

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144538.7 + phase: 1 /pseudo
(632 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
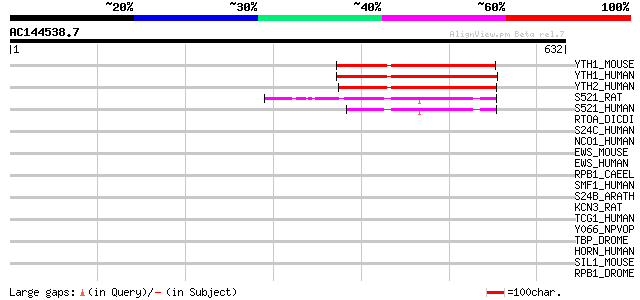
Score E
Sequences producing significant alignments: (bits) Value
YTH1_MOUSE (P59326) YTH domain protein 1 (Dermatomyositis associ... 199 1e-50
YTH1_HUMAN (Q9BYJ9) YTH domain protein 1 (Dermatomyositis associ... 197 5e-50
YTH2_HUMAN (Q9Y5A9) YTH domain protein 2 (High-glucose-regulated... 197 9e-50
S521_RAT (Q9QY02) Putative splicing factor YT521 (RA301-binding ... 75 5e-13
S521_HUMAN (Q96MU7) Putative splicing factor YT521 75 5e-13
RTOA_DICDI (P54681) Protein rtoA (Ratio-A) 43 0.002
S24C_HUMAN (P53992) Protein transport protein Sec24C (SEC24-rela... 41 0.008
NCO1_HUMAN (Q15788) Nuclear receptor coactivator 1 (EC 2.3.1.48)... 40 0.019
EWS_MOUSE (Q61545) RNA-binding protein EWS 40 0.019
EWS_HUMAN (Q01844) RNA-binding protein EWS (EWS oncogene) (Ewing... 40 0.024
RPB1_CAEEL (P16356) DNA-directed RNA polymerase II largest subun... 39 0.032
SMF1_HUMAN (O14497) SWI/SNF-related, matrix-associated, actin-de... 39 0.041
S24B_ARATH (Q9M081) Putative protein transport protein Sec24-lik... 39 0.041
KCN3_RAT (P70605) Small conductance calcium-activated potassium ... 38 0.071
TCG1_HUMAN (O14776) Transcription elongation regulator 1 (TATA b... 38 0.092
Y066_NPVOP (Q83949) Hypothetical 98.6 kDa protein (ORF71) 37 0.16
TBP_DROME (P20227) TATA-box binding protein (TATA-box factor) (T... 37 0.21
HORN_HUMAN (Q86YZ3) Hornerin 37 0.21
SIL1_MOUSE (Q8C0T5) Signal-induced proliferation-associated 1 li... 36 0.27
RPB1_DROME (P04052) DNA-directed RNA polymerase II largest subun... 36 0.27
>YTH1_MOUSE (P59326) YTH domain protein 1 (Dermatomyositis
associated with cancer putative autoantigen-1 homolog)
(DACA-1 homolog)
Length = 559
Score = 199 bits (507), Expect = 1e-50
Identities = 97/182 (53%), Positives = 124/182 (67%), Gaps = 4/182 (2%)
Query: 373 SLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEKQDAC 432
S N +F + K + F+IKSYSED++H+SIKY +W ST +GN++LD A+ K
Sbjct: 376 SYNPKEFDWNLKSGRVFIIKSYSEDDIHRSIKYSIWCSTEHGNKRLDGAFRSMSSKGP-- 433
Query: 433 RIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHI 492
++L FSVN S FCGVAEM PV++ S W QDKW G+F VKW +KDVPN+Q RHI
Sbjct: 434 -VYLLFSVNGSGHFCGVAEMKSPVDYGTSAGVWSQDKWKGKFDVKWIFVKDVPNNQLRHI 492
Query: 493 VLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQ-ERKA 551
LENNDNKPVTNSRDTQEV L++ ++L I +Y+ TSI DDF YE RQ+ + RK
Sbjct: 493 RLENNDNKPVTNSRDTQEVPLEKAKQVLKIIASYKHTTSIFDDFSHYEKRQEEEEVVRKE 552
Query: 552 RQ 553
RQ
Sbjct: 553 RQ 554
>YTH1_HUMAN (Q9BYJ9) YTH domain protein 1 (Dermatomyositis
associated with cancer putative autoantigen-1) (DACA-1)
Length = 559
Score = 197 bits (502), Expect = 5e-50
Identities = 94/183 (51%), Positives = 125/183 (67%), Gaps = 3/183 (1%)
Query: 373 SLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEKQDAC 432
S N +F + K + F+IKSYSED++H+SIKY +W ST +GN++LD+A+ K
Sbjct: 376 SYNPKEFEWNLKSGRVFIIKSYSEDDIHRSIKYSIWCSTEHGNKRLDSAFRCMSSKGP-- 433
Query: 433 RIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHI 492
++L FSVN S FCGVAEM PV++ S W QDKW G+F V+W +KDVPN+Q RHI
Sbjct: 434 -VYLLFSVNGSGHFCGVAEMKSPVDYGTSAGVWSQDKWKGKFDVQWIFVKDVPNNQLRHI 492
Query: 493 VLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKAR 552
LENNDNKPVTNSRDTQEV L++ ++L I +Y+ TSI DDF YE RQ+ + +
Sbjct: 493 RLENNDNKPVTNSRDTQEVPLEKAKQVLKIISSYKHTTSIFDDFAHYEKRQEEEEVVRKE 552
Query: 553 QQS 555
+QS
Sbjct: 553 RQS 555
>YTH2_HUMAN (Q9Y5A9) YTH domain protein 2 (High-glucose-regulated
protein 8) (NY-REN-2 antigen) (CLL-associated antigen
KW-14)
Length = 579
Score = 197 bits (500), Expect = 9e-50
Identities = 94/180 (52%), Positives = 122/180 (67%), Gaps = 3/180 (1%)
Query: 375 NRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEKQDACRI 434
N DF + K + F+IKSYSED++H+SIKY +W ST +GN++LDAAY K +
Sbjct: 399 NPKDFDWNLKHGRVFIIKSYSEDDIHRSIKYNIWCSTEHGNKRLDAAYRSMNGKGP---V 455
Query: 435 FLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVL 494
+L FSVN S FCGVAEM V+++ W QDKW G+F V+W +KDVPNSQ RHI L
Sbjct: 456 YLLFSVNGSGHFCGVAEMKSAVDYNTCAGVWSQDKWKGRFDVRWIFVKDVPNSQLRHIRL 515
Query: 495 ENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKARQQ 554
ENN+NKPVTNSRDTQEV L++ ++L I +Y+ TSI DDF YE RQ+ + K +Q
Sbjct: 516 ENNENKPVTNSRDTQEVPLEKAKQVLKIIASYKHTTSIFDDFSHYEKRQEEEESVKKERQ 575
>S521_RAT (Q9QY02) Putative splicing factor YT521 (RA301-binding
protein)
Length = 738
Score = 75.1 bits (183), Expect = 5e-13
Identities = 68/270 (25%), Positives = 118/270 (43%), Gaps = 30/270 (11%)
Query: 291 SSFGGSAISGLSANDRSFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKNHI 350
S G ++S + RS + S +E R G I+ +++ G AS+ ++
Sbjct: 274 SDSGSESVSFTDGSVRSGSGTDGSDEKKKERK---RARGISPIVFDRS-GSSASE--SYA 327
Query: 351 SSENNAIDGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWAS 410
SE S + + K Q L + +DA+FF+IKS + +NV + GVW++
Sbjct: 328 GSEKKHEKLSSSVRAVRKDQTSKLK-----SVLQDARFFLIKSNNHENVSLAKAKGVWST 382
Query: 411 TPNGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFW----- 465
P +KL+ A+ A+ + L FSV S +F G A + + S W
Sbjct: 383 LPVNEKKLNLAFRSARS------VILIFSVRESGKFQGFARLSSESHHGGSPIHWVLPAG 436
Query: 466 -QQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFK 524
G F + W +++P ++ H+ N++KPV RD QE++L+ G ++ +F
Sbjct: 437 MSAKMLGGVFKIDWICRRELPFTKSAHLTNPWNEHKPVKIGRDGQEIELECGTQLCLLFP 496
Query: 525 NYETDTSILDDFDFYEDRQKAMQERKARQQ 554
E+ D Y+ K +R+ Q
Sbjct: 497 PDES-------IDLYQLIHKMRHKRRMHSQ 519
>S521_HUMAN (Q96MU7) Putative splicing factor YT521
Length = 727
Score = 75.1 bits (183), Expect = 5e-13
Identities = 49/177 (27%), Positives = 84/177 (46%), Gaps = 19/177 (10%)
Query: 384 KDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNAS 443
+DA+FF+IKS + +NV + GVW++ P +KL+ A+ A+ + L FSV S
Sbjct: 353 QDARFFLIKSNNHENVSLAKAKGVWSTLPVNEKKLNLAFRSARS------VILIFSVRES 406
Query: 444 AQFCGVAEMVGPVNFDKSVDFW------QQDKWSGQFPVKWHIIKDVPNSQFRHIVLENN 497
+F G A + + S W G F + W +++P ++ H+ N
Sbjct: 407 GKFQGFARLSSESHHGGSPIHWVLPAGMSAKMLGGVFKIDWICRRELPFTKSAHLTNPWN 466
Query: 498 DNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKARQQ 554
++KPV RD QE++L+ G ++ +F E+ D Y+ K +R+ Q
Sbjct: 467 EHKPVKIGRDGQEIELECGTQLCLLFPPDES-------IDLYQVIHKMRHKRRMHSQ 516
>RTOA_DICDI (P54681) Protein rtoA (Ratio-A)
Length = 400
Score = 43.1 bits (100), Expect = 0.002
Identities = 47/165 (28%), Positives = 71/165 (42%), Gaps = 10/165 (6%)
Query: 209 SFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSAGMASHQQ 268
S + QG GS + G S + + +S S S Q GS S S+G ++
Sbjct: 138 SSNSGSQGSTGSSNSGSQSSTDSSNSGSQSSTDSSNSGSQGSTGSSNSGSESSGSSNSGS 197
Query: 269 RSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSFLSLENSRRHGRETASFCRCN 328
S GSS++ G S+ GSS GS S S+N S S S G E++S +
Sbjct: 198 ESS---GSSNS----GSESSSGSSNSGSESSSGSSNSGS-ESSSGSSNSGSESSSGSSNS 249
Query: 329 GTLDILSEQNRGPRASKLKNHISSENNAIDGSKNNASTAKFQDES 373
G+ N G +S ++ SE+++ GS N+ S + S
Sbjct: 250 GSESSSGSSNSGSESSSGSSNSGSESSS--GSSNSGSESSGSSNS 292
Score = 40.0 bits (92), Expect = 0.019
Identities = 52/201 (25%), Positives = 79/201 (38%), Gaps = 12/201 (5%)
Query: 175 IDQQVESNFFGPRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSE 234
+ +V+S+ S + S S GS S S GS + G S S+ SE
Sbjct: 64 LSSEVDSSDISSSGSNSTASSEGSVSSSSNSGSQSTSNSGSEASGSSNSGSQSTSNSGSE 123
Query: 235 RQ-------RSFMPLSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLS 287
+S S S Q GS S S+ +S+ S+S G S
Sbjct: 124 ASGSSNSGSQSSTDSSNSGSQGSTGSSNSGSQSSTDSSNSGSQSSTDSSNSGSQGSTGSS 183
Query: 288 NPGSSFGGSAISGLSANDRSFLSLEN---SRRHGRETASFCRCNGTLDILSEQNRGPRAS 344
N GS GS+ SG ++ S E+ S G E++S +G+ N G +S
Sbjct: 184 NSGSESSGSSNSGSESSGSSNSGSESSSGSSNSGSESSSGSSNSGSESSSGSSNSGSESS 243
Query: 345 KLKNHISSENNAIDGSKNNAS 365
++ SE+++ GS N+ S
Sbjct: 244 SGSSNSGSESSS--GSSNSGS 262
Score = 35.8 bits (81), Expect = 0.35
Identities = 47/157 (29%), Positives = 61/157 (37%), Gaps = 9/157 (5%)
Query: 174 GIDQQVESNFFGPRASYPSVGSFARGSFPVA-PGSFSFHESQQGFDGSRSGGLWSDSSKP 232
G +S+ G ++S S S ++GS + GS S S G + S S S+SS
Sbjct: 153 GSQSSTDSSNSGSQSSTDSSNSGSQGSTGSSNSGSESSGSSNSGSESSGSSNSGSESSSG 212
Query: 233 SERQ--RSFMPLSPSVPQQPIGSVRSFGPSA------GMASHQQRSLYGFGSSSNPYGRG 284
S S S S + GS S S+ G S S G SSS G
Sbjct: 213 SSNSGSESSSGSSNSGSESSSGSSNSGSESSSGSSNSGSESSSGSSNSGSESSSGSSNSG 272
Query: 285 YLSNPGSSFGGSAISGLSANDRSFLSLENSRRHGRET 321
S+ GSS GS SG S + S S G+ T
Sbjct: 273 SESSSGSSNSGSESSGSSNSGSESSSDSGSSSDGKTT 309
Score = 33.1 bits (74), Expect = 2.3
Identities = 28/112 (25%), Positives = 49/112 (43%), Gaps = 3/112 (2%)
Query: 265 SHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSFLSLENSRRHGRETASF 324
S+ S G SSS+ G SN GS GS+ SG + S S G ++++
Sbjct: 78 SNSTASSEGSVSSSSNSGSQSTSNSGSEASGSSNSGSQSTSNSGSEASGSSNSGSQSSTD 137
Query: 325 CRCNGTLDILSEQNRGPRASKLKNHI---SSENNAIDGSKNNASTAKFQDES 373
+G+ N G ++S ++ SS +++ GS+ + ++ ES
Sbjct: 138 SSNSGSQGSTGSSNSGSQSSTDSSNSGSQSSTDSSNSGSQGSTGSSNSGSES 189
>S24C_HUMAN (P53992) Protein transport protein Sec24C (SEC24-related
protein C)
Length = 1094
Score = 41.2 bits (95), Expect = 0.008
Identities = 54/185 (29%), Positives = 73/185 (39%), Gaps = 18/185 (9%)
Query: 35 TSQSGSFGSGADQPLYPPNVYA---PQAQAFYYGGFDNGNGEWDEYPSYVNNGGIEI-GS 90
T SG SGA P P + P + A G F N +G + YP + G
Sbjct: 138 TQLSGMQISGAVAPAPPSSGLGFGPPTSLASASGSFPN-SGLYGSYPQGQAPPLSQAQGH 196
Query: 91 PGVYNENQSLVFHSGYGFNPQMPYGPYSP-VTSPL--------PSVGGDAQLYSPQQFPY 141
PG+ +S + F P GP P +T PL PSV + SP Q
Sbjct: 197 PGIQTPQRSAPSQAS-SFTPPASGGPRLPSMTGPLLPGQSFGGPSVSQPNHVSSPPQA-- 253
Query: 142 TPPYYNQLVPPPNLPYLNSPT-PVSQPELTNLLGIDQQVESNFFGPRASYPSVGSFARGS 200
PP P LP ++SP P QP+ G + +SN+ GP + P+ GS
Sbjct: 254 LPPGTQMTGPLGPLPPMHSPQQPGYQPQQNGSFGPARGPQSNYGGPYPAAPTFGSQPGPP 313
Query: 201 FPVAP 205
P+ P
Sbjct: 314 QPLPP 318
>NCO1_HUMAN (Q15788) Nuclear receptor coactivator 1 (EC 2.3.1.48)
(NCoA-1) (Steroid receptor coactivator-1) (SRC-1)
(RIP160) (Hin-2 protein)
Length = 1441
Score = 40.0 bits (92), Expect = 0.019
Identities = 34/115 (29%), Positives = 48/115 (41%), Gaps = 24/115 (20%)
Query: 111 QMPYGPYSPVTSPLPSVGGDAQLY--SPQQFPYTPPY-------------YNQLVPPPNL 155
Q +G P +S LP G+ +L +PQQFPY P Y ++QL P
Sbjct: 1137 QQSFGNNLPPSSGLPVQTGNPRLPQGAPQQFPYPPNYGTNPGTPPASTSPFSQLAANPEA 1196
Query: 156 PYLNSPTPVSQPELTNL-----LGIDQQVESNFFGPRASYPSVGSFARGSFPVAP 205
N + VS+ N+ GI+ Q++ N F YP G +G AP
Sbjct: 1197 SLANRNSMVSRGMTGNIGGQFGTGINPQMQQNVF----QYPGAGMVPQGEANFAP 1247
>EWS_MOUSE (Q61545) RNA-binding protein EWS
Length = 655
Score = 40.0 bits (92), Expect = 0.019
Identities = 75/323 (23%), Positives = 110/323 (33%), Gaps = 71/323 (21%)
Query: 23 QGTTTIGHSRDIT----SQSGSFGSGADQPLY--PPNVY----APQAQAFYYGGFDNGNG 72
Q T G D++ + ++G A Y PP Y APQA + G+ G G
Sbjct: 39 QSYGTYGQPTDVSYTQAQTTATYGQTAYATSYGQPPTGYSTPTAPQAYSQPVQGY--GTG 96
Query: 73 EWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMP-YGPYSPVTSPL------- 124
+D + V S S YG P P YG T+P
Sbjct: 97 AYDSTTATVTT------------TQASYAAQSAYGTQPAYPTYGQQPTATAPTRPQDGNK 144
Query: 125 ----------------PSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPE 168
PS+G YS Q P + P PP P S + + +
Sbjct: 145 PAETSQPQSSTGGYNQPSLGYGQSNYSYPQVPGSYPMQPVTAPPSYPPTSYSSSQPTSYD 204
Query: 169 LTNLLGIDQQVESNFFGPRASYPSVGSFARG---SFPVAPGSFSFHESQQGFDGSRSGGL 225
++ + + + +G ++SY S+ + S+P GS+S SQ
Sbjct: 205 QSSYSQQNTYGQPSSYGQQSSYGQQSSYGQQPPTSYPPQTGSYSQAPSQ----------- 253
Query: 226 WSDSSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGY 285
+S S +Q SF PS S+ +G +G S + G + GRG
Sbjct: 254 YSQQSSSYGQQSSFRQDHPS-------SMGVYGQESGGFSGPGENRSLSGPDNRGRGRGG 306
Query: 286 LSNPGSSFG--GSAISGLSANDR 306
G S G G GL A +R
Sbjct: 307 FDRGGMSRGGRGGGRGGLGAGER 329
Score = 33.1 bits (74), Expect = 2.3
Identities = 42/197 (21%), Positives = 78/197 (39%), Gaps = 18/197 (9%)
Query: 184 FGPRASYPSVGSFARGSFPVAP--GSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMP 241
+G + +YP+ G + P P G+ SQ S +GG S + S+
Sbjct: 118 YGTQPAYPTYGQQPTATAPTRPQDGNKPAETSQPQ---SSTGGYNQPSLGYGQSNYSYPQ 174
Query: 242 LSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGL 301
+ S P QP+ + S+ P++ +S S N YG+ SS+G
Sbjct: 175 VPGSYPMQPVTAPPSYPPTSYSSSQPTSYDQSSYSQQNTYGQPSSYGQQSSYG------- 227
Query: 302 SANDRSFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKNHISSENNAIDGSK 361
+S + + +T S+ + S + G ++S ++H SS + G +
Sbjct: 228 ---QQSSYGQQPPTSYPPQTGSYSQAPSQYSQQS-SSYGQQSSFRQDHPSS--MGVYGQE 281
Query: 362 NNASTAKFQDESLNRPD 378
+ + ++ SL+ PD
Sbjct: 282 SGGFSGPGENRSLSGPD 298
>EWS_HUMAN (Q01844) RNA-binding protein EWS (EWS oncogene) (Ewing
sarcoma breakpoint region 1 protein)
Length = 656
Score = 39.7 bits (91), Expect = 0.024
Identities = 72/310 (23%), Positives = 105/310 (33%), Gaps = 69/310 (22%)
Query: 23 QGTTTIGHSRDIT----SQSGSFGSGADQPLY--PPNVY----APQAQAFYYGGFDNGNG 72
Q T G D++ + ++G A Y PP Y APQA + G+ G G
Sbjct: 39 QSYGTYGQPTDVSYTQAQTTATYGQTAYATSYGQPPTGYTTPTAPQAYSQPVQGY--GTG 96
Query: 73 EWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMP-YGPYSPVTSPL------- 124
+D + V S S YG P P YG T+P
Sbjct: 97 AYDTTTATVTT------------TQASYAAQSAYGTQPAYPAYGQQPAATAPTRPQDGNK 144
Query: 125 ----------------PSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPE 168
PS+G YS Q P + P PP P S T + +
Sbjct: 145 PTETSQPQSSTGGYNQPSLGYGQSNYSYPQVPGSYPMQPVTAPPSYPPTSYSSTQPTSYD 204
Query: 169 LTNLLGIDQQVESNFFGPRASYPSVGSFAR---GSFPVAPGSFSFHESQQGFDGSRSGGL 225
++ + + + +G ++SY S+ + S+P GS+S SQ
Sbjct: 205 QSSYSQQNTYGQPSSYGQQSSYGQQSSYGQQPPTSYPPQTGSYSQAPSQ----------- 253
Query: 226 WSDSSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGY 285
+S S +Q SF PS S+ +G +G S + G + GRG
Sbjct: 254 YSQQSSSYGQQSSFRQDHPS-------SMGVYGQESGGFSGPGENRSMSGPDNRGRGRGG 306
Query: 286 LSNPGSSFGG 295
G S GG
Sbjct: 307 FDRGGMSRGG 316
>RPB1_CAEEL (P16356) DNA-directed RNA polymerase II largest subunit
(EC 2.7.7.6)
Length = 1852
Score = 39.3 bits (90), Expect = 0.032
Identities = 57/208 (27%), Positives = 85/208 (40%), Gaps = 16/208 (7%)
Query: 84 GGIEIG---SPGVYNENQSLVFHSGYGFNPQMPYGPYSPVTSPLPSVGGDAQ-LYSPQ-- 137
GG+ G SP + + F+ G G++P P P ++ PS GG + +YSP
Sbjct: 1522 GGMSPGAGFSPAGNTDGGASPFNEG-GWSPASPGDPLGALSPRTPSYGGMSPGVYSPSSP 1580
Query: 138 QFPYTPPYYNQLVP--PPNLPYLNSPTPVSQPELTNLLGIDQQVESNFFGPRASY-PSVG 194
QF T P+Y+ P P P +PVS P + ++ SY P+
Sbjct: 1581 QFSMTSPHYSPTSPSYSPTSPAAGQ-SPVS-PSYSPTSPSYSPTSPSYSPTSPSYSPTSP 1638
Query: 195 SFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSV-PQQPIGS 253
S++ S +P S S+ S + S S S + S ++ P SP+ P P S
Sbjct: 1639 SYSPTSPSYSPTSPSYSPSSPSYSPS-SPSYSPSSPRYSPTSPTYSPTSPTYSPTSPTYS 1697
Query: 254 VRSFGPSAGMASHQQRSLYGFGSSSNPY 281
S P+ S S G+ SS Y
Sbjct: 1698 PTS--PTYSPTSPSYESGGGYSPSSPKY 1723
Score = 35.8 bits (81), Expect = 0.35
Identities = 46/209 (22%), Positives = 76/209 (36%), Gaps = 25/209 (11%)
Query: 13 PESSKFSNSSQGTTTIGHSRDITSQSGSFGSGADQPLYPPNVYAPQAQAFYYGGFDNGNG 72
P S +S SS + + TS + S S P P Y+P + ++ ++G G
Sbjct: 1663 PSSPSYSPSSPRYSPTSPTYSPTSPTYSPTSPTYSPTSP--TYSPTSPSY-----ESGGG 1715
Query: 73 EWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMP-YGPYSPVTSPLPSVGGDA 131
P Y + + Y+ + ++P P Y P SP +P G +
Sbjct: 1716 YSPSSPKYSPSSPTYSPTSPSYSPTSPQYSPTSPQYSPSSPTYTPSSPTYNPTSPRGFSS 1775
Query: 132 QLYSPQQFPYTP--PYYNQLVPPPNLPYLNSPTPVSQPELTNLLGIDQQVESNFFGPRAS 189
YSP Y+P P Y P+ P + +P P + G Q
Sbjct: 1776 PQYSPTSPTYSPTSPSYT-----PSSPQYSPTSPTYTPSPSEQPGTSAQYS--------- 1821
Query: 190 YPSVGSFARGSFPVAPGSFSFHESQQGFD 218
P+ +++ S +P S S+ S +D
Sbjct: 1822 -PTSPTYSPSSPTYSPASPSYSPSSPTYD 1849
>SMF1_HUMAN (O14497) SWI/SNF-related, matrix-associated,
actin-dependent regulator of chromatin subfamily F
member 1 (SWI-SNF complex protein p270) (B120)
Length = 1902
Score = 38.9 bits (89), Expect = 0.041
Identities = 56/225 (24%), Positives = 83/225 (36%), Gaps = 42/225 (18%)
Query: 91 PGVYNENQSLVFHSGYGFNPQMPYGPYS---PVTSP--LPSVGGDAQLYSPQQFPYTPPY 145
P Y + +++ +PQ PYS P +P PS Q PQ PPY
Sbjct: 76 PSGYGQQGQTPYYNQQSPHPQQQQPPYSQQPPSQTPHAQPSYQQQPQSQPPQLQSSQPPY 135
Query: 146 YNQLVPPPN----LPYLN----------SPTPVSQPELTNLLGID--QQVESNFFGPRAS 189
Q PP+ PY + S P SQP+ + QQ + +A+
Sbjct: 136 SQQPSQPPHQQSPAPYPSQQSTTQQHPQSQPPYSQPQAQSPYQQQQPQQPAPSTLSQQAA 195
Query: 190 YPSVGS-------FARGSFP----VAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRS 238
YP S +++ FP ++ SF S S GG + R S
Sbjct: 196 YPQPQSQQSQQTAYSQQRFPPPQELSQDSFGSQASSAPSMTSSKGGQEDMNLSLQSRPSS 255
Query: 239 FMPLSPSVPQQPIGSVRSFGP---SAGMASHQQRSLYGFGSSSNP 280
LS S+ P+G+ + P ++G++S Q G SNP
Sbjct: 256 LPDLSGSIDDLPMGTEGALSPGVSTSGISSSQ-------GEQSNP 293
Score = 36.2 bits (82), Expect = 0.27
Identities = 45/193 (23%), Positives = 68/193 (34%), Gaps = 42/193 (21%)
Query: 110 PQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPEL 169
P G SP S + +P Q P++P P+LP + P+P
Sbjct: 267 PMGTEGALSPGVSTSGISSSQGEQSNPAQSPFSPH------TSPHLPGIRGPSP------ 314
Query: 170 TNLLGIDQQVESNFFGPRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDS 229
P S SV G P++P + ++ +S + S
Sbjct: 315 ---------------SPVGSPASVAQSRSG--PLSPAAVPGNQMPPRPPSGQSDSIMHPS 357
Query: 230 SKPSE--RQRSFMPLSPSVPQ----QPIGSVRSFGPSAG-----MASHQQRSLYGFGSSS 278
S + R +M +P +PQ QP ++ PS G M S+QQ S+ +G
Sbjct: 358 MNQSSIAQDRGYMQRNPQMPQYSSPQPGSALSPRQPSGGQIHTGMGSYQQNSMGSYGPQG 417
Query: 279 NPYG--RGYLSNP 289
YG GY P
Sbjct: 418 GQYGPQGGYPRQP 430
Score = 31.2 bits (69), Expect = 8.6
Identities = 65/291 (22%), Positives = 100/291 (34%), Gaps = 56/291 (19%)
Query: 39 GSFGSGADQP----------LYPPNVYAPQAQAFYYGGFDNGNGEWDEYPSYVNNGGIEI 88
G+ +GA QP +Y P+ Y PQ Q D Y + + G
Sbjct: 915 GNMSTGAPQPNLMPSNPDSGMYSPSRYPPQQQ-------QQQQQRHDSYGNQFSTQGTPS 967
Query: 89 GSPGVYNENQSLVFHSGYGFNPQMPYGPYSPVTSPLPSVGGDAQLYS-PQQFPYTPPYYN 147
GSP + Q+ ++ + G Y P P+ + ++YS P P
Sbjct: 968 GSP--FPSQQTTMYQQQQQNYKRPMDGTYGP-----PAKRHEGEMYSVPYSTGQGQPQQQ 1020
Query: 148 QLVPPPNLPYLNSPTPVSQPELTNLLGIDQQVESNFFGPRASYPSVGSFARGSFPVAPGS 207
QL PP P S +QP QQ N +G +YP+ + A P A G
Sbjct: 1021 QL--PPAQPQPASQQQAAQPS-------PQQDVYNQYG--NAYPATATAATERRP-AGGP 1068
Query: 208 FSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSAGMASHQ 267
+ Q G D S+ P + ++P Q +G GP A
Sbjct: 1069 QNQFPFQFGRD--------RVSAPPGTNAQQ------NMPPQMMG-----GPIQASAEVA 1109
Query: 268 QRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSFLSLENSRRHG 318
Q+ G + Y + GS+ G A G++ D + + + G
Sbjct: 1110 QQGTMWQGRNDMTYNYANRQSTGSAPQGPAYHGVNRTDEMLHTDQRANHEG 1160
>S24B_ARATH (Q9M081) Putative protein transport protein Sec24-like
At4g32640
Length = 1069
Score = 38.9 bits (89), Expect = 0.041
Identities = 63/244 (25%), Positives = 81/244 (32%), Gaps = 36/244 (14%)
Query: 51 PPNVYAPQAQAFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNP 110
PP P +Q +G + YP N + + N+ G G P
Sbjct: 6 PPGAPRPNSQ--------QNSGPPNFYPGSQGNSNALADNMQNLSLNRPPPMMPGSGPRP 57
Query: 111 QMPYGPY-SPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPEL 169
P+G P PS G + SP P PP + P PVSQP
Sbjct: 58 PPPFGQSPQPFPQQSPSYGAPQRGPSPMSRPG---------PPAGMARPGGPPPVSQP-- 106
Query: 170 TNLLGIDQQVESNF-FGPRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSD 228
G V N GP + PS GS R S P P + S GF G +
Sbjct: 107 ---AGFQSNVPLNRPTGPPSRQPSFGS--RPSMPGGPVA-QPAASSSGFPAFGPSGSVAA 160
Query: 229 SSKPSERQRSFMPLSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSN 288
P R +F SP P+GS S PS + GS P G +
Sbjct: 161 GPPPGSRPMAFG--SP----PPVGSGMSMPPSGMIGGPVSNGHQMVGSGGFPRGTQF--- 211
Query: 289 PGSS 292
PG++
Sbjct: 212 PGAA 215
Score = 32.3 bits (72), Expect = 3.9
Identities = 24/63 (38%), Positives = 28/63 (44%), Gaps = 5/63 (7%)
Query: 110 PQMPYGPYSPVTSPLPSVGGDAQLYSPQQFP--YTPPYYNQLVPPPNLPYLNSPTPVSQP 167
P P+G P S LP AQ+ P FP PP Y + P PN N PT + QP
Sbjct: 265 PGAPHG--RPAVSGLPYGPPSAQVAPPLGFPGQMQPPRYG-MGPLPNQSMTNIPTAMGQP 321
Query: 168 ELT 170
T
Sbjct: 322 GAT 324
>KCN3_RAT (P70605) Small conductance calcium-activated potassium
channel protein 3 (SK3)
Length = 732
Score = 38.1 bits (87), Expect = 0.071
Identities = 57/233 (24%), Positives = 82/233 (34%), Gaps = 11/233 (4%)
Query: 123 PLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPELTNLLGIDQQVESN 182
P PS G + Q P PP Q PP P L P Q + QQ +
Sbjct: 22 PCPSSGDEQQQQQQPPPPSAPPAVPQ--QPPG-PLLQPQPPQLQQQQQQQQQQQQQQQQQ 78
Query: 183 FFGPRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPL 242
P P + S V PG H S F S + PS RQ S + L
Sbjct: 79 QQAPLHPLPQLAQLQ--SQLVHPG--LLHSSPTAFRAPNSANS-TAILHPSSRQGSQLNL 133
Query: 243 SPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLS 302
+ + S + GP G + H+Q S SNP+ +S+ + G + LS
Sbjct: 134 NDHLLGHSPSSTATSGPGGG-SRHRQASPLVHRRDSNPFTEIAMSS--CKYSGGVMKPLS 190
Query: 303 ANDRSFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKNHISSENN 355
S +L + G+ F N I+S + L +H ++ +N
Sbjct: 191 RLSASRRNLIEAEPEGQPLQLFSPSNPPEIIISSREDNHAHQTLLHHPNATHN 243
>TCG1_HUMAN (O14776) Transcription elongation regulator 1 (TATA
box-binding protein-associated factor 2S) (Transcription
factor CA150)
Length = 1098
Score = 37.7 bits (86), Expect = 0.092
Identities = 20/63 (31%), Positives = 31/63 (48%), Gaps = 5/63 (7%)
Query: 110 PQMPYG--PYSPVTSPLPSVGGDAQLYSP---QQFPYTPPYYNQLVPPPNLPYLNSPTPV 164
P+ P+G P+ P P+P GG P Q+ P+ PP + + PPP + + PV
Sbjct: 63 PRPPFGRPPFDPNMPPMPPPGGIPPPMGPPHLQRPPFMPPPMSSMPPPPGMMFPPGMPPV 122
Query: 165 SQP 167
+ P
Sbjct: 123 TAP 125
>Y066_NPVOP (Q83949) Hypothetical 98.6 kDa protein (ORF71)
Length = 875
Score = 37.0 bits (84), Expect = 0.16
Identities = 26/71 (36%), Positives = 28/71 (38%), Gaps = 5/71 (7%)
Query: 110 PQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPEL 169
P PYG Y P P P Q PQQ P PP Q P P P P P P
Sbjct: 91 PFYPYGQYWPQQPPQPPPDQPQQPQPPQQPPQQPP---QQQPQPPQPPQQPPQPPQPP-- 145
Query: 170 TNLLGIDQQVE 180
L + QQ+E
Sbjct: 146 PQQLALVQQIE 156
>TBP_DROME (P20227) TATA-box binding protein (TATA-box factor) (TATA
binding factor) (TATA sequence-binding protein)
(Transcription initiation factor TFIID TBP subunit)
Length = 353
Score = 36.6 bits (83), Expect = 0.21
Identities = 40/160 (25%), Positives = 61/160 (38%), Gaps = 24/160 (15%)
Query: 153 PNLPYLNSPTPVSQPELTNLLGIDQQVESN--FFGPRASYPSVGSFARGSFPVAPGSFSF 210
PN + TP+ Q E DQQ+ +N + P S P S APGS S
Sbjct: 7 PNFSIPSIGTPLHQMEA------DQQIVANPVYHPPAVSQPD-------SLMPAPGSSSV 53
Query: 211 HESQQGFDGSRSGGLWSDSSKPS-----ERQRSFMPLSPSVPQQPIGSVRSFGPSAG--- 262
QQ SGG +PS ++ +S+ P + QQ ++S P G
Sbjct: 54 QHQQQQQQSDASGGSGLFGHEPSLPLAHKQMQSYQPSASYQQQQQQQQLQSQAPGGGGST 113
Query: 263 -MASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGL 301
+ Q ++ + P + G+ GG A+S +
Sbjct: 114 PQSMMQPQTPQSMMAHMMPMSERSVGGSGAGGGGDALSNI 153
>HORN_HUMAN (Q86YZ3) Hornerin
Length = 2850
Score = 36.6 bits (83), Expect = 0.21
Identities = 36/95 (37%), Positives = 40/95 (41%), Gaps = 11/95 (11%)
Query: 206 GSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVPQQPIGSVRSFGPS-AGMA 264
GS S H S G GSRSG S ER RS S S Q GS +S G G
Sbjct: 248 GSGSGHSSGYGQHGSRSG-----QSSRGERHRSSSGSSSSYGQHGSGSRQSLGHGRQGSG 302
Query: 265 SHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAIS 299
S Q S GS G G+ S+ G GS+ S
Sbjct: 303 SRQSPSHVRHGS-----GSGHSSSHGQHGSGSSYS 332
Score = 36.2 bits (82), Expect = 0.27
Identities = 50/181 (27%), Positives = 68/181 (36%), Gaps = 21/181 (11%)
Query: 206 GSFSFHESQQGFDGSRSGGLWSDSSKPSERQR-SFMPLSPSVPQQPIGSVRS-------- 256
GS S S G GS SG P QR S SPS + GS RS
Sbjct: 1576 GSGSSQSSGYGRQGSGSG------QSPGHGQRGSGSRQSPSYGRHGSGSGRSSSSGQHGS 1629
Query: 257 -FGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSANDRSFLSLENSR 315
G S+G H+ S G SS + +G G + G GS S +R S +S
Sbjct: 1630 GLGESSGFGHHESSS--GQSSSYSQHGSGSGHSSGYGQHGSRSGQSSRGERHGSSSRSSS 1687
Query: 316 RHGRETASFCRCNGTLDILSEQNRGP---RASKLKNHISSENNAIDGSKNNASTAKFQDE 372
R+G+ + + +G S + P R H SS GS ++S ++
Sbjct: 1688 RYGQHGSGSRQSSGHGRQGSGSGQSPSRGRHGSGLGHSSSHGQHGSGSGRSSSRGPYESR 1747
Query: 373 S 373
S
Sbjct: 1748 S 1748
Score = 35.8 bits (81), Expect = 0.35
Identities = 35/105 (33%), Positives = 39/105 (36%), Gaps = 16/105 (15%)
Query: 206 GSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSPSVPQQPIGSVRSFG------- 258
GS S H S G GSRSG S ERQ S S S Q GS +S G
Sbjct: 482 GSGSGHSSGHGQHGSRSG-----QSSRGERQGSSAGSSSSYGQHGSGSRQSLGHSRHGSG 536
Query: 259 ----PSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAIS 299
PS H+ S + YG G S+ G GS S
Sbjct: 537 SGQSPSPSRGRHESGSRQSSSYGPHGYGSGRSSSRGPYESGSGHS 581
Score = 32.3 bits (72), Expect = 3.9
Identities = 32/96 (33%), Positives = 37/96 (38%), Gaps = 8/96 (8%)
Query: 206 GSFSFHESQQGFDGSRSGGLWSDSSK-PSERQRSFMPLSPSVPQQPIGSVRSFGPSAGMA 264
GS S H S QG GS SG S S RQ S S + G + S
Sbjct: 365 GSGSGHSSSQGQHGSTSGQASSSGQHGSSSRQSSSYGQHESASRHSSGRGQHSSGSGQSP 424
Query: 265 SHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISG 300
H QR G GS +P G+ FG S+ SG
Sbjct: 425 GHGQR---GSGSGQSPSS----GQHGTGFGRSSSSG 453
>SIL1_MOUSE (Q8C0T5) Signal-induced proliferation-associated 1 like
protein 1
Length = 1782
Score = 36.2 bits (82), Expect = 0.27
Identities = 65/305 (21%), Positives = 111/305 (36%), Gaps = 46/305 (15%)
Query: 80 YVNNGGIEIGSPGVYNENQSLVFHSGYG-FNPQMPYGPYSPVTSPLPSVGGDAQLYSPQQ 138
Y N GI + N ++ G +PQ+P SP+TS L + GD ++ P++
Sbjct: 1048 YQMNEGISYEFKFPFRNNNKWQRNASKGAHSPQVPSQLQSPMTSRLNAGKGDGKMPPPER 1107
Query: 139 FPYTP--------PYYNQLVPPPNLPYLNSPTPVSQPELTNLLGIDQQVESNFFGPRASY 190
P P +L P ++ S +++ + N SN +
Sbjct: 1108 AANIPRSISSDGRPLERRLSPGSDIYVTVSSMALARSQCRN-------SPSNLSSSSETG 1160
Query: 191 PSVGSFARGSFPV-------APGSFSFHESQQGFDGS--------RSGGLWSDSSKPSER 235
G++ + S P +P S +Q GS RS +D +P+
Sbjct: 1161 SGGGTYRQKSMPEGFGVSRRSPASIDRQNTQSDISGSGKSTPSWQRSEDSLADQMEPTCH 1220
Query: 236 QRSFMPLSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGG 295
+ + P+ + P G + P ++ ++Q G S + Y SN SS
Sbjct: 1221 LPAVSKVLPAFRESPSGRLMRQDPVVHLSPNKQ------GHSDSHYSSHSSSNTLSSNAS 1274
Query: 296 SAISGLS--ANDRSFLSLEN-------SRRHGRETASFCRCNGTLDILSEQNRGPRASKL 346
SA S DR+ L + S G +TAS+ +G+ L PR+
Sbjct: 1275 SAHSDEKWYDGDRTESDLNSYNYLQGTSADSGIDTASYGPSHGSTASLGASTSSPRSGPG 1334
Query: 347 KNHIS 351
K ++
Sbjct: 1335 KEKVA 1339
>RPB1_DROME (P04052) DNA-directed RNA polymerase II largest subunit
(EC 2.7.7.6)
Length = 1887
Score = 36.2 bits (82), Expect = 0.27
Identities = 40/155 (25%), Positives = 61/155 (38%), Gaps = 14/155 (9%)
Query: 13 PESSKFSNSSQG--TTTIGHSRDITSQSGSFGSGADQPLYPP-NVYAPQAQAFYYGGFDN 69
P+S+ +S SS G T+ +S + QS +G+ +Y P N Y+P + + N
Sbjct: 1626 PQSTGYSPSSSGYSPTSPVYSPTVQFQSSPSFAGSGSNIYSPGNAYSPSSSNYS----PN 1681
Query: 70 GNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMP-YGPYSPVTSPLPSVG 128
PSY + + Y+ + + P P Y P SP S P
Sbjct: 1682 SPSYSPTSPSYSPSSPSYSPTSPCYSPTSPSYSPTSPNYTPVTPSYSPTSPNYSASPQYS 1741
Query: 129 GDAQLYSPQQFPYTP--PYYNQLVPPPNLPYLNSP 161
+ YS Y+P P Y+ PP+ Y SP
Sbjct: 1742 PASPAYSQTGVKYSPTSPTYS----PPSPSYDGSP 1772
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.313 0.132 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 82,962,204
Number of Sequences: 164201
Number of extensions: 4021283
Number of successful extensions: 8672
Number of sequences better than 10.0: 125
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 116
Number of HSP's that attempted gapping in prelim test: 8421
Number of HSP's gapped (non-prelim): 311
length of query: 632
length of database: 59,974,054
effective HSP length: 116
effective length of query: 516
effective length of database: 40,926,738
effective search space: 21118196808
effective search space used: 21118196808
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 69 (31.2 bits)
Medicago: description of AC144538.7


