIKAWFRELPTGVLDSLTPEQVMQC---------NTEDDCTNLVKLLPSTEAALLDWA 276
+ L+K + R+LP V+ E + C + + + LP LL +
Sbjct: 233 ASLLKLYLRDLPEPVVPWSQYEGFLLCGQLMNADEAKAQQELVKQLSTLPRDNYNLLSYI 292
Query: 277 INLMADVVENEQFNKMNARNVAMVFAPNM--TQMADPLTALIHAVQVMNFLKTLI 329
+ ++ N NKM+ N+A V N+ +++ DP + Q+ + +I
Sbjct: 293 CRFLHEIQLNCAVNKMSVDNLATVIGVNLIRSKVEDPAVIMRGTPQIQRVMTMMI 347
>RBP1_MOUSE (Q62172) RalA binding protein 1 (RalBP1) (Ral
interacting protein 1) (Dinitrophenyl S-glutathione
ATPase) (DNP-SG ATPase)
Length = 647
Score = 53.5 bits (127), Expect = 1e-06
Identities = 56/208 (26%), Positives = 97/208 (45%), Gaps = 19/208 (9%)
Query: 139 QPEVPTRVPSASVK-VFGV---SAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGI 194
+PEVP ++ + SVK +FGV A YD G +P + R + G+K EG+
Sbjct: 174 EPEVP-QMDAPSVKPIFGVPLVDAVERTMMYD--GVRLPAVF-RECVDYMEKHGMKCEGV 229
Query: 195 FRITAENSQEAFVRDQLNKGVVPH--GIDVHCLSGLIKAWFRELPTGVLDS-LTPEQVMQ 251
+R++ S+ ++ ++ P+ + + ++ L+K + R+LP +L L P
Sbjct: 230 YRVSGIKSKVDELKAAYDREESPNLEEYEPNTVASLLKQYLRDLPENLLTKELMPRFEEA 289
Query: 252 CN--TE----DDCTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNM 305
C TE + L++ LP LL W I + V+ E KMN +N+++V +P +
Sbjct: 290 CGKTTEMEKVQEFQRLLRELPECNHLLLSWLIVHLDHVIAKELETKMNIQNISIVLSPTV 349
Query: 306 TQMADPLTALIHAVQVMNFLKTLILKML 333
L L VQ T++LK +
Sbjct: 350 QISNRVLYVLFTHVQ--ELFGTVVLKQV 375
>3BP1_HUMAN (Q9Y3L3) SH3-domain binding protein 1 (3BP-1)
Length = 701
Score = 53.5 bits (127), Expect = 1e-06
Identities = 49/206 (23%), Positives = 92/206 (43%), Gaps = 25/206 (12%)
Query: 143 PTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENS 202
P+ + +V+GVS + + G + + L SEG +K EG+FR+ A S
Sbjct: 263 PSMTATHFPRVYGVS---LATHLQELGREIALPIEACVMMLLSEG-MKEEGLFRLAAGAS 318
Query: 203 QEAFVRDQLNKGVVPHGI-----DVHCLSGLIKAWFRELPTGVLD-SLTPEQVMQCNTED 256
++ + PH + D H ++G +K++ RELP ++ L + + + ++
Sbjct: 319 VLKRLKQTMASD--PHSLEEFCSDPHAVAGALKSYLRELPEPLMTFDLYDDWMRAASLKE 376
Query: 257 DCTNLVKL------LPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMT---- 306
L L LP + L + + +A + E ++ NKM N+A+V PN+
Sbjct: 377 PGARLQALQEVCSRLPPENLSNLRYLMKFLARLAEEQEVNKMTPSNIAIVLGPNLLWPPE 436
Query: 307 ---QMADPLTALIHAVQVMNFLKTLI 329
A A + ++QV+ ++ LI
Sbjct: 437 KEGDQAQLDAASVSSIQVVGVVEALI 462
>RBP1_RAT (Q62796) RalA binding protein 1 (RalBP1) (Ral interacting
protein 1) (Cytocentrin) (Dinitrophenyl S-glutathione
ATPase) (DNP-SG ATPase)
Length = 646
Score = 52.0 bits (123), Expect = 3e-06
Identities = 54/207 (26%), Positives = 91/207 (43%), Gaps = 17/207 (8%)
Query: 139 QPEVPTRVPSASVKVFGV---SAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIF 195
+PEVP + +FG A YD G +P + R + G+K EGI+
Sbjct: 174 EPEVPQTDAPSLRPIFGAPFADAVERTMMYD--GIRLPAVF-RECVDYMEKHGMKCEGIY 230
Query: 196 RITAENSQEAFVRDQLNKGVVPH--GIDVHCLSGLIKAWFRELPTGVLDS-LTPEQVMQC 252
R++ S+ ++ ++ P+ + + ++ L+K + R+LP +L L P C
Sbjct: 231 RVSGIKSKVDELKAAYDREESPNLEEYEPNTVASLLKQYLRDLPENLLTKELMPRFEEAC 290
Query: 253 N--TE----DDCTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMT 306
TE + L++ LP LL W I M V+ E KMN +N+++V +P +
Sbjct: 291 GRTTEVEKVQEFQRLLRELPEYNHLLLSWLIVHMDHVIAKELETKMNIQNISIVLSPTVQ 350
Query: 307 QMADPLTALIHAVQVMNFLKTLILKML 333
L L VQ T++LK +
Sbjct: 351 ISNRVLYVLFTHVQ--ELFGTVLLKQV 375
>3BP1_MOUSE (P55194) SH3-domain binding protein 1 (3BP-1)
Length = 601
Score = 52.0 bits (123), Expect = 3e-06
Identities = 48/197 (24%), Positives = 88/197 (44%), Gaps = 24/197 (12%)
Query: 152 KVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVRDQL 211
+V+GVS ++ D G + + L SEG + EG+FR+ A S ++ +
Sbjct: 192 RVYGVSLRT---HLQDLGRDIALPIEACVLLLLSEGMQEEEGLFRLAAGASVLKRLKQTM 248
Query: 212 NKGVVPHGIDVHC-----LSGLIKAWFRELPTGVLDS-LTPEQVMQCNTEDDCTNLVKL- 264
PH ++ C ++G +K++ RELP ++ S L + + + ++ L L
Sbjct: 249 ASD--PHSLEEFCSGPHAVAGALKSYLRELPEPLMTSDLYDDWMRAASLKEPGARLEALH 306
Query: 265 -----LPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMT-------QMADPL 312
LP L + + +A + E + NKM N+A+V PN+ A
Sbjct: 307 DVCSRLPQENFNNLRYLMKFLALLAEEQDVNKMTPSNIAIVLGPNLLWPPEKEGDQAQLD 366
Query: 313 TALIHAVQVMNFLKTLI 329
A + ++QV+ ++ LI
Sbjct: 367 AASVSSIQVVGVVEALI 383
>YLE5_CAEEL (P46941) Hypothetical protein C38D4.5 in chromosome III
Length = 837
Score = 51.6 bits (122), Expect = 4e-06
Identities = 45/168 (26%), Positives = 73/168 (42%), Gaps = 14/168 (8%)
Query: 153 VFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVR---D 209
VFG S S C ++ NS+ +R+ ++ GL+ +GI+R++ S +R D
Sbjct: 607 VFG-STLSAICQHE---NSLVPKFIRVITEVIESKGLETDGIYRVSGNLSAVQKIRCQAD 662
Query: 210 QLNKGVVPHGIDVHCLSGLIKAWFRELPTGVLD-SLTPEQVMQCNTEDDCTNLVKL---- 264
Q N + D+H L+G +K +FREL + SL E + T K
Sbjct: 663 QDNYKALVSEEDIHVLTGALKLFFRELTDPLFPISLHKEYTSAMQMPNATTRFKKFEELL 722
Query: 265 --LPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMAD 310
LP+ L + + V + N+M N+A+VF P + D
Sbjct: 723 SRLPNENRETLKMLLRHLNRVASHSSQNRMQQHNLAIVFGPTLFHNGD 770
>RHG1_HUMAN (Q07960) Rho-GTPase-activating protein 1
(GTPase-activating protein rhoOGAP) (Rho-related small
GTPase protein activator) (CDC42 GTPase-activating
protein) (p50-rhoGAP)
Length = 439
Score = 50.8 bits (120), Expect = 8e-06
Identities = 52/205 (25%), Positives = 88/205 (42%), Gaps = 14/205 (6%)
Query: 135 PSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGI 194
P+ +P R P + + FGVS + +Q ++ +P +L L + L EGI
Sbjct: 224 PATAPKPMPPRPPLPNQQ-FGVSLQHLQEKNPEQ-EPIPIVLRETVAYLQAHA-LTTEGI 280
Query: 195 FRITAENSQEAFVRDQLNKGVVPHGID----VHCLSGLIKAWFRELPTGVLD-SLTPEQV 249
FR +A V+ + N G+ P D +H + ++K + RELP +L L P V
Sbjct: 281 FRRSANTQVVREVQQKYNMGL-PVDFDQYNELHLPAVILKTFLRELPEPLLTFDLYPHVV 339
Query: 250 MQCNTEDD-----CTNLVKLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPN 304
N ++ +++ LP +L + + + + NKM N+A+VF PN
Sbjct: 340 GFLNIDESQRVPATLQVLQTLPEENYQVLRFLTAFLVQISAHSDQNKMTNTNLAVVFGPN 399
Query: 305 MTQMADPLTALIHAVQVMNFLKTLI 329
+ D L + F K L+
Sbjct: 400 LLWAKDAAITLKAINPINTFTKFLL 424
>MY9B_HUMAN (Q13459) Myosin IXb (Unconventional myosin-9b)
Length = 2158
Score = 50.8 bits (120), Expect = 8e-06
Identities = 51/220 (23%), Positives = 100/220 (45%), Gaps = 18/220 (8%)
Query: 147 PSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAF 206
P A FGV S+ D+ SVP +L ++ + + G L EG++R + ++
Sbjct: 1691 PGAEPGHFGVCVDSLT---SDKA-SVPIVLEKLLEHVEMHG-LYTEGLYRKSGAANRTRE 1745
Query: 207 VRDQLNK---GVVPHGIDVHCLSGLIKAWFRELPTGVL------DSLTPEQVMQCNTEDD 257
+R L V +H ++G++K W RELP ++ D L ++ + +
Sbjct: 1746 LRQALQTDPAAVKLENFPIHAITGVLKQWLRELPEPLMTFAQYGDFLRAVELPEKQEQLA 1805
Query: 258 CTNLV-KLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQM---ADPLT 313
V + LP L+ I + V E N+M+ +A++FAP + + +DPLT
Sbjct: 1806 AIYAVLEHLPEANHNSLERLIFHLVKVALLEDVNRMSPGALAIIFAPCLLRCPDNSDPLT 1865
Query: 314 ALIHAVQVMNFLKTLILKMLREREESIDNARLLSHSMDFA 353
++ +++ ++ LI + +R+ + ++ L + A
Sbjct: 1866 SMKDVLKITTCVEMLIKEQMRKYKVKMEEISQLEAAESIA 1905
>GAP_CAEEL (P34288) GTPase-activating protein GAP (CeGAP)
Length = 1317
Score = 50.1 bits (118), Expect = 1e-05
Identities = 42/163 (25%), Positives = 76/163 (45%), Gaps = 21/163 (12%)
Query: 188 GLKAEGIFRITAENSQEAFVRDQL-NKG-----------VVPHGIDVHCLSGLIKAWFRE 235
G+ GI+RI + +++ L N+G + P DV+ +S L+K + R+
Sbjct: 689 GMDTVGIYRIPGNTAAVNALKESLSNRGFDSVDLSKVESLDPRWRDVNVVSSLLKMFLRK 748
Query: 236 LPTGVL-DSLTPEQV------MQCNTEDDCTNLVKLLPSTEAALLDWAINLMADVVENEQ 288
LP +L D L P + N NL++ LP L + I ++++ ++
Sbjct: 749 LPEPLLTDKLYPFFIDANRISTHHNRLHKLRNLLRKLPRPHYDTLRFLIVHLSEITKHSD 808
Query: 289 FNKMNARNVAMVFAPNMTQMADP--LTALIHAVQVMNFLKTLI 329
NKM RN+A++F P++ + +D T + H ++TLI
Sbjct: 809 VNKMECRNLALMFGPSIVRPSDDNMATMVTHMSDQCKIIETLI 851
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.314 0.129 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 56,874,808
Number of Sequences: 164201
Number of extensions: 2489419
Number of successful extensions: 32554
Number of sequences better than 10.0: 584
Number of HSP's better than 10.0 without gapping: 439
Number of HSP's successfully gapped in prelim test: 149
Number of HSP's that attempted gapping in prelim test: 21555
Number of HSP's gapped (non-prelim): 4331
length of query: 486
length of database: 59,974,054
effective HSP length: 114
effective length of query: 372
effective length of database: 41,255,140
effective search space: 15346912080
effective search space used: 15346912080
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 68 (30.8 bits)
Medicago: description of AC144592.1


