

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144539.1 + phase: 0
(163 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
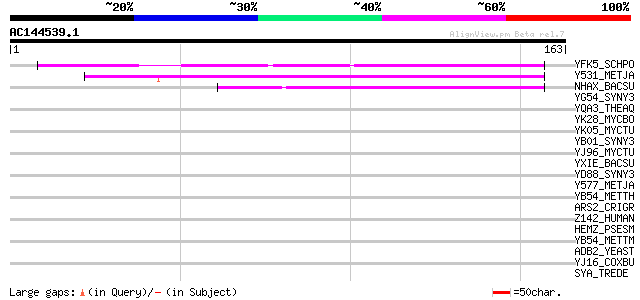
Score E
Sequences producing significant alignments: (bits) Value
YFK5_SCHPO (P87132) Hypothetical protein C167.05 in chromosome I 52 5e-07
Y531_METJA (Q57951) Hypothetical protein MJ0531 47 1e-05
NHAX_BACSU (O07552) Stress response protein nhaX 43 4e-04
YG54_SYNY3 (P72817) Hypothetical protein sll1654 41 0.001
YQA3_THEAQ (P74897) Hypothetical 14.6 kDa protein in QAH/OAS sul... 40 0.002
YK28_MYCBO (P64922) Hypothetical protein Mb2028c 38 0.009
YK05_MYCTU (P64921) Hypothetical protein Rv2005c/MT2061 38 0.009
YB01_SYNY3 (P72745) Hypothetical protein slr1101 37 0.015
YJ96_MYCTU (Q10862) Hypothetical protein Rv1996/MT2052/Mb2019 37 0.020
YXIE_BACSU (P42297) Hypothetical protein yxiE precursor 36 0.034
YD88_SYNY3 (P74148) Hypothetical protein sll1388 36 0.044
Y577_METJA (Q57997) Protein MJ0577 35 0.098
YB54_METTH (O27222) Hypothetical protein MTH1154 33 0.28
ARS2_CRIGR (Q60436) Arsenite-resistance protein 2 (Fragment) 31 1.4
Z142_HUMAN (P52746) Zinc finger protein 142 (HA4654) 30 1.8
HEMZ_PSESM (Q888A2) Ferrochelatase (EC 4.99.1.1) (Protoheme ferr... 30 2.4
YB54_METTM (Q50777) Hypothetical 16.1 kDa protein in MTR region ... 30 3.1
ADB2_YEAST (P36000) Probable beta-adaptin (Clathrin assembly pro... 29 4.1
YJ16_COXBU (P45680) Hypothetical protein CBU1916 29 5.4
SYA_TREDE (P61708) Alanyl-tRNA synthetase (EC 6.1.1.7) (Alanine-... 29 5.4
>YFK5_SCHPO (P87132) Hypothetical protein C167.05 in chromosome I
Length = 601
Score = 52.4 bits (124), Expect = 5e-07
Identities = 37/149 (24%), Positives = 73/149 (48%), Gaps = 14/149 (9%)
Query: 9 IAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLGEFIN 68
+ +D S S+ A +W V +++ GD LI++ + E + + +
Sbjct: 435 LTLDLSSESLHAAEWAVGILLRNGDTLIIV------------DVIECDDPSARAVKDRME 482
Query: 69 SDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLTMGNR 128
S+ + E K +LK+ + + + +V + V ++ A+ + E I+ + + MG+R
Sbjct: 483 SEQLETLE-KITKYILKLLSKTVLEVEVNIEV-IHHEKAKHLIIEMIDYIEPSLVVMGSR 540
Query: 129 GLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
G L+ ++GS SNY+VN +S PV V +
Sbjct: 541 GRSHLKGVLLGSFSNYLVNKSSVPVMVAR 569
>Y531_METJA (Q57951) Hypothetical protein MJ0531
Length = 170
Score = 47.4 bits (111), Expect = 1e-05
Identities = 38/153 (24%), Positives = 67/153 (42%), Gaps = 18/153 (11%)
Query: 23 WTVDNIVKEGDNLILIIIRP------------------EEYEHGEMQLWEVTGSPLTPLG 64
+ +++I++ G+NL I+ P +E++ ++ V SP L
Sbjct: 13 YILNSIIRNGENLYKKIVIPTDGSDVSLEAAKHAINIAKEFDAEVYAIYVVDVSPFVGLP 72
Query: 65 EFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLT 124
+ +L + + E LK E+ V + ++ G ++ E E+ D +
Sbjct: 73 AEGSWELISELLKEEGQEALKKVKKMAEEWGVKIHTEMLEGVPANEIVEFAEKKKADLIV 132
Query: 125 MGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
MG G L R ++GSV+ V+ NA CPV VVK
Sbjct: 133 MGTTGKTGLERILLGSVAERVIKNAHCPVLVVK 165
>NHAX_BACSU (O07552) Stress response protein nhaX
Length = 166
Score = 42.7 bits (99), Expect = 4e-04
Identities = 27/96 (28%), Positives = 47/96 (48%), Gaps = 1/96 (1%)
Query: 62 PLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLD 121
PL + S P YE +T+ EV+ A + +++ + + GD E + E ++ D
Sbjct: 72 PLISDVTSPEPMIYEDRTE-EVIAEARMMLNEQQADGDIDILEGDPAESIIEHANRISAD 130
Query: 122 GLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
+ G+R L++ I GSVS + + PV +VK
Sbjct: 131 MIVTGSRDQNRLKKLIFGSVSEKLSAKSDIPVLIVK 166
>YG54_SYNY3 (P72817) Hypothetical protein sll1654
Length = 157
Score = 41.2 bits (95), Expect = 0.001
Identities = 23/75 (30%), Positives = 37/75 (48%)
Query: 82 EVLKIATTAIEQKKVVVLVKVYWGDAREKLCEAIEQVPLDGLTMGNRGLGTLRRAIMGSV 141
++L+ A Q+ + G A +C+ ++V D + MG RGLG + SV
Sbjct: 82 KLLEAAQAVFSQQGIATKTIEREGMASFTICDVADEVNADLIVMGCRGLGLTTEGVAESV 141
Query: 142 SNYVVNNASCPVTVV 156
+ V+N + CPV VV
Sbjct: 142 TARVINLSPCPVLVV 156
>YQA3_THEAQ (P74897) Hypothetical 14.6 kDa protein in QAH/OAS
sulfhydrylase 3'region
Length = 137
Score = 40.0 bits (92), Expect = 0.002
Identities = 34/121 (28%), Positives = 52/121 (42%), Gaps = 6/121 (4%)
Query: 39 IIRPEEYEHGEMQLWEVTGSPLTP-LGE-FINSDLPKKYEIKTDPEVLKIATTAIEQKKV 96
+ + E HG L P+ LGE F L ++ E +A T + ++
Sbjct: 21 VAKAEAQAHGARLLVVHAYEPVPDYLGEPFFEEALKRRLERAEKVRAEAMALTGVPREDA 80
Query: 97 VVLVKVYWGDAREKLCEAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVV 156
++L G E + +A D + MG RGLG + +GS S VV A CPV +V
Sbjct: 81 LLLQ----GRPAEAILQAAIGEKADLIVMGTRGLGAVGSLFLGSQSQKVVAEAPCPVLLV 136
Query: 157 K 157
+
Sbjct: 137 R 137
>YK28_MYCBO (P64922) Hypothetical protein Mb2028c
Length = 295
Score = 38.1 bits (87), Expect = 0.009
Identities = 16/36 (44%), Positives = 24/36 (66%)
Query: 123 LTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKS 158
+ +G+ G G L R ++GSVS+ +V A CPV V+ S
Sbjct: 114 VVLGSSGRGALARGLLGSVSSSLVRRAGCPVAVIHS 149
Score = 31.2 bits (69), Expect = 1.1
Identities = 15/37 (40%), Positives = 23/37 (61%)
Query: 123 LTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSS 159
+ +G+ G G L ++GSVSN V++ A PV V + S
Sbjct: 259 VVVGSHGRGGLTGMLLGSVSNAVLHAARVPVIVARQS 295
>YK05_MYCTU (P64921) Hypothetical protein Rv2005c/MT2061
Length = 295
Score = 38.1 bits (87), Expect = 0.009
Identities = 16/36 (44%), Positives = 24/36 (66%)
Query: 123 LTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKS 158
+ +G+ G G L R ++GSVS+ +V A CPV V+ S
Sbjct: 114 VVLGSSGRGALARGLLGSVSSSLVRRAGCPVAVIHS 149
Score = 31.2 bits (69), Expect = 1.1
Identities = 15/37 (40%), Positives = 23/37 (61%)
Query: 123 LTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKSS 159
+ +G+ G G L ++GSVSN V++ A PV V + S
Sbjct: 259 VVVGSHGRGGLTGMLLGSVSNAVLHAARVPVIVARQS 295
>YB01_SYNY3 (P72745) Hypothetical protein slr1101
Length = 108
Score = 37.4 bits (85), Expect = 0.015
Identities = 19/52 (36%), Positives = 30/52 (57%)
Query: 105 GDAREKLCEAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVV 156
G + +C+ +Q D + +G+RG L ++GSV NYV ++A C V VV
Sbjct: 53 GSPGKIICQRAKQDNSDIIVVGHRGRWGLSEILLGSVGNYVFHHAHCCVFVV 104
>YJ96_MYCTU (Q10862) Hypothetical protein Rv1996/MT2052/Mb2019
Length = 317
Score = 37.0 bits (84), Expect = 0.020
Identities = 34/149 (22%), Positives = 60/149 (39%), Gaps = 3/149 (2%)
Query: 9 IAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLGEFIN 68
+ +D SPCS A +W + L ++ + P E +E + E +
Sbjct: 12 VGVDGSPCSHTAVEWAARDAQMRNVALRVVQVVPPVITAPEGWAFEYSRFQEAQKREIVE 71
Query: 69 SDLPKKYEIKTDPEVLKIATTAIEQKKVVVLV-KVYWGDAREKLCEAIEQVPLDGLTMGN 127
+ + K+A A + + +V G L QV + + +G
Sbjct: 72 HSYLVAQAHQIVEQAHKVALEASSSGRAAQITGEVLHGQIVPTLANISRQVAM--VVLGY 129
Query: 128 RGLGTLRRAIMGSVSNYVVNNASCPVTVV 156
RG G + A++GSVS+ +V +A PV V+
Sbjct: 130 RGQGAVAGALLGSVSSSLVRHAHGPVAVI 158
>YXIE_BACSU (P42297) Hypothetical protein yxiE precursor
Length = 148
Score = 36.2 bits (82), Expect = 0.034
Identities = 37/152 (24%), Positives = 70/152 (45%), Gaps = 13/152 (8%)
Query: 9 IAMDFSPCSIKAFQWTVDNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLGEFIN 68
+A+D S S KA V ++ L ++ + E VT S LT + ++
Sbjct: 7 VAIDGSDMSAKALDAAVHLAKEQQAELSILHVGREAV---------VTTSSLTGI-VYVP 56
Query: 69 SDLPKKYEIKTDPEVLKIATTAIEQ--KKVVVLVKVYW-GDAREKLCEAIEQVPLDGLTM 125
+ + E LKI A E+ +K V +Y G+ ++ ++ + + +
Sbjct: 57 EHFIDEIRNEVKKEGLKILENAKEKAAEKGVQAETIYANGEPAHEILNHAKEKGVSLIVV 116
Query: 126 GNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
G+RG+ L+ ++GSVS+ V ++CPV +V+
Sbjct: 117 GSRGISGLKEMMLGSVSHKVSQLSTCPVLIVR 148
>YD88_SYNY3 (P74148) Hypothetical protein sll1388
Length = 154
Score = 35.8 bits (81), Expect = 0.044
Identities = 15/37 (40%), Positives = 24/37 (64%)
Query: 121 DGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
D + +G RGL L +GSVS+YV+++ C V +V+
Sbjct: 117 DLVVLGRRGLKGLAEVFLGSVSSYVIHHVQCSVLIVQ 153
>Y577_METJA (Q57997) Protein MJ0577
Length = 162
Score = 34.7 bits (78), Expect = 0.098
Identities = 17/53 (32%), Positives = 30/53 (56%)
Query: 105 GDAREKLCEAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVK 157
G E++ + E +D + MG+ G L+ ++GSV+ V+ ++ PV VVK
Sbjct: 106 GIPHEEIVKIAEDEGVDIIIMGSHGKTNLKEILLGSVTENVIKKSNKPVLVVK 158
>YB54_METTH (O27222) Hypothetical protein MTH1154
Length = 146
Score = 33.1 bits (74), Expect = 0.28
Identities = 27/105 (25%), Positives = 50/105 (46%), Gaps = 14/105 (13%)
Query: 54 EVTGSPLTPLGEFINSDLPKKYEIKTDPEVLKIATTAIEQKKVVVLVKVYWGDAREKLCE 113
E+ LT G I D+ K T PE ++ A+ ++ GD +++ +
Sbjct: 55 EMMVKELTQRGNEILRDMEKGL---TGPENPNVSFRAVMRE----------GDPADEIVK 101
Query: 114 AIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKS 158
E+ +D + MG G + + ++GSVS VV+ A C + +V++
Sbjct: 102 VAEEEDVDVIVMGT-GKSLVDKHLLGSVSEKVVHYAPCTIHLVRT 145
>ARS2_CRIGR (Q60436) Arsenite-resistance protein 2 (Fragment)
Length = 290
Score = 30.8 bits (68), Expect = 1.4
Identities = 19/56 (33%), Positives = 29/56 (50%), Gaps = 4/56 (7%)
Query: 42 PEEYEHGEMQLWEVT-GSPLTPLGEFINSDL---PKKYEIKTDPEVLKIATTAIEQ 93
P HGE+ W+ T LTPL S L P+++ +KT+ EV K T+ ++
Sbjct: 81 PNRISHGEVLEWQKTFEEKLTPLLSVRESFLRKRPRRWVVKTEQEVEKFVTSNTQE 136
>Z142_HUMAN (P52746) Zinc finger protein 142 (HA4654)
Length = 1687
Score = 30.4 bits (67), Expect = 1.8
Identities = 14/34 (41%), Positives = 20/34 (58%)
Query: 42 PEEYEHGEMQLWEVTGSPLTPLGEFINSDLPKKY 75
P E E E L V+GSP+ P G + ++ PKK+
Sbjct: 946 PREPEETEEPLATVSGSPVPPAGNSLPTEAPKKH 979
>HEMZ_PSESM (Q888A2) Ferrochelatase (EC 4.99.1.1) (Protoheme
ferro-lyase) (Heme synthetase)
Length = 340
Score = 30.0 bits (66), Expect = 2.4
Identities = 23/91 (25%), Positives = 39/91 (42%), Gaps = 9/91 (9%)
Query: 36 ILIIIRPEEYEHGEMQLWEVTGSPLTPLGEFINSDLPKKYEIKT--------DPEVLKIA 87
+++I RPE+ H +W GSPL L + + + K++ +P + +
Sbjct: 49 LILIKRPEQSAHAYASIWWDEGSPLVVLSKRLQQAMKKEWSHGPVELAMRYGEPSIETVL 108
Query: 88 TTAIEQK-KVVVLVKVYWGDAREKLCEAIEQ 117
T EQ K V L +Y A + IE+
Sbjct: 109 TRLSEQGFKKVTLAPLYPQFADSTVTTVIEE 139
>YB54_METTM (Q50777) Hypothetical 16.1 kDa protein in MTR region
(ORF143)
Length = 143
Score = 29.6 bits (65), Expect = 3.1
Identities = 15/54 (27%), Positives = 32/54 (58%), Gaps = 1/54 (1%)
Query: 105 GDAREKLCEAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKS 158
G+ +++ + E+ +D + MG G + + ++GSVS VV+ A C + +V++
Sbjct: 90 GNPADEIVKLAEEEDVDVIIMGT-GKSLVDKHLLGSVSEKVVHYAPCTIHLVRT 142
>ADB2_YEAST (P36000) Probable beta-adaptin (Clathrin assembly
protein large beta chain) (Clathrin assembly protein
complex 2 beta large chain)
Length = 726
Score = 29.3 bits (64), Expect = 4.1
Identities = 15/33 (45%), Positives = 22/33 (66%), Gaps = 1/33 (3%)
Query: 81 PEVLK-IATTAIEQKKVVVLVKVYWGDAREKLC 112
P+VLK IAT +EQKK+V L + + + +LC
Sbjct: 65 PDVLKNIATIDVEQKKLVYLYVMNYAETHPELC 97
>YJ16_COXBU (P45680) Hypothetical protein CBU1916
Length = 146
Score = 28.9 bits (63), Expect = 5.4
Identities = 19/62 (30%), Positives = 34/62 (54%), Gaps = 3/62 (4%)
Query: 99 LVKVYWGDAREKLCEAIEQVPLDGLTMGNRGLGTLRRAIMGSVSNYVVNNASCPVTVVKS 158
+VKV G A+ + E + +D + +G+ G ++ ++GS SN V++ A C V V+
Sbjct: 87 IVKV--GPAKFLILEQAKNWGVDLIIVGSHGRHGIQ-LLLGSTSNAVLHGAKCDVLAVRI 143
Query: 159 SG 160
G
Sbjct: 144 KG 145
>SYA_TREDE (P61708) Alanyl-tRNA synthetase (EC 6.1.1.7)
(Alanine--tRNA ligase) (AlaRS)
Length = 600
Score = 28.9 bits (63), Expect = 5.4
Identities = 24/88 (27%), Positives = 40/88 (45%), Gaps = 2/88 (2%)
Query: 26 DNIVKEGDNLILIIIRPEEYEHGEMQLWEVTGSPLTPLGEFINSDLPKKYEIKTDPEVLK 85
D++ ++G N+ +R ++ H E + E I +DLP E+ E K
Sbjct: 479 DHVQQKGSNITAERLR-FDFSHPEPMTEAEKKEVERLVNEAIKADLPVTMEVMPLEEAKK 537
Query: 86 IATTAIEQKKVVVLVKVY-WGDAREKLC 112
I A+ +K +VKVY GD ++C
Sbjct: 538 IGAMALFGEKYEDVVKVYKIGDFSTEVC 565
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.135 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,885,862
Number of Sequences: 164201
Number of extensions: 767319
Number of successful extensions: 1919
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 15
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 1899
Number of HSP's gapped (non-prelim): 29
length of query: 163
length of database: 59,974,054
effective HSP length: 102
effective length of query: 61
effective length of database: 43,225,552
effective search space: 2636758672
effective search space used: 2636758672
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144539.1


