

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144591.1 - phase: 0
(167 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
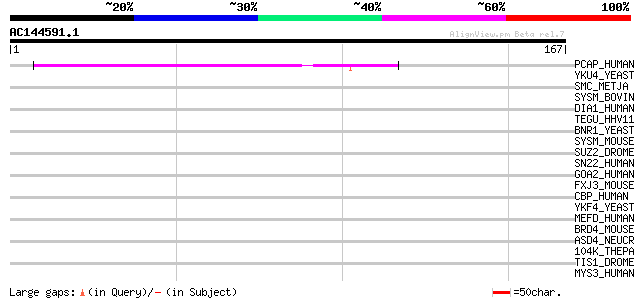
Score E
Sequences producing significant alignments: (bits) Value
PCAP_HUMAN (Q96RN5) Positive cofactor 2 glutamine/Q-rich-associa... 43 4e-04
YKU4_YEAST (P36041) Hypothetical 69.8 kDa protein in LOS1-TOR2 i... 37 0.021
SMC_METJA (Q59037) Chromosome partition protein smc homolog 37 0.027
SYSM_BOVIN (Q9N0F3) Seryl-tRNA synthetase, mitochondrial precurs... 35 0.061
DIA1_HUMAN (O60610) Diaphanous protein homolog 1 (Diaphanous-rel... 35 0.080
TEGU_HHV11 (P10220) Large tegument protein (Virion protein UL36) 34 0.18
BNR1_YEAST (P40450) BNI1 related protein 1 33 0.23
SYSM_MOUSE (Q9JJL8) Seryl-tRNA synthetase, mitochondrial precurs... 33 0.30
SUZ2_DROME (P25172) Suppressor 2 of zeste protein (Protein poste... 33 0.30
SN22_HUMAN (P51531) Possible global transcription activator SNF2... 33 0.30
GOA2_HUMAN (Q08379) Golgi autoantigen, golgin subfamily A member... 33 0.30
FXJ3_MOUSE (Q8BUR3) Forkhead box protein J3 33 0.30
CBP_HUMAN (Q92793) CREB-binding protein (EC 2.3.1.48) 33 0.30
YKF4_YEAST (P35732) Hypothetical 84.0 kDa protein in NUP120-CSE4... 33 0.40
MEFD_HUMAN (Q14814) Myocyte-specific enhancer factor 2D 33 0.40
BRD4_MOUSE (Q9ESU6) Bromodomain-containing protein 4 (Mitotic ch... 33 0.40
ASD4_NEUCR (Q9HEV5) GATA type zinc finger protein Asd4 (Ascus de... 33 0.40
104K_THEPA (P15711) 104 kDa microneme-rhoptry antigen 33 0.40
TIS1_DROME (P47980) TIS11 protein (dTIS11) 32 0.52
MYS3_HUMAN (Q92794) MYST histone acetyltransferase 3 (Runt-relat... 32 0.52
>PCAP_HUMAN (Q96RN5) Positive cofactor 2 glutamine/Q-rich-associated
protein (PC2 glutamine/Q-rich-associated protein)
(TPA-inducible gene-1) (TIG-1) (Activator-recruited
cofactor 105 kDa component) (ARC105) (CTG repeat protein
7a)
Length = 788
Score = 42.7 bits (99), Expect = 4e-04
Identities = 30/111 (27%), Positives = 53/111 (47%), Gaps = 4/111 (3%)
Query: 8 AHMQTLAQTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQPPR 67
A +Q Q + Q +Q L +L + + ++ R++ L + + Q QQ+ Q +
Sbjct: 202 AVVQQQQQLQQQQQQQQHLIKLHHQNQQQIQQQQQQLQRIAQLQLQQQQQQQQQQQQQQQ 261
Query: 68 QPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKG-NYVQTRPPPPRPEKLP 117
Q Q P QQ P + P P ++ LP L+ ++ Q PPP+P++ P
Sbjct: 262 QALQAQPPIQQPPMQQPQPPP---SQALPQQLQQMHHTQHHQPPPQPQQPP 309
Score = 35.0 bits (79), Expect = 0.080
Identities = 28/112 (25%), Positives = 54/112 (48%), Gaps = 17/112 (15%)
Query: 10 MQTLAQTHKMDQLEQELHELRKEMTALRAK--VEELTNRVSSLMVTKDLPQFQQR----- 62
+Q +AQ Q +Q+ + +++ AL+A+ +++ + ++ LPQ Q+
Sbjct: 238 LQRIAQLQLQQQQQQQQQQQQQQQQALQAQPPIQQPPMQQPQPPPSQALPQQLQQMHHTQ 297
Query: 63 -----PQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPP 109
PQP + P QP Q P+++ P+ + A+ LP G + T+PP
Sbjct: 298 HHQPPPQPQQPPVAQNQPSQLPPQSQTQPL-VSQAQALP----GQMLYTQPP 344
>YKU4_YEAST (P36041) Hypothetical 69.8 kDa protein in LOS1-TOR2
intergenic region
Length = 632
Score = 37.0 bits (84), Expect = 0.021
Identities = 23/58 (39%), Positives = 27/58 (45%), Gaps = 12/58 (20%)
Query: 56 LPQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRP 113
LPQFQQ+PQ + +FTQ P P A F L KG + PPPP P
Sbjct: 412 LPQFQQQPQQMQPMAFTQHP------------PNNNAFFNGLLNKGKSETSTPPPPPP 457
Score = 28.5 bits (62), Expect = 7.4
Identities = 19/59 (32%), Positives = 23/59 (38%), Gaps = 2/59 (3%)
Query: 53 TKDLPQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPP 111
T DLPQ Q P PP F P P P+P + +L Q +PP P
Sbjct: 526 TPDLPQQQYMPPPPPPGFFPMHP--NFPNGPMPPLPQGFPIPPNGMLPVTGQQPQPPYP 582
>SMC_METJA (Q59037) Chromosome partition protein smc homolog
Length = 1169
Score = 36.6 bits (83), Expect = 0.027
Identities = 16/39 (41%), Positives = 25/39 (64%)
Query: 16 THKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTK 54
T K +QLE+E+ L +E + KV ++ NR++ LMV K
Sbjct: 891 TEKKEQLEKEIETLERERREILRKVRDIENRINELMVEK 929
>SYSM_BOVIN (Q9N0F3) Seryl-tRNA synthetase, mitochondrial precursor
(EC 6.1.1.11) (Serine--tRNA ligase) (SerRSmt)
Length = 518
Score = 35.4 bits (80), Expect = 0.061
Identities = 17/41 (41%), Positives = 27/41 (65%)
Query: 24 QELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQ 64
QEL +LR+++ +L + E +T V +L+V +D Q QQ PQ
Sbjct: 94 QELRQLREQIRSLEEEKEAVTEAVRALVVNQDNSQVQQDPQ 134
>DIA1_HUMAN (O60610) Diaphanous protein homolog 1
(Diaphanous-related formin 1) (DRF1)
Length = 1248
Score = 35.0 bits (79), Expect = 0.080
Identities = 32/116 (27%), Positives = 42/116 (35%), Gaps = 12/116 (10%)
Query: 16 THKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQR-----PQPPRQPS 70
T ++ +L +EL + +KEM +L A + V S P P PP
Sbjct: 536 TGEVAKLTKELEDAKKEMASLSAAAITVPPSVPSRAPVPPAPPLPGDSGTIIPPPPAPGD 595
Query: 71 FTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPP-------PRPEKLPAG 119
T P P P+P A P L G+ PPP P P LP G
Sbjct: 596 STTPPPPPPPPPPPPPLPGGTAISPPPPLSGDATIPPPPPLPEGVGIPSPSSLPGG 651
Score = 34.3 bits (77), Expect = 0.14
Identities = 30/106 (28%), Positives = 42/106 (39%), Gaps = 8/106 (7%)
Query: 20 DQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQPPRQPSFTQQPR--Q 77
D L E ++ E L A+V +LT V+ L TK+L ++ + T P
Sbjct: 512 DALHSEKQQIATEKQDLEAEVSQLTGEVAKL--TKELEDAKKEMASLSAAAITVPPSVPS 569
Query: 78 QAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRP----EKLPAG 119
+AP P+P +P T PPPP P LP G
Sbjct: 570 RAPVPPAPPLPGDSGTIIPPPPAPGDSTTPPPPPPPPPPPPPLPGG 615
>TEGU_HHV11 (P10220) Large tegument protein (Virion protein UL36)
Length = 3164
Score = 33.9 bits (76), Expect = 0.18
Identities = 24/72 (33%), Positives = 33/72 (45%), Gaps = 2/72 (2%)
Query: 47 VSSLMVTKDLPQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQT 106
VS L + PQ Q +PQP QP QP+ Q P+ + P P + P Q
Sbjct: 2905 VSRLSAPQPQPQPQPQPQPQPQPQPQPQPQPQ-PQPQPQPQPQPQPQPQPQPQPQPQPQP 2963
Query: 107 RP-PPPRPEKLP 117
+P P P+P+ P
Sbjct: 2964 QPQPQPQPQPQP 2975
>BNR1_YEAST (P40450) BNI1 related protein 1
Length = 1375
Score = 33.5 bits (75), Expect = 0.23
Identities = 28/103 (27%), Positives = 35/103 (33%), Gaps = 8/103 (7%)
Query: 37 RAKVEELTNRVSSLMVTKDLPQFQQRPQPPRQPSFTQQ--PRQQAPRTRFDPIPMKYAEF 94
R + L N + + LPQ P PP P Q +A I
Sbjct: 746 RLSSDSLDNGIQLVPEVVKLPQLPPPPPPPPPPPLPQSLLTEAEAKPDGVSCIAAPAPPP 805
Query: 95 LPSLLKGNYVQTRPPPPRPEKLPAGFRADLSCVFHQGAPGHDV 137
LP L K PPPP P LP S ++G HD+
Sbjct: 806 LPDLFKTKTCGAVPPPPPPPPLPE------SLSMNKGPSNHDL 842
>SYSM_MOUSE (Q9JJL8) Seryl-tRNA synthetase, mitochondrial precursor
(EC 6.1.1.11) (Serine--tRNA ligase) (SerRSmt)
Length = 518
Score = 33.1 bits (74), Expect = 0.30
Identities = 15/41 (36%), Positives = 26/41 (62%)
Query: 24 QELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQ 64
QEL +LR+++ +L A+ E + V +L+ +D Q Q+ PQ
Sbjct: 94 QELRQLREQIRSLEAEKEAVAEAVRALLANQDSDQVQKDPQ 134
>SUZ2_DROME (P25172) Suppressor 2 of zeste protein (Protein
posterior sex combs)
Length = 1365
Score = 33.1 bits (74), Expect = 0.30
Identities = 32/124 (25%), Positives = 50/124 (39%), Gaps = 21/124 (16%)
Query: 9 HMQTLAQTHKMDQLEQELHELRKEMT----ALRAKVEELTNRVSSLMV----------TK 54
H ++H + E +L +L+ + T AL + E ++SL+V T
Sbjct: 540 HKLDFKRSHSLASGELDLQKLKLDSTSTSEALNRTLGEEARSINSLVVGGAPTPPPTPTA 599
Query: 55 DLPQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPM-------KYAEFLPSLLKGNYVQTR 107
+ Q QQ+ Q +QP QQ +QQ + +F +P PSL K T
Sbjct: 600 EPEQQQQQQQQQQQPQQQQQQQQQQQQQQFVVLPKIKDLTLPTSPPLPPSLFKAYTPSTT 659
Query: 108 PPPP 111
P P
Sbjct: 660 PTAP 663
>SN22_HUMAN (P51531) Possible global transcription activator SNF2L2
(SNF2-alpha) (SWI/SNF related matrix associated actin
dependent regulator of chromatin subfamily A member 2)
Length = 1586
Score = 33.1 bits (74), Expect = 0.30
Identities = 30/103 (29%), Positives = 42/103 (40%), Gaps = 15/103 (14%)
Query: 25 ELHELRKEMTALRAKV------EELTNRVSSLMVTKDLPQFQQRPQPPRQPSFTQQPRQQ 78
+LH+LR ++ A + E L V L Q QQ+ Q +Q QQ +QQ
Sbjct: 178 QLHQLRAQILAYKMLARGQPLPETLQLAVQGKRTLPGLQQQQQQQQQQQQQQQQQQQQQQ 237
Query: 79 APRTRFDPIPMKYAEFLPSLLKGNYVQTRPP----PPRPEKLP 117
P P P + P+L+ N P P P+KLP
Sbjct: 238 QP-----PQPQTQQQQQPALVNYNRPSGPGPELSGPSTPQKLP 275
>GOA2_HUMAN (Q08379) Golgi autoantigen, golgin subfamily A member 2
(Golgi matrix protein GM130) (Gm130 autoantigen)
(Golgin-95)
Length = 990
Score = 33.1 bits (74), Expect = 0.30
Identities = 18/60 (30%), Positives = 31/60 (51%), Gaps = 3/60 (5%)
Query: 18 KMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQPPRQPSFTQQPRQ 77
+M Q+ +++H LR+E ++V+EL ++ L P P+PP PS +Q Q
Sbjct: 398 RMQQMSEQVHTLREEKECSMSRVQELETSLAELRNQMAEP---PPPEPPAGPSEVEQQLQ 454
>FXJ3_MOUSE (Q8BUR3) Forkhead box protein J3
Length = 623
Score = 33.1 bits (74), Expect = 0.30
Identities = 21/68 (30%), Positives = 31/68 (44%), Gaps = 2/68 (2%)
Query: 56 LPQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEK 115
LPQ QRPQ P P Q + Q P ++ P P ++ + P+ PPPP+
Sbjct: 391 LPQHPQRPQHPA-PHPQQHSQLQPPHSQHPP-PHQHIQHHPNHQHQTLAHQPPPPPQQVS 448
Query: 116 LPAGFRAD 123
+G +D
Sbjct: 449 CNSGVSSD 456
>CBP_HUMAN (Q92793) CREB-binding protein (EC 2.3.1.48)
Length = 2442
Score = 33.1 bits (74), Expect = 0.30
Identities = 30/107 (28%), Positives = 45/107 (42%), Gaps = 18/107 (16%)
Query: 27 HELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQP-------PRQPSFTQQPRQQA 79
H+LR++ R + +L R + M T+++PQ Q P P P Q T Q Q
Sbjct: 1848 HKLRQQQIQHRLQQAQLMRRRMATMNTRNVPQ-QSLPSPTSAPPGTPTQQPSTPQTPQPP 1906
Query: 80 PRTRFDPIPMKYAEFLPSLLK---------GNYVQTRPPPPRPEKLP 117
+ + P+ M A F PS+ + G P PP P + P
Sbjct: 1907 AQPQPSPVSMSPAGF-PSVARTQPPTTVSTGKPTSQVPAPPPPAQPP 1952
>YKF4_YEAST (P35732) Hypothetical 84.0 kDa protein in NUP120-CSE4
intergenic region
Length = 738
Score = 32.7 bits (73), Expect = 0.40
Identities = 24/73 (32%), Positives = 33/73 (44%), Gaps = 17/73 (23%)
Query: 23 EQELHELRKEMTALRAKVEELT---NRVSSLMVTKDLPQFQ--------------QRPQP 65
EQ EL +E + A EE+T +V V + P+ Q Q+PQ
Sbjct: 336 EQTAEELEQEQDNVAAPEEEVTVVEEKVEISAVISEPPEDQANTVPQPQQQSQQPQQPQQ 395
Query: 66 PRQPSFTQQPRQQ 78
P+QP QQP+QQ
Sbjct: 396 PQQPQQPQQPQQQ 408
Score = 29.3 bits (64), Expect = 4.4
Identities = 12/21 (57%), Positives = 15/21 (71%)
Query: 57 PQFQQRPQPPRQPSFTQQPRQ 77
PQ Q+PQ P+QP QQP+Q
Sbjct: 393 PQQPQQPQQPQQPQQQQQPQQ 413
Score = 28.5 bits (62), Expect = 7.4
Identities = 25/90 (27%), Positives = 36/90 (39%), Gaps = 22/90 (24%)
Query: 11 QTLAQTHKMDQLEQELHEL---RKEMTALRAKVE----------ELTNRVSSLMVTKDLP 57
Q + ++LEQE + +E+T + KVE + N V P
Sbjct: 331 QVKEEEQTAEELEQEQDNVAAPEEEVTVVEEKVEISAVISEPPEDQANTVPQPQQQSQQP 390
Query: 58 QFQQRPQPPRQPSFTQ---------QPRQQ 78
Q Q+PQ P+QP Q QP+QQ
Sbjct: 391 QQPQQPQQPQQPQQPQQQQQPQQPQQPQQQ 420
Score = 28.1 bits (61), Expect = 9.7
Identities = 18/47 (38%), Positives = 25/47 (52%), Gaps = 4/47 (8%)
Query: 57 PQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNY 103
PQ QQ+PQ P+QP QQ QQ + + P+ + A+ L NY
Sbjct: 405 PQQQQQPQQPQQP---QQQLQQQQQQQQQPVQAQ-AQAQEEQLSQNY 447
>MEFD_HUMAN (Q14814) Myocyte-specific enhancer factor 2D
Length = 521
Score = 32.7 bits (73), Expect = 0.40
Identities = 21/55 (38%), Positives = 29/55 (52%), Gaps = 3/55 (5%)
Query: 57 PQFQQRPQPPRQ-PSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPP 110
PQ Q+PQPP+Q P QQP+ Q P+ P P + + +P L N + P P
Sbjct: 370 PQQPQQPQPPQQQPPQPQQPQPQQPQQPQQP-PQQQSHLVPVSL-SNLIPGSPLP 422
Score = 28.5 bits (62), Expect = 7.4
Identities = 11/21 (52%), Positives = 15/21 (71%)
Query: 61 QRPQPPRQPSFTQQPRQQAPR 81
Q+PQ P+QP Q P+QQ P+
Sbjct: 365 QQPQQPQQPQQPQPPQQQPPQ 385
>BRD4_MOUSE (Q9ESU6) Bromodomain-containing protein 4 (Mitotic
chromosome-associated protein) (MCAP)
Length = 1400
Score = 32.7 bits (73), Expect = 0.40
Identities = 23/64 (35%), Positives = 30/64 (45%), Gaps = 7/64 (10%)
Query: 58 QFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPP--RPEK 115
Q Q +PQPP P QP P+ + P P LPS+ ++Q PPPP +P
Sbjct: 970 QQQLQPQPPPPPPPQPQP---PPQQQHQPPPRPV--HLPSMPFSAHIQQPPPPPGQQPTH 1024
Query: 116 LPAG 119
P G
Sbjct: 1025 PPPG 1028
Score = 28.9 bits (63), Expect = 5.7
Identities = 14/37 (37%), Positives = 18/37 (47%)
Query: 57 PQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAE 93
P QQ+ QPP P P+Q AP + P P A+
Sbjct: 770 PPQQQQQQPPPPPPPPSMPQQTAPAMKSSPPPFITAQ 806
Score = 28.1 bits (61), Expect = 9.7
Identities = 15/54 (27%), Positives = 22/54 (39%), Gaps = 6/54 (11%)
Query: 62 RPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEK 115
RP+PP+ P + P R P+ + P Q+ PPP P+K
Sbjct: 1166 RPEPPKHPENIKAPVHLPQRPEMKPVDIGRPVIRPP------EQSAPPPGAPDK 1213
>ASD4_NEUCR (Q9HEV5) GATA type zinc finger protein Asd4 (Ascus
development protein 4)
Length = 426
Score = 32.7 bits (73), Expect = 0.40
Identities = 31/124 (25%), Positives = 52/124 (41%), Gaps = 25/124 (20%)
Query: 6 LSAHMQTLAQTHKMDQLEQELHELRKEMTALRAKVEELT---NRVSSLMVTKDLPQFQQ- 61
L A + + + H+ QL++EL E + L+ +++EL V + ++D + ++
Sbjct: 234 LRAQIDAMGEAHQ--QLQKELEESHRRENMLKRRLDELEVELKDVKDALESQDNGRHKKI 291
Query: 62 ------RPQPPRQPSFTQQPRQQAPRTRFDPI--PMKYAEFLPSLLKGNYVQTRPPPPRP 113
+ +P + QQP QQ P PI PM E P+ P P P
Sbjct: 292 RLDENVKTEPYAEVVEPQQPEQQQPAPAEQPIPTPMAIDEATPA-----------PAPAP 340
Query: 114 EKLP 117
E P
Sbjct: 341 EAAP 344
>104K_THEPA (P15711) 104 kDa microneme-rhoptry antigen
Length = 924
Score = 32.7 bits (73), Expect = 0.40
Identities = 20/74 (27%), Positives = 35/74 (47%), Gaps = 6/74 (8%)
Query: 41 EELTNRVSSLMVTKDLPQFQQRPQPPRQPSFTQQPRQQAPRTRFDP-IPMKYAEFLPSLL 99
E +++ +L P+ + P+ P++P ++ PR +P R P +P L L
Sbjct: 558 EHKPSKIPTLSKKPSGPKDPKHPRDPKEPRKSKSPRTASPTRRPSPKLPQ-----LSKLP 612
Query: 100 KGNYVQTRPPPPRP 113
K ++ PPP RP
Sbjct: 613 KSTSPRSPPPPTRP 626
>TIS1_DROME (P47980) TIS11 protein (dTIS11)
Length = 437
Score = 32.3 bits (72), Expect = 0.52
Identities = 19/84 (22%), Positives = 43/84 (50%), Gaps = 1/84 (1%)
Query: 11 QTLAQTHKMDQLEQELHELRKEMTALR-AKVEELTNRVSSLMVTKDLPQFQQRPQPPRQP 69
+T++Q ++ Q +Q+ H+ +++ A + +T + ++ + + L + Q P PP+QP
Sbjct: 72 RTISQPAQLIQQQQQQHQQQQQQQQPPVASLVTITENLGNMNLHRKLERTQSEPLPPQQP 131
Query: 70 SFTQQPRQQAPRTRFDPIPMKYAE 93
T + + + R + KY E
Sbjct: 132 MNTSRYKTELCRPFEEAGECKYGE 155
>MYS3_HUMAN (Q92794) MYST histone acetyltransferase 3 (Runt-related
transcription factor binding protein 2) (Monocytic
leukemia zinc finger protein) (Zinc finger protein 220)
Length = 2004
Score = 32.3 bits (72), Expect = 0.52
Identities = 20/61 (32%), Positives = 25/61 (40%), Gaps = 16/61 (26%)
Query: 57 PQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEKL 116
PQ Q P P QP+ P QQ P+ + P P Q PPPP P++
Sbjct: 1656 PQQPQPPPPQPQPAPQPPPPQQQPQQQPQPQP----------------QQPPPPPPPQQQ 1699
Query: 117 P 117
P
Sbjct: 1700 P 1700
Score = 30.4 bits (67), Expect = 2.0
Identities = 29/109 (26%), Positives = 49/109 (44%), Gaps = 6/109 (5%)
Query: 57 PQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEKL 116
PQ Q PQPP QQP+QQ P P + P L + + + P P ++
Sbjct: 1664 PQPQPAPQPPPPQ---QQPQQQPQPQPQQPPPPPPPQQQPPLSQCSMNNSFTPAPMIMEI 1720
Query: 117 P-AGFRADLSCVFHQGAPGHDVERCYALRNAVQDLVEAKILSFTDLNPN 164
P +G ++S ++ PG Y+ +A L + + L+ T ++P+
Sbjct: 1721 PESGSTGNIS--IYERIPGDFGAGSYSQPSATFSLAKLQQLTNTIMDPH 1767
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.132 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,945,998
Number of Sequences: 164201
Number of extensions: 979119
Number of successful extensions: 6529
Number of sequences better than 10.0: 209
Number of HSP's better than 10.0 without gapping: 52
Number of HSP's successfully gapped in prelim test: 165
Number of HSP's that attempted gapping in prelim test: 5937
Number of HSP's gapped (non-prelim): 568
length of query: 167
length of database: 59,974,054
effective HSP length: 102
effective length of query: 65
effective length of database: 43,225,552
effective search space: 2809660880
effective search space used: 2809660880
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144591.1


