

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144541.14 - phase: 0
(63 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
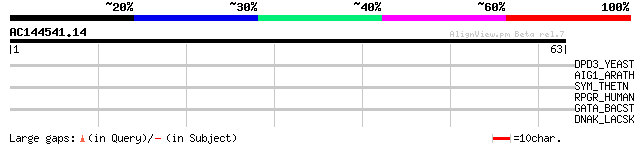
Score E
Sequences producing significant alignments: (bits) Value
DPD3_YEAST (P47110) DNA polymerase delta subunit 3 29 2.0
AIG1_ARATH (P54120) AIG1 protein 29 2.0
SYM_THETN (Q8RDD1) Methionyl-tRNA synthetase (EC 6.1.1.10) (Meth... 28 4.5
RPGR_HUMAN (Q92834) X-linked retinitis pigmentosa GTPase regulator 28 5.8
GATA_BACST (Q93LE2) Glutamyl-tRNA(Gln) amidotransferase subunit ... 28 5.8
DNAK_LACSK (O87777) Chaperone protein dnaK (Heat shock protein 7... 27 9.9
>DPD3_YEAST (P47110) DNA polymerase delta subunit 3
Length = 350
Score = 29.3 bits (64), Expect = 2.0
Identities = 17/54 (31%), Positives = 31/54 (56%), Gaps = 6/54 (11%)
Query: 1 PNGERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNEL 54
P ++P + + ++KD G+R +A +L K+ K ++DKE S++NEL
Sbjct: 140 PTARSKTTPEETTGRKSKSKDMGLRSTA-----LLAKMKKDRDDKET-SRQNEL 187
>AIG1_ARATH (P54120) AIG1 protein
Length = 353
Score = 29.3 bits (64), Expect = 2.0
Identities = 15/49 (30%), Positives = 31/49 (62%), Gaps = 1/49 (2%)
Query: 12 LSKFLGRNKDDGVRRSAV-SGKKILLKLDKTKEDKEAESKRNELLNFLN 59
L +LG N D ++R + G++++L +KTK+D++ + +ELL ++
Sbjct: 180 LEDYLGDNMPDFLKRVLILCGQRMILFDNKTKDDEKKTKQVHELLKLID 228
>SYM_THETN (Q8RDD1) Methionyl-tRNA synthetase (EC 6.1.1.10)
(Methionine--tRNA ligase) (MetRS)
Length = 638
Score = 28.1 bits (61), Expect = 4.5
Identities = 12/38 (31%), Positives = 23/38 (59%)
Query: 26 RSAVSGKKILLKLDKTKEDKEAESKRNELLNFLNASFD 63
R + G K+L ++ + + ES+RNE++NF+ A +
Sbjct: 162 RLSAYGDKLLKYYEEHPDFIQPESRRNEMINFIKAGLE 199
>RPGR_HUMAN (Q92834) X-linked retinitis pigmentosa GTPase regulator
Length = 1020
Score = 27.7 bits (60), Expect = 5.8
Identities = 14/51 (27%), Positives = 24/51 (46%)
Query: 4 ERSSSPVQLSKFLGRNKDDGVRRSAVSGKKILLKLDKTKEDKEAESKRNEL 54
ER S ++S F G D G+ + K + K + K+D +S+R +
Sbjct: 660 EREKSTKKMSPFFGNLPDRGMNTESEENKDFVKKRESCKQDVIFDSERESV 710
>GATA_BACST (Q93LE2) Glutamyl-tRNA(Gln) amidotransferase subunit A
(EC 6.3.5.-) (Glu-ADT subunit A)
Length = 485
Score = 27.7 bits (60), Expect = 5.8
Identities = 12/32 (37%), Positives = 21/32 (65%)
Query: 13 SKFLGRNKDDGVRRSAVSGKKILLKLDKTKED 44
+++LG D+GVR+S ++ +L KL T E+
Sbjct: 265 NEYLGEGVDEGVRQSVLAALAVLEKLGATWEE 296
>DNAK_LACSK (O87777) Chaperone protein dnaK (Heat shock protein 70)
(Heat shock 70 kDa protein) (HSP70)
Length = 613
Score = 26.9 bits (58), Expect = 9.9
Identities = 15/50 (30%), Positives = 30/50 (60%), Gaps = 3/50 (6%)
Query: 13 SKFLGRNKDDGVR---RSAVSGKKILLKLDKTKEDKEAESKRNELLNFLN 59
+K LG NK+ + S +S ++I + + +E++EA++KR E ++ N
Sbjct: 459 AKDLGTNKEQKITIKSNSGLSDEEIDRMMKEAQENEEADTKRKEEVDLKN 508
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.307 0.129 0.336
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,692,052
Number of Sequences: 164201
Number of extensions: 207357
Number of successful extensions: 607
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 604
Number of HSP's gapped (non-prelim): 6
length of query: 63
length of database: 59,974,054
effective HSP length: 39
effective length of query: 24
effective length of database: 53,570,215
effective search space: 1285685160
effective search space used: 1285685160
T: 11
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC144541.14


