

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
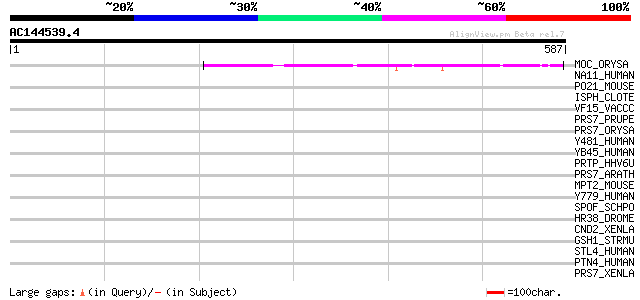
Query= AC144539.4 - phase: 0
(587 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
MOC_ORYSA (Q84MM9) Protein MONOCULM 1 100 2e-20
NA11_HUMAN (P59045) NACHT-, LRR- and PYD-containing protein 11 (... 36 0.25
PO21_MOUSE (P25425) POU domain, class 2, transcription factor 1 ... 34 0.93
ISPH_CLOTE (Q895G2) 4-hydroxy-3-methylbut-2-enyl diphosphate red... 34 0.93
VF15_VACCC (P21020) Protein F15 34 1.2
PRS7_PRUPE (O64982) 26S protease regulatory subunit 7 (26S prote... 34 1.2
PRS7_ORYSA (Q9FXT9) 26S protease regulatory subunit 7 (26S prote... 34 1.2
Y481_HUMAN (O75069) Hypothetical protein KIAA0481 (Cerebral prot... 33 1.6
YB45_HUMAN (Q9ULS5) Hypothetical protein KIAA1145 (Fragment) 33 2.7
PRTP_HHV6U (P52384) Probable processing and transport protein 33 2.7
PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 (26S prote... 33 2.7
MPT2_MOUSE (Q8BG95) Protein phosphatase 1 regulatory subunit 12B... 33 2.7
Y779_HUMAN (O94876) Hypothetical protein KIAA0779 (Fragment) 32 3.5
SPOF_SCHPO (Q10411) Sporulation-specific protein 15 32 3.5
HR38_DROME (P49869) Probable nuclear hormone receptor HR38 (dHR38) 32 4.6
CND2_XENLA (O13067) Condensin subunit 2 (Chromosome-associated p... 32 4.6
GSH1_STRMU (Q8DW15) Probable glutamate--cysteine ligase (EC 6.3.... 32 6.0
STL4_HUMAN (Q96C24) Synaptotagmin-like protein 4 (Exophilin 2) (... 31 7.9
PTN4_HUMAN (P29074) Protein tyrosine phosphatase, non-receptor t... 31 7.9
PRS7_XENLA (P46472) 26S protease regulatory subunit 7 (MSS1 prot... 31 7.9
>MOC_ORYSA (Q84MM9) Protein MONOCULM 1
Length = 441
Score = 99.8 bits (247), Expect = 2e-20
Identities = 99/408 (24%), Positives = 175/408 (42%), Gaps = 47/408 (11%)
Query: 206 SLAESLLACAEKVGYQQYERARKLLSQI-ESLSSKTGNPVKRVVHYFAEALCQRIDKETG 264
S + LLACA+ + AR+ + + +S G+ R+ ++FA AL R+D + G
Sbjct: 49 STRDLLLACADLLQRGDLPAARRAAEIVLAAAASPRGDAADRLAYHFARALALRVDAKAG 108
Query: 265 RFSVSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLENVNDAKKI 324
V + + A +A + PF + + T QA+LE V+ A+++
Sbjct: 109 HGHVVVGGGAARPAS----------SGAYLAFNQIAPFLRFAHLTANQAILEAVDGARRV 158
Query: 325 HVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTGKRLKDFAQ 384
H++DL+ G QW L+QA+ R + L ++ +G + + TG RL+ FA+
Sbjct: 159 HILDLDAVHGVQWPPLLQAIAERADPALGPPEVRVTGAG---ADRDTLLRTGNRLRAFAR 215
Query: 385 SLNIPFSFDIVVVSDLLHIREEL-------------------FKIDSEETVAVYSQFALR 425
S+++PF F +++S + + +ET+AV L
Sbjct: 216 SIHLPFHFTPLLLSCATTAPHHVAGTSTGAAAAASTAAAATGLEFHPDETLAVNCVMFLH 275
Query: 426 SKIQQPDKLETIMRVIRTINPIVMVVAEIEA------NHNSKSFVNRFIEALFYFSAYFD 479
+ + D+L ++ ++ ++P V+ +AE EA + R A+ ++SA F+
Sbjct: 276 N-LAGHDELAAFLKWVKAMSPAVVTIAEREAGGGGGGGDHIDDLPRRVGVAMDHYSAVFE 334
Query: 480 CFETCMKGDEKNRFILESMYFSHGIRNIVAEEGAERKSRNVKIDVWRAFFTRFGMVETEL 539
E + + R +E I V G R R I+ W G L
Sbjct: 335 ALEATVPPGSRERLAVEQEVLGREIEAAVGPSGG-RWWRG--IERWGGAARAAGFAARPL 391
Query: 540 SMKSLYQAELVAKRF--ACGYACTFDMNGHCLLVGWKGTPINSVSVWK 585
S ++ QA L+ + + GY + G C L GW+ P+ SVS W+
Sbjct: 392 SAFAVSQARLLLRLHYPSEGYLVQ-EARGACFL-GWQTRPLLSVSAWQ 437
>NA11_HUMAN (P59045) NACHT-, LRR- and PYD-containing protein 11
(PYRIN-containing APAF1-like protein 6)
(Nucleotide-binding oligomerization domain protein 17)
Length = 1033
Score = 36.2 bits (82), Expect = 0.25
Identities = 27/123 (21%), Positives = 52/123 (41%), Gaps = 1/123 (0%)
Query: 52 IESLCSNFGFFQDDPSYQENEFLKFHHQQQESYQDYETFDNLHFDMVHFGDTKKDQPFYQ 111
++ C+ F P+Y + + +++E Y D+ F +++ K + +
Sbjct: 453 VQEFCTAIAFLMAVPNYLIPSGSREYKEKREQYSDFNQVFTFIFGLLNANRRKILETSFG 512
Query: 112 TPLAPVDILKNYGKGFKRLLPDDEGNILHPVNDFGLVTNNENEKKLLSTIEIMKMAGTRF 171
L VD K Y G+ + L D + H + F + N E++ + TI M T +
Sbjct: 513 YQLPMVDSFKWYSVGYMKHLDRDPEKLTHHMPLFYCLYEN-REEEFVKTIVDALMEVTVY 571
Query: 172 IQS 174
+QS
Sbjct: 572 LQS 574
>PO21_MOUSE (P25425) POU domain, class 2, transcription factor 1
(Octamer-binding transcription factor 1) (Oct-1) (OTF-1)
(NF-A1)
Length = 770
Score = 34.3 bits (77), Expect = 0.93
Identities = 39/167 (23%), Positives = 67/167 (39%), Gaps = 14/167 (8%)
Query: 175 SSSSKSASGLILNHPFGFSFSGLSDEEKENVSLAESLLACAEKVGYQQYERARKLLSQIE 234
SS S ++S LN P G GL+ K+ S+ ++ EK + +K S+
Sbjct: 358 SSDSTASSPSALNSP-GLGAEGLNRRRKKRTSIETNIRVALEK----SFMENQKPTSEDI 412
Query: 235 SLSSKTGNPVKRVVHYFAEALCQRIDKETGRFSVSSNNMQKMESLFDPQEVSKDLNPAMI 294
+L ++ N K V+ + C R KE SS P + +++
Sbjct: 413 TLIAEQLNMEKEVIRVW---FCNRRQKEKRINPPSSGGTSS-----SPIKAIFPSPASLV 464
Query: 295 ALYEDLPFSQVSIFTCVQALLENVNDA-KKIHVIDLEIRKGCQWTIL 340
A L S + V +L + A + + D ++R+GC W +L
Sbjct: 465 ATTPSLVTSSTATTLTVNPVLPLTSAAVTNLSLTDQDLRRGCSWEVL 511
>ISPH_CLOTE (Q895G2) 4-hydroxy-3-methylbut-2-enyl diphosphate
reductase (EC 1.17.1.2)
Length = 635
Score = 34.3 bits (77), Expect = 0.93
Identities = 35/148 (23%), Positives = 66/148 (43%), Gaps = 16/148 (10%)
Query: 268 VSSNNMQKMESLFDPQEVSKDLNPAMIALYEDLPFSQVSIFTCVQALLENVNDAKKIHVI 327
+S + + + + +E ++ ++ + ED+ +SI+ + + + + HV
Sbjct: 365 ISVIELNRENAYVELKEAFENKEKVVVKVKEDVNGGLISIYKNIVRVFIPASHVELRHVD 424
Query: 328 DLEIRKGCQWTI-LMQALQSRNECPL-----ELLK----ITAIESGNSDTSKHIVEDTGK 377
DL I KGC+ T+ +++ + RN + +LLK E+ +S I E +
Sbjct: 425 DLSIYKGCELTVNIIEFEEGRNNTRIVASRRDLLKEEQSKVEEETWSSLEKDTIKEGEVR 484
Query: 378 RLKDFAQSLNIPFSFDIVVVSDLLHIRE 405
RL DF +NI V LLH+ E
Sbjct: 485 RLTDFGAFVNIN------GVDGLLHVSE 506
>VF15_VACCC (P21020) Protein F15
Length = 158
Score = 33.9 bits (76), Expect = 1.2
Identities = 26/90 (28%), Positives = 43/90 (46%), Gaps = 4/90 (4%)
Query: 139 LHPVNDFGLVTNNENEKKLLSTIEIMKMAGTRFIQSSSSSKSASGLILNHPFGFS---FS 195
+ P + +T N+N+ +L+ +K+ R+I S+S S S I NH F S +S
Sbjct: 23 MKPFKNMNKITINQNDNCILANRCFVKIDTPRYIPSTSISSSNIIRIRNHDFTLSELLYS 82
Query: 196 GLSDEEKE-NVSLAESLLACAEKVGYQQYE 224
++ + L +L C +KV QQ E
Sbjct: 83 PFHFQQPQFQYLLPGFVLTCIDKVSKQQKE 112
>PRS7_PRUPE (O64982) 26S protease regulatory subunit 7 (26S
proteasome subunit 7) (26S proteasome AAA-ATPase subunit
RPT1) (Regulatory particle triple-A ATPase subunit 1)
Length = 425
Score = 33.9 bits (76), Expect = 1.2
Identities = 24/102 (23%), Positives = 47/102 (45%), Gaps = 10/102 (9%)
Query: 281 DPQEVSKDLNP--------AMIALYEDLPFSQ--VSIFTCVQALLENVNDAKKIHVIDLE 330
+P+++ + NP A++ Y P+S V+ L + VND I D
Sbjct: 4 EPEDIKDEKNPRPLDEDDIALLKTYGLGPYSTHIKKAEKEVKDLAKKVNDLCGIKESDTG 63
Query: 331 IRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIV 372
+ QW ++ + E PL++ + T I + NS+ +K+++
Sbjct: 64 LAAPSQWDLVSDKQMMQEEQPLQVARCTKIINPNSEDAKYVI 105
>PRS7_ORYSA (Q9FXT9) 26S protease regulatory subunit 7 (26S
proteasome subunit 7) (26S proteasome AAA-ATPase subunit
RPT1) (Regulatory particle triple-A ATPase subunit 1)
Length = 426
Score = 33.9 bits (76), Expect = 1.2
Identities = 15/62 (24%), Positives = 32/62 (51%)
Query: 311 VQALLENVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKH 370
++ + + +ND I D + QW ++ + E PL++ + T I S N+D +K+
Sbjct: 45 IKEMAKKINDLCGIKESDTGLAPPSQWDLVSDKQMMQEEQPLQVARCTKIISPNTDDAKY 104
Query: 371 IV 372
++
Sbjct: 105 VI 106
>Y481_HUMAN (O75069) Hypothetical protein KIAA0481 (Cerebral
protein-11) (hucep-11) (Fragment)
Length = 650
Score = 33.5 bits (75), Expect = 1.6
Identities = 15/35 (42%), Positives = 21/35 (59%)
Query: 201 EKENVSLAESLLACAEKVGYQQYERARKLLSQIES 235
+ E +L + L + EKV YQ YERAR + +ES
Sbjct: 518 QNEMTNLKQELASMEEKVAYQSYERARDIQEAVES 552
>YB45_HUMAN (Q9ULS5) Hypothetical protein KIAA1145 (Fragment)
Length = 467
Score = 32.7 bits (73), Expect = 2.7
Identities = 14/39 (35%), Positives = 23/39 (58%)
Query: 201 EKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSK 239
+ E +L + L + EKV YQ YER+R + +ES ++
Sbjct: 338 QHETANLKQELASIEEKVAYQAYERSRDIQEALESCQTR 376
>PRTP_HHV6U (P52384) Probable processing and transport protein
Length = 726
Score = 32.7 bits (73), Expect = 2.7
Identities = 28/134 (20%), Positives = 59/134 (43%), Gaps = 12/134 (8%)
Query: 161 IEIMKMAGTRFIQSSSSSKSASGLILNHPFGFSFSGLSDEEKENVSLAESLLACAEKVGY 220
+E +K + S ++ +G+++ H + F G+ ++ N+ A +L + +
Sbjct: 21 LECLKFCDPVIVLSDMANFKKNGIVILHLYQTFFEGIKEQ---NLLCASALTVYMQVLLK 77
Query: 221 QQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALCQRIDKETGRFSVSSNN----MQKM 276
YE+ L + +ES YF + LC R ET ++S NN + ++
Sbjct: 78 AMYEQVLLLDAALESFMVDQDRK-----KYFEKVLCLRRCAETLSINISLNNGVEFIVQL 132
Query: 277 ESLFDPQEVSKDLN 290
+L D +++ +N
Sbjct: 133 STLNDIEQLISKIN 146
>PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 (26S
proteasome subunit 7) (26S proteasome AAA-ATPase subunit
RPT1a) (Regulatory particle triple-A ATPase subunit 1a)
Length = 426
Score = 32.7 bits (73), Expect = 2.7
Identities = 15/62 (24%), Positives = 32/62 (51%)
Query: 311 VQALLENVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKH 370
++ L + +ND I D + QW ++ + E PL++ + T I S N++ +K+
Sbjct: 45 IKDLAKKINDLCGIKESDTGLAPPSQWDLVSDKQMMQEEQPLQVARCTKIISPNTEDAKY 104
Query: 371 IV 372
++
Sbjct: 105 VI 106
>MPT2_MOUSE (Q8BG95) Protein phosphatase 1 regulatory subunit 12B
(Myosin phosphatase targeting subunit 2) (Fragment)
Length = 484
Score = 32.7 bits (73), Expect = 2.7
Identities = 38/156 (24%), Positives = 68/156 (43%), Gaps = 14/156 (8%)
Query: 142 VNDFGLVTNNENEKKLLSTIEIMKMAGTRFIQSSSSSKSASGLILNHP---FGFSFSGLS 198
V D GLV + E +K + K + I+S +SK SGL N +
Sbjct: 289 VADEGLVEHLEMLQKKQDVLRSEKETRNKLIESDLNSKFQSGLFKNKEKMLYEEEIPKSQ 348
Query: 199 DEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEALCQR 258
D E+EN ES + +E+ E +S+ E+ P V H +E +
Sbjct: 349 DTEEEN---KESSSSSSEE------EEGEDEVSESETEKEADKKPEATVNHSNSEIKSRI 399
Query: 259 IDK--ETGRFSVSSNNMQKMESLFDPQEVSKDLNPA 292
+++ + + S+++ +++ SLF+ E KD +P+
Sbjct: 400 MEQIPAPAQNTFSASSARRLSSLFNKAEEPKDESPS 435
>Y779_HUMAN (O94876) Hypothetical protein KIAA0779 (Fragment)
Length = 320
Score = 32.3 bits (72), Expect = 3.5
Identities = 13/39 (33%), Positives = 24/39 (61%)
Query: 201 EKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSK 239
+ E ++L + L + EK+ YQ YERAR + +E+ ++
Sbjct: 187 QNEILNLKQELASMEEKIAYQSYERARDIQEALEACQTR 225
>SPOF_SCHPO (Q10411) Sporulation-specific protein 15
Length = 1957
Score = 32.3 bits (72), Expect = 3.5
Identities = 29/124 (23%), Positives = 54/124 (43%), Gaps = 8/124 (6%)
Query: 357 ITAIESGNSDTSKHIVE--DTGKRLKDFAQSLNIPFSFDIVVVSDLLHIREEL--FKIDS 412
I ++E NS+ + E D RL D +SL+ + V+ SDL+ + +EL K D
Sbjct: 1157 IVSLEQSNSNNEALVEERSDLANRLSDMKKSLSDSDNVISVIRSDLVRVNDELDTLKKDK 1216
Query: 413 EETVAVYSQFALRSKIQQPDKLETIMRVIRTINPIVMVVAEIEANHNSKSFVNRFIEALF 472
+ YS+ + D L+++ + N + + E+ V+ ++ F
Sbjct: 1217 DSLSTQYSEVCQ----DRDDLLDSLKGCEESFNKYAVSLRELCTKSEIDVPVSEILDDNF 1272
Query: 473 YFSA 476
F+A
Sbjct: 1273 VFNA 1276
>HR38_DROME (P49869) Probable nuclear hormone receptor HR38 (dHR38)
Length = 1073
Score = 32.0 bits (71), Expect = 4.6
Identities = 16/53 (30%), Positives = 27/53 (50%), Gaps = 3/53 (5%)
Query: 86 DYETFDNLHFDMVHFGDTKKDQPFYQTPLAPVDILKNYGK---GFKRLLPDDE 135
D D H++ + K Q FYQ + VD++K + + G+ LLP+D+
Sbjct: 858 DPSCLDYSHYEEQSMSEADKVQQFYQLLTSSVDVIKQFAEKIPGYFDLLPEDQ 910
>CND2_XENLA (O13067) Condensin subunit 2 (Chromosome-associated
protein H) (Chromosome assembly protein XCAP-H) (Barren
homolog)
Length = 699
Score = 32.0 bits (71), Expect = 4.6
Identities = 23/87 (26%), Positives = 39/87 (44%), Gaps = 6/87 (6%)
Query: 321 AKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKHIVEDTGKRLK 380
AK +D++ K W++L +S+ E P +I A + + KRL
Sbjct: 598 AKTAKKMDMKRLKSSMWSLLANCPESQEEMPSSKEEIDAALITDEQVFSSVTHGLQKRLP 657
Query: 381 D-FAQSLNIPFSFDIVVVSDLLHIREE 406
AQ+L++P +F + LLH+ E
Sbjct: 658 PVMAQNLSVPLAF-----ACLLHLANE 679
>GSH1_STRMU (Q8DW15) Probable glutamate--cysteine ligase (EC
6.3.2.2) (Gamma-glutamylcysteine synthetase) (Gamma-ECS)
(GCS)
Length = 754
Score = 31.6 bits (70), Expect = 6.0
Identities = 32/140 (22%), Positives = 59/140 (41%), Gaps = 8/140 (5%)
Query: 412 SEETVAVYSQFALRSK-IQQPDKLETIMRVIRTINPIVMVVAEIEA--NHNSKSFVNRFI 468
++ET+ F L + P KL + + +N + + +E NS S + +
Sbjct: 295 TQETIDSVHLFILAMLWLDTPKKLNQALDKAQKLNDKIALSHPLEKLPKENSASLIIEAM 354
Query: 469 EAL---FYFSAYFDCFETCMKGDEKN-RFILESMYFSHGIRNIVAEEGAERKSRNVKIDV 524
EAL F +Y+D +K +N + L F H I++ E ++K ++
Sbjct: 355 EALIKHFKLPSYYDDLLIAIKKQVENPKLTLSGRLFEH-IKHASLEHFGQKKGQDYHNYA 413
Query: 525 WRAFFTRFGMVETELSMKSL 544
W+ ++ G ELS + L
Sbjct: 414 WQNYYALKGYENMELSTQML 433
>STL4_HUMAN (Q96C24) Synaptotagmin-like protein 4 (Exophilin 2)
(Granuphilin)
Length = 671
Score = 31.2 bits (69), Expect = 7.9
Identities = 23/77 (29%), Positives = 39/77 (49%), Gaps = 6/77 (7%)
Query: 195 SGLSDEEKENVSLAESLLACAEKVGYQQYERARKLLSQIESLSSKTGNPVKRVVHYFAEA 254
S LS+EEK+ L S+L E+V +R R+L +++ + K KR ++++
Sbjct: 8 SFLSEEEKD---LILSVLQRDEEVRKADEKRIRRLKNELLEIKRKGA---KRGSQHYSDR 61
Query: 255 LCQRIDKETGRFSVSSN 271
C R + GR S +N
Sbjct: 62 TCARCQESLGRLSPKTN 78
>PTN4_HUMAN (P29074) Protein tyrosine phosphatase, non-receptor type
4 (EC 3.1.3.48) (Protein-tyrosine phosphatase MEG1)
(PTPase-MEG1) (MEG)
Length = 926
Score = 31.2 bits (69), Expect = 7.9
Identities = 16/43 (37%), Positives = 23/43 (53%), Gaps = 3/43 (6%)
Query: 40 FSVAEDLGDYNGIESL---CSNFGFFQDDPSYQENEFLKFHHQ 79
F+V +LGDY+ E+L S++ F + P E E K H Q
Sbjct: 152 FAVQSELGDYDQSENLSGYLSDYSFIPNQPQDFEKEIAKLHQQ 194
>PRS7_XENLA (P46472) 26S protease regulatory subunit 7 (MSS1
protein)
Length = 433
Score = 31.2 bits (69), Expect = 7.9
Identities = 18/74 (24%), Positives = 36/74 (48%), Gaps = 5/74 (6%)
Query: 311 VQALLENVNDAKKIHVIDLEIRKGCQWTILMQALQSRNECPLELLKITAIESGNSDTSKH 370
+Q LL+ +N+ I D + W + ++E PL++ + T I + +S+ K+
Sbjct: 52 IQQLLKKINELTGIKESDTGLAPPALWDLAADKQTLQSEQPLQVARCTKIINADSEDPKY 111
Query: 371 IVEDTGKRLKDFAQ 384
I+ +K FA+
Sbjct: 112 II-----NVKQFAK 120
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.135 0.391
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 67,058,641
Number of Sequences: 164201
Number of extensions: 2782482
Number of successful extensions: 7256
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 15
Number of HSP's that attempted gapping in prelim test: 7241
Number of HSP's gapped (non-prelim): 25
length of query: 587
length of database: 59,974,054
effective HSP length: 116
effective length of query: 471
effective length of database: 40,926,738
effective search space: 19276493598
effective search space used: 19276493598
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 69 (31.2 bits)
Medicago: description of AC144539.4


