

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144538.13 + phase: 0 /pseudo
(1574 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
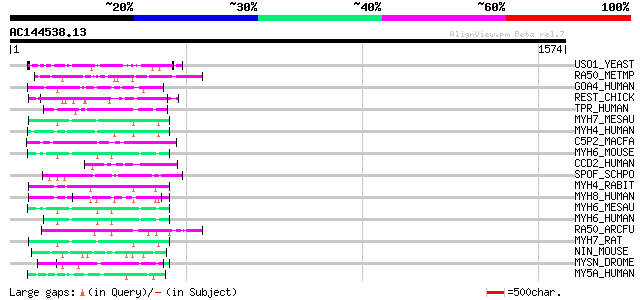
Sequences producing significant alignments: (bits) Value
USO1_YEAST (P25386) Intracellular protein transport protein USO1 63 6e-09
RA50_METMP (P62134) DNA double-strand break repair rad50 ATPase 57 3e-07
GOA4_HUMAN (Q13439) Golgi autoantigen, golgin subfamily A member... 56 9e-07
REST_CHICK (O42184) Restin (Cytoplasmic linker protein-170) (CLI... 53 8e-06
TPR_HUMAN (P12270) Nucleoprotein TPR 52 2e-05
MYH7_MESAU (P13540) Myosin heavy chain, cardiac muscle beta isof... 51 3e-05
MYH4_HUMAN (Q9Y623) Myosin heavy chain, skeletal muscle, fetal (... 51 3e-05
C5P2_MACFA (Q9BE52) CDK5 regulatory subunit associated protein 2... 50 5e-05
MYH6_MOUSE (Q02566) Myosin heavy chain, cardiac muscle alpha iso... 50 6e-05
CCD2_HUMAN (Q96LB3) Coiled-coil domain containing protein 2 (Cap... 50 6e-05
SPOF_SCHPO (Q10411) Sporulation-specific protein 15 49 8e-05
MYH4_RABIT (Q28641) Myosin heavy chain, skeletal muscle, juvenile 49 8e-05
MYH8_HUMAN (P13535) Myosin heavy chain, skeletal muscle, perinat... 49 1e-04
MYH6_MESAU (P13539) Myosin heavy chain, cardiac muscle alpha iso... 49 1e-04
MYH6_HUMAN (P13533) Myosin heavy chain, cardiac muscle alpha iso... 49 1e-04
RA50_ARCFU (O29230) DNA double-strand break repair rad50 ATPase 48 2e-04
MYH7_RAT (P02564) Myosin heavy chain, cardiac muscle beta isofor... 48 2e-04
NIN_MOUSE (Q61043) Ninein 48 2e-04
MYSN_DROME (Q99323) Myosin heavy chain, non-muscle (Zipper prote... 48 2e-04
MY5A_HUMAN (Q9Y4I1) Myosin Va (Myosin 5A) (Dilute myosin heavy c... 48 2e-04
>USO1_YEAST (P25386) Intracellular protein transport protein USO1
Length = 1790
Score = 63.2 bits (152), Expect = 6e-09
Identities = 94/428 (21%), Positives = 175/428 (39%), Gaps = 72/428 (16%)
Query: 54 EKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHV 113
+KK ED + GK + ALSRE + CKN ++ D V+H+
Sbjct: 855 KKKAEDGINKMGKDLF-----------ALSREMQAVEENCKNLQKEKDKSNVNHQ--KET 901
Query: 114 KETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRS 173
K + DI + + E + I+E I N L S EK+ E V R +S
Sbjct: 902 KSLKEDIAAK------ITEIKAINENLEEMKIQCNNL-SKEKEHISKELVEYKSRF--QS 952
Query: 174 WRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEM 233
++V +TE +D+ ++ ++E K+ + ++ L+ +S++QE E
Sbjct: 953 HDNLVAKLTEKLKSLANNYKDMQAENESLIKAVEESKNESSIQLSNLQNKIDSMSQEKE- 1011
Query: 234 NFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGC 293
NF ++ +++ Q L++ I +Q K ++ +++K E
Sbjct: 1012 NFQIERGSIEKNIEQ---------LKKTISDLEQTKEEIISKSDSSKDE----------- 1051
Query: 294 YKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNE-EVNL 352
Y+++ L K K T + E + T E+ +A+L NE E L
Sbjct: 1052 -----YESQISLLKEKLETATTANDENVNKISELTK-TREELEAELAAYKNLKNELETKL 1105
Query: 353 DPCSSC-----EKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDL---- 403
+ E EHL + LE+E + + LR E LEKE++DL
Sbjct: 1106 ETSEKALKEVKENEEHLKEEKI----QLEKEATETKQQLNSLRANLESLEKEHEDLAAQL 1161
Query: 404 ------IKNIQHSESKEVSENKN---NSQKENIILKENVLKLKNDISNFVKSTETFQKIM 454
I N + ++E+S+ + ++Q+EN +K+ +L+ ++ ++E +
Sbjct: 1162 KKYEEQIANKERQYNEEISQLNDEITSTQQENESIKKKNDELEGEVKAMKSTSEEQSNLK 1221
Query: 455 GSQVSVFD 462
S++ +
Sbjct: 1222 KSEIDALN 1229
Score = 52.4 bits (124), Expect = 1e-05
Identities = 96/457 (21%), Positives = 196/457 (42%), Gaps = 62/457 (13%)
Query: 56 KKEDWSEEEGKRMLLNFKAKLFLTMALSREEY--DRVQECKNAKEIWDTLKVHHEGTSHV 113
K + S+ E L N + K+ +M+ +E + +R KN +++ T+ E T
Sbjct: 983 KAVEESKNESSIQLSNLQNKID-SMSQEKENFQIERGSIEKNIEQLKKTIS-DLEQTKEE 1040
Query: 114 KETRIDIGVRKFE-----LFEMKETETI--DEMYGRFTIIMNELRSLEKDFTIHERVRKI 166
++ D ++E L E ET T DE + + + LE + ++ ++
Sbjct: 1041 IISKSDSSKDEYESQISLLKEKLETATTANDENVNKISELTKTREELEAELAAYKNLKNE 1100
Query: 167 LRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQES 226
L T K LK+++ + LK ++ L+++ + K ++ +L+ N ES
Sbjct: 1101 LE---------TKLETSEKALKEVKENE--EHLKEEKIQLEKEATETKQQLNSLRANLES 1149
Query: 227 LNQE----------LEMNFDNQQQELDEEDHQ--DQIILLTRKLQRMIQRRDQNKRNFPA 274
L +E E N++++ +EE Q D+I ++ + + ++ D+ + A
Sbjct: 1150 LEKEHEDLAAQLKKYEEQIANKERQYNEEISQLNDEITSTQQENESIKKKNDELEGEVKA 1209
Query: 275 RKENAKTE--LDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTN 332
K ++ + L KS++ + + K N +TN+ S + + E +T
Sbjct: 1210 MKSTSEEQSNLKKSEIDALNLQ----------IKELKKKN-ETNEASLLESIKSVESETV 1258
Query: 333 E-DEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRK 391
+ E D C + + E+ +S +K + L++E E+++ E
Sbjct: 1259 KIKELQDECNFKEKEVSELEDKLKASEDKNSKYLE--------LQKESEKIKEELDAKTT 1310
Query: 392 EKEILEKENKDLIKNIQHSESKEVSENKNNSQKENIILKENVLKLKNDISNFVKSTETFQ 451
E +I ++ +L K + SES E+S K S +E +E + KLKN+I ++ E +
Sbjct: 1311 ELKIQLEKITNLSKAKEKSES-ELSRLKKTSSEERKNAEEQLEKLKNEIQIKNQAFEKER 1369
Query: 452 KIMGSQVSVFDKACIGFKTNQKQKLYENFFIPKKNKN 488
K++ S + + ++K E+ I +N+N
Sbjct: 1370 KLLNEGSSTITQ-----EYSEKINTLEDELIRLQNEN 1401
Score = 50.8 bits (120), Expect = 3e-05
Identities = 90/431 (20%), Positives = 188/431 (42%), Gaps = 67/431 (15%)
Query: 50 EDLNEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEG 109
++ N K+KE SE E K K +L + ++E ++++E +AK LK+ E
Sbjct: 1265 DECNFKEKEV-SELEDKLKASEDKNSKYLEL---QKESEKIKEELDAKTT--ELKIQLEK 1318
Query: 110 TSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTI--IMNELRSLEKDFTIHERVRKIL 167
+++ + + + EL +K+T + + + + NE++ + F E+ RK+L
Sbjct: 1319 ITNLSKAKEK---SESELSRLKKTSSEERKNAEEQLEKLKNEIQIKNQAF---EKERKLL 1372
Query: 168 RCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESL 227
+ IT+ K LED + L+ L ++ + +S++ + + + L
Sbjct: 1373 N-------EGSSTITQEYSEKINTLEDELIRLQNENELKAKEIDNTRSELEKVSLSNDEL 1425
Query: 228 NQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQ 287
+E + + Q E+ ++D+I TR ++++ NKR+ + KE + +
Sbjct: 1426 LEEKQNTIKSLQDEI--LSYKDKI---TRNDEKLLSIERDNKRDLESLKEQLRAAQESKA 1480
Query: 288 VTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDN 347
G KL ++ KS + +S + ++NE E KS
Sbjct: 1481 KVEEGLKKLEEESSKEKAELEKSKEMMKKLESTI--------ESNETE-------LKSSM 1525
Query: 348 EEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLEL-----RKEKEILEKENKD 402
E + S EK E S + E++ + L++E +L EK+I E ++K
Sbjct: 1526 ETIR----KSDEKLEQ-------SKKSAEEDIKNLQHEKSDLISRINESEKDIEELKSKL 1574
Query: 403 LIKNIQHSESKEVSENKNNSQ-------KENIILKENVLKLKNDISN---FVKSTETFQK 452
I+ SE + V + NN+Q +EN +LK + ++ ++ + +KS + ++
Sbjct: 1575 RIEAKSGSELETVKQELNNAQEKIRINAEENTVLKSKLEDIERELKDKQAEIKSNQEEKE 1634
Query: 453 IMGSQVSVFDK 463
++ S++ ++
Sbjct: 1635 LLTSRLKELEQ 1645
Score = 44.7 bits (104), Expect = 0.002
Identities = 71/367 (19%), Positives = 157/367 (42%), Gaps = 26/367 (7%)
Query: 108 EGTSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIH-ERVRKI 166
+ + +K ID + E + E ++E + +E+ S + T + E++ I
Sbjct: 1398 QNENELKAKEIDNTRSELEKVSLSNDELLEEKQNTIKSLQDEILSYKDKITRNDEKLLSI 1457
Query: 167 LRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQES 226
R R + + A++ K ++E+ + L+ + + +K M L++ ES
Sbjct: 1458 ERDNKRDLESLKEQLRAAQE-SKAKVEEGLKKLEEESSKEKAELEKSKEMMKKLESTIES 1516
Query: 227 LNQELEMNFD-----NQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKT 281
EL+ + + +++ E ++ ++ I L + +I R ++++++ K +
Sbjct: 1517 NETELKSSMETIRKSDEKLEQSKKSAEEDIKNLQHEKSDLISRINESEKDIEELKSKLRI 1576
Query: 282 ELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCL 341
E +L K E N ++ + + + + + E E+ +D+QA++
Sbjct: 1577 EAKSGS-------ELETVKQELN-NAQEKIRINAEENTVLKSKLEDIERELKDKQAEI-- 1626
Query: 342 MAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQE----CERLRNENLELRKEKEILE 397
KS+ EE L S ++ E D+ +Q E+E + + E +L ++ +LE
Sbjct: 1627 --KSNQEEKEL-LTSRLKELEQELDSTQQKAQKSEEERRAEVRKFQVEKSQLDEKAMLLE 1683
Query: 398 KENKDLIKNIQHSESKEVSENK-NNSQKENI-ILKENVLKLKNDISNFVKSTETFQKIMG 455
+ DL+ Q + E + K +SQ++ I L + + LK + S ++ E +I
Sbjct: 1684 TKYNDLVNKEQAWKRDEDTVKKTTDSQRQEIEKLAKELDNLKAENSKLKEANEDRSEIDD 1743
Query: 456 SQVSVFD 462
+ V D
Sbjct: 1744 LMLLVTD 1750
Score = 37.4 bits (85), Expect = 0.33
Identities = 84/425 (19%), Positives = 170/425 (39%), Gaps = 32/425 (7%)
Query: 82 LSREEYDRVQ-ECKNAKEIWDTLKVHHEGT-SHVKETRIDIGVRKFELFEMKE--TETID 137
+S EE +++Q +C K +L+ E T ++ E I + EL E + +
Sbjct: 727 ISFEEVEKLQRQCTKLKGEITSLQTETESTHENLTEKLIALTNEHKELDEKYQILNSSHS 786
Query: 138 EMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLEDLIG 197
+ F+I+ EL+++ ++R +L + + TA+ E K + ED I
Sbjct: 787 SLKENFSILETELKNVRDSLDEMTQLRDVLETKDKENQ---TALLEYKSTIH-KQEDSIK 842
Query: 198 SL-KAHEVLLQEDKSS----NKSKMIALKTNQESLNQELEMNFDNQQQELDEED--HQDQ 250
+L K E +L + K + NK ++E Q +E N N Q+E D+ + HQ +
Sbjct: 843 TLEKGLETILSQKKKAEDGINKMGKDLFALSREM--QAVEENCKNLQKEKDKSNVNHQKE 900
Query: 251 IILLTRKLQRMIQRRDQNKRNFPARKENAKTELDK-SQVTCYGCYKLGHYKNECPLNKRK 309
T+ L+ I + + E K + + S+ + +L YK+ +
Sbjct: 901 ----TKSLKEDIAAKITEIKAINENLEEMKIQCNNLSKEKEHISKELVEYKSRFQSHDNL 956
Query: 310 SSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLF---D 366
+ L KS + + + E L + E ++ + K + + +
Sbjct: 957 VAKLTEKLKSLANNYKDMQA-----ENESLIKAVEESKNESSIQLSNLQNKIDSMSQEKE 1011
Query: 367 NLFYSSQMLEQECERLRN--ENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQK 424
N +E+ E+L+ +LE KE+ I + ++ Q S KE E +
Sbjct: 1012 NFQIERGSIEKNIEQLKKTISDLEQTKEEIISKSDSSKDEYESQISLLKEKLETATTAND 1071
Query: 425 ENIILKENVLKLKNDISNFVKSTETFQKIMGSQVSVFDKACIGFKTNQKQKLYENFFIPK 484
EN+ + K + ++ + + + + + +++ +KA K N++ E + K
Sbjct: 1072 ENVNKISELTKTREELEAELAAYKNLKNELETKLETSEKALKEVKENEEHLKEEKIQLEK 1131
Query: 485 KNKNT 489
+ T
Sbjct: 1132 EATET 1136
Score = 35.8 bits (81), Expect = 0.95
Identities = 77/375 (20%), Positives = 152/375 (40%), Gaps = 81/375 (21%)
Query: 49 DEDLNEKKKEDWSEEEGKRMLLNFKAKLFLT-MALSREEYDRVQECKNAKEIWDTLKVHH 107
DE L EK+ S ++ +L++K K+ L E D ++ ++ KE L+
Sbjct: 1423 DELLEEKQNTIKSLQDE---ILSYKDKITRNDEKLLSIERDNKRDLESLKE---QLRAAQ 1476
Query: 108 EGTSHVKETRIDIGVRKFELFEMKETETID---EMYGRF--TIIMNELRSLEKDFTIHER 162
E + V+E G++K E KE ++ EM + TI NE TI +
Sbjct: 1477 ESKAKVEE-----GLKKLEEESSKEKAELEKSKEMMKKLESTIESNETELKSSMETIRKS 1531
Query: 163 VRKI----------LRCLPRSWRHIVTAITEA-KDLKKLRLEDLIGSLKAHEVLLQEDKS 211
K+ ++ L +++ I E+ KD+++L+ + I + E+ + +
Sbjct: 1532 DEKLEQSKKSAEEDIKNLQHEKSDLISRINESEKDIEELKSKLRIEAKSGSELETVKQEL 1591
Query: 212 SNKSKMIALKTNQ----ESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQ 267
+N + I + + +S +++E ++Q E+ + +Q++ LLT +L+ + Q D
Sbjct: 1592 NNAQEKIRINAEENTVLKSKLEDIERELKDKQAEI--KSNQEEKELLTSRLKELEQELDS 1649
Query: 268 NKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEP 327
++ +E + E+ K QV
Sbjct: 1650 TQQKAQKSEEERRAEVRKFQV--------------------------------------- 1670
Query: 328 EEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENL 387
E++ DE+A L+ N+ VN + ++ D + ++ QE E+L E
Sbjct: 1671 -EKSQLDEKA--MLLETKYNDLVNKEQAWKRDE-----DTVKKTTDSQRQEIEKLAKELD 1722
Query: 388 ELRKEKEILEKENKD 402
L+ E L++ N+D
Sbjct: 1723 NLKAENSKLKEANED 1737
>RA50_METMP (P62134) DNA double-strand break repair rad50 ATPase
Length = 993
Score = 57.4 bits (137), Expect = 3e-07
Identities = 119/539 (22%), Positives = 230/539 (42%), Gaps = 97/539 (17%)
Query: 71 NFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEM 130
N+K L L + + +E +++ K+ E ++ LK E S +KE +GV K L +
Sbjct: 284 NYKKYLELELKI-KELNNKLIGHKSNYESYNKLKTIEE--SLLKE----LGVLKESLKDN 336
Query: 131 KET-----ETIDEMYGRFTII------MNELRSLEK---DFTIHERVRKILRCLPRSWRH 176
K+ E + E + I+ + EL +EK + IH++ + L + +
Sbjct: 337 KKNPDELKENLKENDEKILILDKIKEKIKELEFIEKQIYEIKIHKKTVETLFDSVKIYDD 396
Query: 177 IVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFD 236
+ E K KK E+L+ E LQ + + K+K+I+ T+ E + +++ N +
Sbjct: 397 SIKTFEELKT-KKNSYENLLKEKFDLEKKLQNE-TDEKTKLISELTDFEKIEEKI--NLE 452
Query: 237 NQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYK- 295
N+ +E ED ++I KL ++ +++ + +N+K EL+K++ +C+ C
Sbjct: 453 NELKE-KYEDLSEKI----DKLNEIVLKKESKISEY----KNSKAELEKTKDSCHVCQSK 503
Query: 296 ---------LGHYKNECP----------------LNKRKSSNLQTNQ-KSFMTTWDEPEE 329
L Y +E LNK++ ++ N+ SF + E +E
Sbjct: 504 ITEEKKQELLEKYNSEIQNEQLSTESLKKQLEIILNKKEKMKVKLNEIDSFKLKYGELKE 563
Query: 330 QTNEDEQADLCLMAKSD---------------NEEVNLDPCSSCEKTEHLFDNLFYSSQM 374
+ N + + ++ ++ N+E++L + + E+ + N YSSQ
Sbjct: 564 KKNYSLKVEESIIETTEKLNELTGKINEYSSLNDEISLIE-NKLKNLENDYKNCNYSSQF 622
Query: 375 LEQECE-RLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNN--SQKENIILKE 431
L + E + LEL K I+ + I+N + S E KN + + I LK+
Sbjct: 623 LTKNDESEFLTKKLELSK---IIGDYDSSKIENEKKSLENLKDELKNTIYNLEREINLKK 679
Query: 432 NVLKLKNDISNFVKSTETFQKIMGSQVSVFDKACIGFKTNQKQKLYENFFIPKKNKNTCE 491
+ ++NDIS+ + E + K ++ S F+ N+ + EN+ ++ +
Sbjct: 680 ELKNIQNDISSKIGIVECYVK-WETEKSDFE--------NKLSECKENYEKYMESLAVLK 730
Query: 492 RYKCTYCEKYGHLEPFCFRK--RKRLASQKFKNNYTNFHKKETLNNTN---HQGPKKIW 545
Y TY + +L F +K K+ +K T K N N H+ K+++
Sbjct: 731 NYSKTYSVEINNLNEFLNQKIAEKQQFCEKLLETRTEIEKNIQTVNYNPELHENAKRLY 789
Score = 35.8 bits (81), Expect = 0.95
Identities = 34/107 (31%), Positives = 58/107 (53%), Gaps = 10/107 (9%)
Query: 347 NEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNE--NLELRKEKEILEKENKDLI 404
+E++N+ S E L L ++LE E+L+NE E+ KE+ + + EN + +
Sbjct: 164 SEKMNIVKKSYEETLLKLEGELTQEPEILEN-LEKLKNEVSESEILKEEILKKYENLEKL 222
Query: 405 KNIQHSESKEVSEN--KNNSQKENIILKENVLKLKN---DISNFVKS 446
K ++SE ++ E +NN KEN LK+ + ++KN +I NF S
Sbjct: 223 KLEKNSEILQMEEKFAENNQLKEN--LKDIISEIKNINLEIQNFKNS 267
>GOA4_HUMAN (Q13439) Golgi autoantigen, golgin subfamily A member 4
(Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (72.1
protein)
Length = 2230
Score = 55.8 bits (133), Expect = 9e-07
Identities = 97/407 (23%), Positives = 168/407 (40%), Gaps = 53/407 (13%)
Query: 50 EDLNEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREE--YDRVQECKNAKEIWDTLKVHH 107
E+L +KK +E E K L +A+ + T L E +QE KN + L VH
Sbjct: 538 EELELQKKAILTESENKLRDLQQEAETYRTRILELESSLEKSLQENKNQSK---DLAVHL 594
Query: 108 EG--TSHVKETRIDIGVRKFELFEMKE------TETIDEMYGRFTIIMNELRSLEKDFTI 159
E H KE + + K EL +K TE + + ++ M +LR EK
Sbjct: 595 EAEKNKHNKEITVMVEKHKTELESLKHQQDALWTEKLQVLKQQYQTEMEKLR--EKCEQE 652
Query: 160 HERVRKILRCLPRSWRHIVTAIT-EAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMI 218
E + K + ++ + T E D+K+ LE L L EVL K + ++
Sbjct: 653 KETLLKDKEIIFQAHIEEMNEKTLEKLDVKQTELESLSSELS--EVLKARHKL--EEELS 708
Query: 219 ALKTNQESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKEN 278
LK + + QELE D Q+ HQ Q+ + ++ + IQR ++ ++ + E
Sbjct: 709 VLKDQTDKMKQELEAKMDEQKNH-----HQQQVDSIIKEHEVSIQRTEKALKDQINQLEL 763
Query: 279 AKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQAD 338
E DK H +N KR LQ + + + + T+E +A
Sbjct: 764 LLKERDKHLKE-----HQAHVENLEADIKRSEGELQ-QASAKLDVFQSYQSATHEQTKAY 817
Query: 339 LCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQE------CERLRNENLELRKE 392
+A+ + ++L+ + + + Q+ E E C L ++++
Sbjct: 818 EEQLAQLQQKLLDLET-----------ERILLTKQVAEVEAQKKDVCTELDAHKIQVQDL 866
Query: 393 KEILEKENKDLIKNI----QHSESKEVSENKNNSQKENIIL-KENVL 434
+ LEK+N ++ + + Q ESK NK Q + I++ KEN++
Sbjct: 867 MQQLEKQNSEMEQKVKSLTQVYESKLEDGNKEQEQTKQILVEKENMI 913
Score = 45.4 bits (106), Expect = 0.001
Identities = 70/365 (19%), Positives = 146/365 (39%), Gaps = 26/365 (7%)
Query: 189 KLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDHQ 248
+ R+ +L SL E LQE+K+ +K + L+ + N+E+ + + + EL+ HQ
Sbjct: 566 RTRILELESSL---EKSLQENKNQSKDLAVHLEAEKNKHNKEITVMVEKHKTELESLKHQ 622
Query: 249 DQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKR 308
Q L T KLQ + Q+ + E K L K + + H +
Sbjct: 623 -QDALWTEKLQVLKQQYQTEMEKLREKCEQEKETLLKDKEIIF----QAHIEEMNEKTLE 677
Query: 309 KSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEH--LFD 366
K QT +S + E + ++ E+ L ++D + L+ +K H D
Sbjct: 678 KLDVKQTELESLSSELSEVLKARHKLEEELSVLKDQTDKMKQELEAKMDEQKNHHQQQVD 737
Query: 367 NLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKEN 426
++ ++ Q E+ + ++ + + +L++ +K L ++ H E+ E ++ + +
Sbjct: 738 SIIKEHEVSIQRTEKALKD--QINQLELLLKERDKHLKEHQAHVENLEADIKRSEGELQQ 795
Query: 427 IILKENVLKLKNDISNFVKSTETFQKIMGSQVSVFDKACIGFKTNQKQKLYENFFIPKKN 486
K +V + ++ +T K Q++ + + +T + + + +
Sbjct: 796 ASAKLDVFQ------SYQSATHEQTKAYEEQLAQLQQKLLDLETERILLTKQVAEVEAQK 849
Query: 487 KNTC---ERYKCTYCEKYGHLEPFCFRKRKRLASQKFKNNYTNFHKKETLNNTNHQGPKK 543
K+ C + +K + LE K+ QK K+ + K N + K+
Sbjct: 850 KDVCTELDAHKIQVQDLMQQLE-----KQNSEMEQKVKSLTQVYESKLEDGNKEQEQTKQ 904
Query: 544 IWVPK 548
I V K
Sbjct: 905 ILVEK 909
Score = 43.5 bits (101), Expect = 0.005
Identities = 73/391 (18%), Positives = 168/391 (42%), Gaps = 43/391 (10%)
Query: 158 TIHERVRKILRCLPR------SWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKS 211
T+ +RV++ L R S + T +T K+ + +L++ + L+ + L +K+
Sbjct: 283 TLQQRVKRQENLLKRCKETIQSHKEQCTLLTSEKEALQEQLDERLQELEKIKDLHMAEKT 342
Query: 212 SNKSKMIALKTNQESLNQELEMNFDNQQQELDE--EDHQDQIILLTRKLQRMIQRRDQNK 269
+++ K E L Q+ M ++++ E E +++I L ++++M + ++ +
Sbjct: 343 KLITQLRDAKNLIEQLEQDKGMVIAETKRQMHETLEMKEEEIAQLRSRIKQMTTQGEELR 402
Query: 270 RNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEE 329
+ A EL+K+ T +K+ + K+ M + E
Sbjct: 403 EQKEKSERAAFEELEKALSTA-----------------QKTEEARRKLKAEMDEQIKTIE 445
Query: 330 QTNEDEQADLCL-MAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECE---RLRNE 385
+T+E+E+ L +++ E V++ SS E+ L + ++ +E E +L+
Sbjct: 446 KTSEEERISLQQELSRVKQEVVDVMKKSSEEQIAKL--QKLHEKELARKEQELTKKLQTR 503
Query: 386 NLELRKEKEI-LEKENKDLIKNIQHSESKEVSENKNNSQKENIILKENVLKLKNDISNFV 444
E +++ ++ LEK + +K Q E +E + ++ IL E+ +N + +
Sbjct: 504 EREFQEQMKVALEKSQSEYLKISQEKEQQESLALEELELQKKAILTES----ENKLRDLQ 559
Query: 445 KSTETFQKIMGSQVSVFDKACIGFKTNQKQKLYENFFIPKKNKN-----TCERYKCTYCE 499
+ ET++ + S +K+ + NQ + L + K N E++K T E
Sbjct: 560 QEAETYRTRILELESSLEKS-LQENKNQSKDLAVHLEAEKNKHNKEITVMVEKHK-TELE 617
Query: 500 KYGHLEPFCFRKRKRLASQKFKNNYTNFHKK 530
H + + ++ ++ Q+++ +K
Sbjct: 618 SLKHQQDALWTEKLQVLKQQYQTEMEKLREK 648
Score = 42.0 bits (97), Expect = 0.013
Identities = 91/453 (20%), Positives = 187/453 (41%), Gaps = 53/453 (11%)
Query: 53 NEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSH 112
N++KK + +++ K M K KL A ++E + + KE K+ +
Sbjct: 951 NQEKKMEKVKQKAKEMQETLKKKLLDQEAKLKKELENTALELSQKEKQFNAKMLE--MAQ 1008
Query: 113 VKETRIDIGVRKFELFEMKETETIDEMYGRFT--II----------------MNELRSLE 154
I V + E + ++ E++ E++ R +I ++E++ E
Sbjct: 1009 ANSAGISDAVSRLETNQKEQIESLTEVHRRELNDVISIWEKKLNQQAEELQEIHEIQLQE 1068
Query: 155 KDFTIHERVRKILR--CLPRSWRHIVTAITEAKDLKKLRLEDLIGSLK---AHEVLLQED 209
K+ + E +KIL C +T + E + L +L LK AH L +D
Sbjct: 1069 KEQEVAELKQKILLFGCEKEEMNKEITWLKEEGVKQDTTLNELQEQLKQKSAHVNSLAQD 1128
Query: 210 KSSNKSKMIALKTNQESLNQELEMNFDNQQQELD----EEDHQDQIILLTRKL------- 258
++ K+ + L+ + LN+ L+ N Q+Q ++ E+ + ++ LT KL
Sbjct: 1129 ETKLKAHLEKLEVD---LNKSLKENTFLQEQLVELKMLAEEDKRKVSELTSKLKTTDEEF 1185
Query: 259 QRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKS-SNLQTNQ 317
Q + +++ ++ + K ++ + C K E N+ + S+ +TN
Sbjct: 1186 QSLKSSHEKSNKSLEDKSLEFKKLSEELAIQLDICCKKTEALLEAKTNELINISSSKTN- 1244
Query: 318 KSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHL-FDNLFYSSQMLE 376
+ ++ + +T + ++A L E L + + T ++ F + + E
Sbjct: 1245 -AILSRISHCQHRTTKVKEALLIKTCTVSELEAQLRQLTEEQNTLNISFQQATHQLEEKE 1303
Query: 377 QECERLRNENLELRKEKEILEKENKDLIKNIQHSES------KEVSENKNNSQKENIILK 430
+ + ++ + L EKE L+KE + + ES KE+SEN N ++K
Sbjct: 1304 NQIKSMKADIESLVTEKEALQKEGGNQQQAASEKESCITQLKKELSENIN----AVTLMK 1359
Query: 431 ENVLKLKNDISNFVKSTETFQKIMGSQVSVFDK 463
E + + K +IS+ K + + +S+ +K
Sbjct: 1360 EELKEKKVEISSLSKQLTDLNVQLQNSISLSEK 1392
Score = 40.8 bits (94), Expect = 0.030
Identities = 79/385 (20%), Positives = 149/385 (38%), Gaps = 54/385 (14%)
Query: 197 GSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDHQDQIILLTR 256
G K E+L Q K+ A + + LN+E E F NQ++++++ + +
Sbjct: 919 GQKKEIEILTQ--------KLSAKEDSIHILNEEYETKFKNQEKKMEKVKQK------AK 964
Query: 257 KLQRMIQRR--DQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQ 314
++Q ++++ DQ + +KE T L+ SQ E N + Q
Sbjct: 965 EMQETLKKKLLDQEAK---LKKELENTALELSQ-------------KEKQFNAKMLEMAQ 1008
Query: 315 TNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQM 374
N E E ++ + + N+ +++ ++ E L + Q
Sbjct: 1009 ANSAGISDAVSRLETNQKEQIESLTEVHRRELNDVISIWEKKLNQQAEELQEIHEIQLQE 1068
Query: 375 LEQECERLRNENLELRKEKEILEKE---------NKDLIKNIQHSESKEVSENKNNSQKE 425
EQE L+ + L EKE + KE +D N + K+ S + N+ ++
Sbjct: 1069 KEQEVAELKQKILLFGCEKEEMNKEITWLKEEGVKQDTTLNELQEQLKQKSAHVNSLAQD 1128
Query: 426 NIILKENVLKLKNDISNFVKSTETFQK--IMGSQVSVFDKACIGFKTNQKQKLYENFFIP 483
LK ++ KL+ D++ +K Q+ + ++ DK + T++ + E F
Sbjct: 1129 ETKLKAHLEKLEVDLNKSLKENTFLQEQLVELKMLAEEDKRKVSELTSKLKTTDEEFQSL 1188
Query: 484 K----KNKNTCERYKCTY---CEKYGHLEPFCFRKRKRLASQKFKNNYTNFHKKET---L 533
K K+ + E + E+ C +K + L K N N +T L
Sbjct: 1189 KSSHEKSNKSLEDKSLEFKKLSEELAIQLDICCKKTEALLEAK-TNELINISSSKTNAIL 1247
Query: 534 NNTNHQGPKKIWVPKALLTSTAGMS 558
+ +H + V +ALL T +S
Sbjct: 1248 SRISHCQHRTTKVKEALLIKTCTVS 1272
Score = 40.4 bits (93), Expect = 0.039
Identities = 66/380 (17%), Positives = 152/380 (39%), Gaps = 13/380 (3%)
Query: 55 KKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGT-SHV 113
K ED E E L K K +A +++ E K + + +GT SH+
Sbjct: 1623 KALEDRLESESAAKLAELKRKAEQKIAAIKKQLLSQMEEKEEQ--------YKKGTESHL 1674
Query: 114 KETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRS 173
E + R+ E+ ++E E T+I+ +T E C+ ++
Sbjct: 1675 SELNTKLQEREREVHILEEKLKSVESSQSETLIVPRSAKNVAAYTEQEEADS-QGCVQKT 1733
Query: 174 WRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEM 233
+ ++ + + +K +L +G K V + + + + E+ E +
Sbjct: 1734 YEEKISVL-QRNLTEKEKLLQRVGQEKEETVSSHFEMRCQYQERLIKLEHAEAKQHEDQS 1792
Query: 234 NFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGC 293
+ Q+EL+E++ + +I+ + + Q K+N ++ + L + ++TC
Sbjct: 1793 MIGHLQEELEEKNKKYSLIVAQHVEKEGGKNNIQAKQNLENVFDDVQKTLQEKELTCQIL 1852
Query: 294 YKLGHYKNECPLNKRKSSNLQTNQ-KSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNL 352
+ + C + +++ ++ + S ++ ++ +L + + +L
Sbjct: 1853 EQKIKELDSCLVRQKEVHRVEMEELTSKYEKLQALQQMDGRNKPTELLEENTEEKSKSHL 1912
Query: 353 DPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSES 412
E ++L + E+E ++L E + L+K+ +L KE++ ++ I E
Sbjct: 1913 VQPKLLSNMEAQHNDLEFKLAGAEREKQKLGKEIVRLQKDLRMLRKEHQQELE-ILKKEY 1971
Query: 413 KEVSENKNNSQKENIILKEN 432
+ E K ++E++ LK N
Sbjct: 1972 DQEREEKIKQEQEDLELKHN 1991
Score = 39.7 bits (91), Expect = 0.066
Identities = 63/308 (20%), Positives = 131/308 (42%), Gaps = 72/308 (23%)
Query: 214 KSKMIALKTNQESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFP 273
K K + T ++ +EL++ + + +E E+D +QI LL +L DQ + F
Sbjct: 1446 KKKAQSRFTQHQNTVKELQIQLELKSKEAYEKD--EQINLLKEEL-------DQQNKRFD 1496
Query: 274 ARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKS----FMTTWDEPEE 329
K + E DKS++ ++K SNL+T KS M D +
Sbjct: 1497 CLK--GEMEDDKSKM------------------EKKESNLETELKSQTARIMELEDHITQ 1536
Query: 330 QTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHL--------------------FDNLF 369
+T E E + L K+ N++ +++ +K +H +N
Sbjct: 1537 KTIEIESLNEVL--KNYNQQKDIEHKELVQKLQHFQELGEEKDNRVKEAEEKILTLENQV 1594
Query: 370 YSSQM-LEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENII 428
YS + LE + + L + NL ++ + E+E K L ++ + +++E K ++++
Sbjct: 1595 YSMKAELETKKKELEHVNLSVKSK----EEELKALEDRLESESAAKLAELKRKAEQKIAA 1650
Query: 429 LKENVL-KLKNDISNFVKSTETFQKIMGS-------QVSVFDKACIGFKTNQKQKLYENF 480
+K+ +L +++ + K TE+ + + +V + ++ +++Q E
Sbjct: 1651 IKKQLLSQMEEKEEQYKKGTESHLSELNTKLQEREREVHILEEKLKSVESSQS----ETL 1706
Query: 481 FIPKKNKN 488
+P+ KN
Sbjct: 1707 IVPRSAKN 1714
Score = 38.9 bits (89), Expect = 0.11
Identities = 93/487 (19%), Positives = 190/487 (38%), Gaps = 84/487 (17%)
Query: 62 EEEGKRMLLNFKAKLFLTMALSREEY---DRVQECKNAKEIWDTLKVHHEGTSH---VKE 115
+EE +L + F LS+E+ ++V + N W K T H VKE
Sbjct: 1404 DEEKCELLDQVQDLSFKVDTLSKEKISALEQVDDWSNKFSEWKK-KAQSRFTQHQNTVKE 1462
Query: 116 TRIDIGVRKFELFEMKET-----ETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCL 170
+I + ++ E +E E E +D+ RF + E+ E D + E+ L
Sbjct: 1463 LQIQLELKSKEAYEKDEQINLLKEELDQQNKRFDCLKGEM---EDDKSKMEKKESNLETE 1519
Query: 171 PRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLN-- 228
+S + + + K + +E L +EVL K+ N+ K I K + L
Sbjct: 1520 LKSQTARIMELEDHITQKTIEIESL------NEVL----KNYNQQKDIEHKELVQKLQHF 1569
Query: 229 QELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQV 288
QEL DN+ +E +E+ + + + K + ++++ N + + + + + ++
Sbjct: 1570 QELGEEKDNRVKEAEEKILTLENQVYSMKAELETKKKELEHVNLSVKSKEEELKALEDRL 1629
Query: 289 TCYGCYKLGHYKNECP-----LNKRKSSNLQTNQKSFM-----------TTWDEPEEQTN 332
KL K + + K+ S ++ ++ + T E E + +
Sbjct: 1630 ESESAAKLAELKRKAEQKIAAIKKQLLSQMEEKEEQYKKGTESHLSELNTKLQEREREVH 1689
Query: 333 ---------EDEQADLCLMAKS--------DNEEVNLDPCSSC---EKTEHLFDNLFYSS 372
E Q++ ++ +S + EE + C EK L NL
Sbjct: 1690 ILEEKLKSVESSQSETLIVPRSAKNVAAYTEQEEADSQGCVQKTYEEKISVLQRNLTEKE 1749
Query: 373 QMLE---QECERLRNENLELR----------KEKEILEKENKDLIKNIQHS---ESKEVS 416
++L+ QE E + + E+R + E + E++ +I ++Q ++K+ S
Sbjct: 1750 KLLQRVGQEKEETVSSHFEMRCQYQERLIKLEHAEAKQHEDQSMIGHLQEELEEKNKKYS 1809
Query: 417 -----ENKNNSQKENIILKENVLKLKNDISNFVKSTETFQKIMGSQVSVFDKACIGFKTN 471
+ K NI K+N+ + +D+ ++ E +I+ ++ D + K
Sbjct: 1810 LIVAQHVEKEGGKNNIQAKQNLENVFDDVQKTLQEKELTCQILEQKIKELDSCLVRQKEV 1869
Query: 472 QKQKLYE 478
+ ++ E
Sbjct: 1870 HRVEMEE 1876
Score = 38.5 bits (88), Expect = 0.15
Identities = 75/351 (21%), Positives = 149/351 (42%), Gaps = 54/351 (15%)
Query: 105 VHHEGTSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVR 164
V +GTS + V++ E + ETI + T++ +E +L++ + ER++
Sbjct: 271 VVEDGTSVKTLETLQQRVKRQENLLKRCKETIQSHKEQCTLLTSEKEALQEQ--LDERLQ 328
Query: 165 KILRCLPRSWRHIVTAITEAKDLKKL--RLEDLIGSLKA------HEVL--LQEDKSSNK 214
++ + IT+ +D K L +LE G + A HE L +E+ + +
Sbjct: 329 ELEKIKDLHMAEKTKLITQLRDAKNLIEQLEQDKGMVIAETKRQMHETLEMKEEEIAQLR 388
Query: 215 SKMIALKTNQESLNQELEMNFDNQQQELD------------------EEDHQDQIILLTR 256
S++ + T E L ++ E + +EL+ E D Q + I T
Sbjct: 389 SRIKQMTTQGEELREQKEKSERAAFEELEKALSTAQKTEEARRKLKAEMDEQIKTIEKTS 448
Query: 257 KLQRMIQRRDQNKRN---FPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNL 313
+ +R+ +++ ++ K++++ ++ K Q KL H K + + L
Sbjct: 449 EEERISLQQELSRVKQEVVDVMKKSSEEQIAKLQ-------KL-HEKELARKEQELTKKL 500
Query: 314 QTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQ 373
QT ++ F E + E Q++ +++ ++ +L + E+ E + S+
Sbjct: 501 QTREREFQ----EQMKVALEKSQSEYLKISQEKEQQESL----ALEELELQKKAILTESE 552
Query: 374 M----LEQECERLRNENLELRKEKEILEKENKDLIKNIQ-HSESKEVSENK 419
L+QE E R LEL E +ENK+ K++ H E+++ NK
Sbjct: 553 NKLRDLQQEAETYRTRILELESSLEKSLQENKNQSKDLAVHLEAEKNKHNK 603
>REST_CHICK (O42184) Restin (Cytoplasmic linker protein-170)
(CLIP-170)
Length = 1433
Score = 52.8 bits (125), Expect = 8e-06
Identities = 90/426 (21%), Positives = 184/426 (43%), Gaps = 64/426 (15%)
Query: 54 EKKKEDWSEEEGKRM-LLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSH 112
EK E +GK LL+ + L A+++ + +E + KE + + E
Sbjct: 800 EKVSNLTKELQGKEQKLLDLEKNL---SAVNQVKDSLEKELQLLKEKFTSAVDGAENAQR 856
Query: 113 VKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEK--------DFTIHERVR 164
+ I+ +K E F + +E ++++ T++ +L+ E+ + +
Sbjct: 857 AMQETINKLNQKEEQFALMSSE-LEQLKSNLTVMETKLKEREEREQQLTEAKVKLENDIA 915
Query: 165 KILRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSL-KAHEVLLQEDKS-------SNKSK 216
+I++ S ++ E + LK+ +LE + L KA+E +Q K+ + +S+
Sbjct: 916 EIMKSSGDSSAQLMKMNDELR-LKERQLEQIQLELTKANEKAVQLQKNVEQTAQKAEQSQ 974
Query: 217 MIALKTNQESLNQELEMNFD-NQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPAR 275
LKT+QE L + + D +Q E + ++D ++ MI + D + + F
Sbjct: 975 QETLKTHQEELKKMQDQLTDMKKQMETSQNQYKDLQAKYEKETSEMITKHDADIKGFKQN 1034
Query: 276 KENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDE 335
+A+ L +Q +K+ L+T + + EQ D+
Sbjct: 1035 LLDAEEALKAAQ--------------------KKNDELETQAEEL----KKQAEQAKADK 1070
Query: 336 QAD--LCLMAKSDNEEVNLDPCSSCEKTEHL--FDNLFYSSQMLEQECERLRNENL---- 387
+A+ L M K E+ + EK E L +N +++ L+ E + L+ NL
Sbjct: 1071 RAEEVLQTMEKVTKEKDAIHQ----EKIETLASLENSRQTNEKLQNELDMLKQNNLKNEE 1126
Query: 388 ELRKEKEILEKENKDLIKNIQHSESKEVSENKNNS-----QKENIILKENVLKLKNDISN 442
EL K KE+L ENK + + + E+ +++ + + Q+EN+ L E + + ++++++
Sbjct: 1127 ELTKSKELLNLENKKVEELKKEFEALKLAAAQKSQQLAALQEENVKLAEELGRSRDEVTS 1186
Query: 443 FVKSTE 448
K E
Sbjct: 1187 HQKLEE 1192
Score = 47.8 bits (112), Expect = 2e-04
Identities = 95/442 (21%), Positives = 181/442 (40%), Gaps = 96/442 (21%)
Query: 87 YDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEMK----ETETIDEMYGR 142
+++ + K +E+ DT S + E D+ +R E+ E++ ++ ID++
Sbjct: 476 FEKTKADKLQRELEDTRVATVSEKSRIMELERDLALRVKEVAELRGRLESSKHIDDVDTS 535
Query: 143 FTIIMN-----------------ELRSLEKDF-TIHERVRKILRCLPRSWRHIVT----- 179
+++ E+ SL++ F + E +RK ++ L S +
Sbjct: 536 LSLLQEISSLQEKMAAAGKEHQREMSSLKEKFESSEEALRKEIKTLSASNERMGKENESL 595
Query: 180 ------AITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSS-NK------SKMIALKTNQES 226
A E D+ +L L ++ +H+ ++E K S NK ++ LKT E
Sbjct: 596 KTKLDHANKENSDVIELWKSKLESAIASHQQAMEELKVSFNKGVGAQTAEFAELKTQMEK 655
Query: 227 --LNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTE-- 282
L+ E EM+ +QE ++ H +I L KL + + ++Q N A+ E+ + +
Sbjct: 656 VKLDYENEMSNLKLKQENEKSQHLKEIEALKAKLLEVTEEKEQTLENLKAKLESVEDQHL 715
Query: 283 ------LDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQ 336
L+K Q +L + +C N QT +T + + +E++
Sbjct: 716 VEMEDTLNKLQEAEIKVKELDVLQAKC--------NEQTKLIGSLT----QQIRASEEKL 763
Query: 337 ADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENL-ELRKEKEI 395
DL + K+ N E L+ L Q E++ + L E + L KE +
Sbjct: 764 LDLAALQKA-NSEGKLE-----------IQKLSEQLQAAEKQIQNLETEKVSNLTKELQG 811
Query: 396 LEKENKDLIKNIQHSESKEVSENKNNSQKENIILKENVLKLKNDISNFVKSTETFQKIMG 455
E++ DL KN+ V++ K++ +KE +LKE ++ V E Q+ M
Sbjct: 812 KEQKLLDLEKNL-----SAVNQVKDSLEKELQLLKEK-------FTSAVDGAENAQRAMQ 859
Query: 456 SQVSVFDKACIGFKTNQKQKLY 477
++ K NQK++ +
Sbjct: 860 ETIN---------KLNQKEEQF 872
Score = 43.5 bits (101), Expect = 0.005
Identities = 67/291 (23%), Positives = 124/291 (42%), Gaps = 29/291 (9%)
Query: 158 TIHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKM 217
T E ++K+ L + + T+ + KDL+ ++ + H+ ++ K +
Sbjct: 980 THQEELKKMQDQLTDMKKQMETSQNQYKDLQAKYEKETSEMITKHDADIKGFKQNLLDAE 1039
Query: 218 IALKTNQESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRD---QNKRNFPA 274
ALK Q+ N ELE + +++ ++ + + + ++++ + +D Q K A
Sbjct: 1040 EALKAAQKK-NDELETQAEELKKQAEQAKADKRAEEVLQTMEKVTKEKDAIHQEKIETLA 1098
Query: 275 RKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNED 334
EN++ +K Q K + KNE L K K N+K EE E
Sbjct: 1099 SLENSRQTNEKLQNEL-DMLKQNNLKNEEELTKSKELLNLENKKV--------EELKKEF 1149
Query: 335 EQADLCLMAKSDN----EEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELR 390
E L KS +E N+ ++ + S Q LE+E L N+ LE++
Sbjct: 1150 EALKLAAAQKSQQLAALQEENVKLAEELGRSR----DEVTSHQKLEEERSVLNNQLLEMK 1205
Query: 391 KEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENIILKENVLKLKNDIS 441
K + L+KE + ++Q S S + +QK+ E + KL+N+I+
Sbjct: 1206 KRESTLKKEIDEERASLQKSIS---DTSALITQKD-----EELEKLRNEIT 1248
Score = 37.4 bits (85), Expect = 0.33
Identities = 76/364 (20%), Positives = 140/364 (37%), Gaps = 38/364 (10%)
Query: 150 LRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQED 209
L SLE +E+++ L L ++ +T++K+L L + + LK L+
Sbjct: 1097 LASLENSRQTNEKLQNELDMLKQNNLKNEEELTKSKELLNLENKK-VEELKKEFEALKLA 1155
Query: 210 KSSNKSKMIALKTNQESLNQELEMNFDN--QQQELDEEDHQDQIILLTRKLQRMIQRRDQ 267
+ ++ AL+ L +EL + D Q+L+EE +L +L M +R
Sbjct: 1156 AAQKSQQLAALQEENVKLAEELGRSRDEVTSHQKLEEERS-----VLNNQLLEMKKREST 1210
Query: 268 NKRNFPARKENAKTEL-DKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDE 326
K+ + + + + D S + +L +NE + + ++++ +T Q T +
Sbjct: 1211 LKKEIDEERASLQKSISDTSALITQKDEELEKLRNEITVLRGENASAKTLQSVVKTLESD 1270
Query: 327 PEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFY--SSQMLEQECERLRN 384
+ + + + L AKS+ P + + + NL S++ +QE + L +
Sbjct: 1271 KLKLEEKVKNLEQKLKAKSEQ------PLTVTSPSGDIAANLLQDESAEDKQQEIDFLNS 1324
Query: 385 ENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENIILKENVLKLKNDISNFV 444
++L++ E L N++ E + N N + N +E L K
Sbjct: 1325 VIVDLQRRNEEL---------NLKIQRMCEAALNGNEEETINYDSEEEGLSKK------- 1368
Query: 445 KSTETFQKIMGSQVSVFDKACIGFKTNQKQKLYENFFIPKKNKNTCERYKCTYCEKYGHL 504
+ F I G FD Q Q L E ER C CE +GH
Sbjct: 1369 -TPRLFCDICGC----FDLHDTEDCPTQAQMLEEPPHSTYHGSRREERPYCDTCEMFGHW 1423
Query: 505 EPFC 508
C
Sbjct: 1424 TADC 1427
>TPR_HUMAN (P12270) Nucleoprotein TPR
Length = 2349
Score = 51.6 bits (122), Expect = 2e-05
Identities = 77/375 (20%), Positives = 163/375 (42%), Gaps = 51/375 (13%)
Query: 97 KEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKD 156
KE + K+ +E ++E D+ + ++ + +D R+ ++ + + ++
Sbjct: 684 KEKAENEKIQNEQLEKLQEQVTDLRSQNTKI-----STQLDFASKRYEMLQDNVEGYRRE 738
Query: 157 FT-IHERVRKILRCLPRSWRHIVTAITE---------------AKDLKK----LRLEDLI 196
T +HER +K L + I+ +T+ A++LKK L+L ++
Sbjct: 739 ITSLHERNQK-LTATTQKQEQIINTMTQDLRGANEKLAVAEVRAENLKKEKEMLKLSEVR 797
Query: 197 GSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDE--EDHQDQIILL 254
S + E LL E + N L TN +++ LE + +Q L E + +I L
Sbjct: 798 LS-QQRESLLAEQRGQN-----LLLTNLQTIQGILERSETETKQRLSSQIEKLEHEISHL 851
Query: 255 TRKLQRMIQRRDQNKRNFPARKENAKTELD-KSQVTCYGCYKLGHYKNECPLNKRKSSNL 313
+KL+ +++R RN + + K +LD ++ + L + + E K+ SN+
Sbjct: 852 KKKLENEVEQRHTLTRNLDVQLLDTKRQLDTETNLHLNTKELLKNAQKEIATLKQHLSNM 911
Query: 314 QTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQ 373
+ S + + +N+++ DL + E+VN L + L S+
Sbjct: 912 EVQVASQSSQRTGKGQPSNKEDVDDLVSQLRQTEEQVN-----------DLKERLKTSTS 960
Query: 374 MLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENIILKENV 433
+EQ + + L KEK++ E+ K++ + KE +E + +K+ + +++
Sbjct: 961 NVEQYQAMVTSLEESLNKEKQVTEEVRKNIEVRL-----KESAEFQTQLEKKLMEVEKEK 1015
Query: 434 LKLKNDISNFVKSTE 448
+L++D ++S E
Sbjct: 1016 QELQDDKRRAIESME 1030
Score = 37.4 bits (85), Expect = 0.33
Identities = 76/328 (23%), Positives = 129/328 (39%), Gaps = 52/328 (15%)
Query: 124 KFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITE 183
K E F + ID + GR ++ S ++ F I +R L S +V E
Sbjct: 24 KLEKFLADQQSEIDGLKGRHEKF--KVESEQQYFEIEKR-------LSHSQERLVNETRE 74
Query: 184 AKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELD 243
+ L+ L LE L LKA L E NK IA + N ++ F ++EL+
Sbjct: 75 CQSLR-LELEKLNNQLKA----LTE---KNKELEIA-----QDRNIAIQSQFTRTKEELE 121
Query: 244 EEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNEC 303
E +I +L + ++ ++ + + + + T + Q+ KL +
Sbjct: 122 AEKRD--LIRTNERLSQELEYLTEDVKRLNEKLKESNTTKGELQL------KLDELQASD 173
Query: 304 PLNKRKSSNLQTNQKSF--MTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKT 361
K + L+ ++ TW E +T DE L L + NE + L C+ K
Sbjct: 174 VSVKYREKRLEQEKELLHSQNTWLNTELKTKTDEL--LALGREKGNEILELK-CNLENKK 230
Query: 362 EHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVS-ENKN 420
E E RL + L+ E L+K +DL+ ++ ++ ++ S E K
Sbjct: 231 E---------------EVSRLEEQMNGLKTSNEHLQKHVEDLLTKLKEAKEQQASMEEKF 275
Query: 421 NSQKENIILKENVLKLKNDISNFVKSTE 448
+++ I N+ K D S KS E
Sbjct: 276 HNELNAHIKLSNLYKSAADDSE-AKSNE 302
>MYH7_MESAU (P13540) Myosin heavy chain, cardiac muscle beta isoform
(MyHC-beta)
Length = 1934
Score = 50.8 bits (120), Expect = 3e-05
Identities = 88/440 (20%), Positives = 173/440 (39%), Gaps = 50/440 (11%)
Query: 53 NEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECK-NAKEIWDTLKVHHEGTS 111
N+K++ D ++ L A++ AL + +++E + +E+ + L+ +
Sbjct: 1070 NDKQQLDEKLKKKDFELNALNARIEDEQALGSQLQKKLKELQARIEELEEELEAERTARA 1129
Query: 112 HVKETRIDIGVRKFELFEMKE------TETIDEMYGRFTIIMNELRSLEKDFTIHERVRK 165
V++ R D+ E+ E E + I+ R R LE+ HE
Sbjct: 1130 KVEKLRSDLSRELEEISERLEEAGGATSVQIEMNKKREAEFQKMRRDLEEATLQHEATAA 1189
Query: 166 ILRCLPRSWRHIVTAITEAKDLKKL-RLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQ 224
LR +H + + + L R++ + K+ L +D +SN ++I K N
Sbjct: 1190 ALRK-----KHADSVAELGEQIDNLQRVKQKLEKEKSEFKLELDDVTSNMEQIIKAKANL 1244
Query: 225 ESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELD 284
E + + LE + + + +E Q + LT + ++ + R KE ++L
Sbjct: 1245 EKMCRTLEDQMNEHRSKAEET--QRSVNDLTSQRAKLQTENGELSRQLD-EKEALISQLT 1301
Query: 285 KSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADL---CL 341
+ ++T +L K + + + L +S D EQ E+ +A C+
Sbjct: 1302 RGKLTY--TQQLEDLKRQLEEEVKAKNTLAHALQSARHDCDLLREQYEEETEAKAELQCV 1359
Query: 342 MAKSDNE------EVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEI 395
++K+++E + D E+ E L Q E+ E + + L K K
Sbjct: 1360 LSKANSEVAQWRTKYETDAIQRTEELEEAKKKLAQRLQDAEEAVEAVNAKCSSLEKTKHR 1419
Query: 396 LEKENKDLIKNIQHSESKEVSENKN-----------------------NSQKENIILKEN 432
L+ E +DL+ +++ S + + +K +SQKE L
Sbjct: 1420 LQNEIEDLMVDVERSNAAAAALDKKQRNFDKILAEWKQKYEESQSELESSQKEARSLSTE 1479
Query: 433 VLKLKNDISNFVKSTETFQK 452
+ KLKN ++ ETF++
Sbjct: 1480 LFKLKNAYEESLEHLETFKR 1499
Score = 35.4 bits (80), Expect = 1.2
Identities = 69/315 (21%), Positives = 126/315 (39%), Gaps = 56/315 (17%)
Query: 178 VTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDN 237
V +T++K + +++DL GSL+ + + + + + + LK QES+ M+ +N
Sbjct: 1016 VNTLTKSKVKLEQQVDDLEGSLEQEKKVRMDLERAKRKLEGDLKLTQESI-----MDLEN 1070
Query: 238 QQQELDE----------------EDHQ---DQIILLTRKLQRMIQRRDQN---KRNFPAR 275
+Q+LDE ED Q Q+ ++LQ I+ ++ +R A+
Sbjct: 1071 DKQQLDEKLKKKDFELNALNARIEDEQALGSQLQKKLKELQARIEELEEELEAERTARAK 1130
Query: 276 KENAKTELDK--SQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNE 333
E +++L + +++ G + +NK++ + Q ++ EE T +
Sbjct: 1131 VEKLRSDLSRELEEISERLEEAGGATSVQIEMNKKREAEFQKMRRDL-------EEATLQ 1183
Query: 334 DEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNE------NL 387
E L K S + DNL Q LE+E + E N+
Sbjct: 1184 HEATAAALRKKH---------ADSVAELGEQIDNLQRVKQKLEKEKSEFKLELDDVTSNM 1234
Query: 388 E-LRKEKEILEKENKDLIKNIQHSESK----EVSENKNNSQKENIILKENVLKLKNDISN 442
E + K K LEK + L + SK + S N SQ+ + + L + D
Sbjct: 1235 EQIIKAKANLEKMCRTLEDQMNEHRSKAEETQRSVNDLTSQRAKLQTENGELSRQLDEKE 1294
Query: 443 FVKSTETFQKIMGSQ 457
+ S T K+ +Q
Sbjct: 1295 ALISQLTRGKLTYTQ 1309
Score = 34.3 bits (77), Expect = 2.8
Identities = 49/204 (24%), Positives = 88/204 (43%), Gaps = 35/204 (17%)
Query: 69 LLNFKAKLFLTMA-LSREEYDRVQECKNAKE-----------IWDTLKVHHEGTSHVKET 116
L+N K K+ ++ L E + VQEC+NA+E + + LK + ++H++
Sbjct: 1722 LINQKKKMDADLSQLQTEVEEAVQECRNAEEKAKKAITDAAMMAEELKKEQDTSAHLERM 1781
Query: 117 RIDIGVRKFEL-FEMKETETIDEMYGRFTI--IMNELRSLEKDFTIHE-RVRKILRCLPR 172
+ ++ +L + E E I G+ + + +R LE + + R + ++ + +
Sbjct: 1782 KKNMEQTIKDLQHRLDEAEQIALKGGKKQLQKLEARVRELENELEAEQKRNAESVKGMRK 1841
Query: 173 SWRHI--VTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQE 230
S R I +T TE LRL+DL+ L+ K+ A K E ++
Sbjct: 1842 SERRIKELTYQTEEDRKNLLRLQDLVDKLQL--------------KVKAYKRQAEEAEEQ 1887
Query: 231 LEMN---FDNQQQELDEEDHQDQI 251
N F Q ELDE + + I
Sbjct: 1888 ANTNLSKFRKVQHELDEAEERADI 1911
Score = 33.1 bits (74), Expect = 6.2
Identities = 58/273 (21%), Positives = 112/273 (40%), Gaps = 31/273 (11%)
Query: 191 RLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDHQDQ 250
R++D + +A L+E KM++L + L +++ DN D E+ DQ
Sbjct: 857 RVKDALEKSEARRKELEE-------KMVSLLQEKNDLQLQVQAEQDNLA---DAEERCDQ 906
Query: 251 IIL----LTRKLQRMIQR-RDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPL 305
+I L K++ M +R D+ + N + K E + S++ K E L
Sbjct: 907 LIKNKIQLEAKVKEMTERLEDEEEMNAELTAKKRKLEDECSEL------KRDIDDLELTL 960
Query: 306 NKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLF 365
K + T K T EE DE K +E + + E
Sbjct: 961 AKVEKDKHATENKVKNLT----EEMAGLDEIIAKLTKEKKALQEAHQQALDDLQAEEDKV 1016
Query: 366 DNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKE 425
+ L S LEQ+ + L + +K + LE+ + L +++ ++ + + + +N+ Q+
Sbjct: 1017 NTLTKSKVKLEQQVDDLEGSLEQEKKVRMDLERAKRKLEGDLKLTQ-ESIMDLENDKQQL 1075
Query: 426 NIILKENVLKLKNDISNFVKSTETFQKIMGSQV 458
+ LK+ +L N + + ++ +GSQ+
Sbjct: 1076 DEKLKKKDFEL-----NALNARIEDEQALGSQL 1103
>MYH4_HUMAN (Q9Y623) Myosin heavy chain, skeletal muscle, fetal
(Myosin heavy chain IIb) (MyHC-IIb)
Length = 1939
Score = 50.8 bits (120), Expect = 3e-05
Identities = 85/444 (19%), Positives = 174/444 (39%), Gaps = 58/444 (13%)
Query: 50 EDLNEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECK-NAKEIWDTLKVHHE 108
+ LNEK K+ E + N + K+ AL+ + +++E + +E+ + ++
Sbjct: 1078 QQLNEKLKKKEFE------MSNLQGKIEDEQALAMQLQKKIKELQARIEELEEEIEAERA 1131
Query: 109 GTSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILR 168
+ ++ R D+ E+ +E ++E G + + + E +F +++R+ L
Sbjct: 1132 SRAKAEKQRSDLSRELEEI-----SERLEEAGGATSAQIELNKKREAEF---QKMRRDLE 1183
Query: 169 CLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLN 228
A+ + L I SL+ + L+++KS K ++ L +N E+++
Sbjct: 1184 ESTLQHEATAAALRKKHADSVAELGKQIDSLQRVKQKLEKEKSELKMEINDLASNMETVS 1243
Query: 229 QELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQV 288
+ + NF+ + L ED +I + QR+I K +LD+
Sbjct: 1244 KA-KANFEKMCRTL--EDQLSEIKTKEEEQQRLINELSAQKARLHTESGEFSRQLDEKDA 1300
Query: 289 TCYGCYK--------LGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLC 340
+ + K + + S L +S D EQ E+++A
Sbjct: 1301 MVSQLSRGKQAFTQQIEELKRQLEEETKAKSTLAHALQSARHDCDLLREQYEEEQEAKAE 1360
Query: 341 L---MAKSDNE------EVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRK 391
L M+K+++E + D E+ E L Q E+ E + ++ L K
Sbjct: 1361 LQRGMSKANSEVAQWRTKYETDAIQRTEELEEAKKKLAQRLQDAEEHVEAVNSKCASLEK 1420
Query: 392 EKEILEKENKDLIKNIQHSESKEVSENKNN-----------------------SQKENII 428
K+ L+ E +DL+ +++ S + ++ +K SQKE+
Sbjct: 1421 TKQRLQNEVEDLMIDVERSNAACIALDKKQRNFDKVLAEWKQKYEETQAELEASQKESRS 1480
Query: 429 LKENVLKLKNDISNFVKSTETFQK 452
L + K+KN + ET ++
Sbjct: 1481 LSTELFKVKNAYEESLDHLETLKR 1504
Score = 42.7 bits (99), Expect = 0.008
Identities = 62/282 (21%), Positives = 113/282 (39%), Gaps = 54/282 (19%)
Query: 178 VTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDN 237
V +T+AK + +++DL GSL+ + L + + + + LK QES M+ +N
Sbjct: 1021 VNTLTKAKTKLEQQVDDLEGSLEQEKKLCMDLERAKRKLEGDLKLAQES-----TMDTEN 1075
Query: 238 QQQELDE----------------EDHQDQIILLTRKLQRMIQRRDQNKRNFPA-RKENAK 280
+Q+L+E ED Q + L +K++ + R ++ + A R AK
Sbjct: 1076 DKQQLNEKLKKKEFEMSNLQGKIEDEQALAMQLQKKIKELQARIEELEEEIEAERASRAK 1135
Query: 281 TELDKSQVTCYGCYKL-----------GHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEE 329
E +S ++ +L G + LNK++ + Q ++ EE
Sbjct: 1136 AEKQRSDLS----RELEEISERLEEAGGATSAQIELNKKREAEFQKMRRDL-------EE 1184
Query: 330 QTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLEL 389
T + E L K S + D+L Q LE+E L+ E +L
Sbjct: 1185 STLQHEATAAALRKKH---------ADSVAELGKQIDSLQRVKQKLEKEKSELKMEINDL 1235
Query: 390 RKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENIILKE 431
E + K + K + E ++SE K +++ ++ E
Sbjct: 1236 ASNMETVSKAKANFEKMCRTLED-QLSEIKTKEEEQQRLINE 1276
Score = 37.0 bits (84), Expect = 0.43
Identities = 89/456 (19%), Positives = 178/456 (38%), Gaps = 50/456 (10%)
Query: 54 EKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHV 113
EK KE+ ++ E KR L K M +E + +Q A+ D L E +
Sbjct: 861 EKTKEELAKTEAKRKELEEK------MVTLMQEKNDLQLQVQAEA--DALADAEERCDQL 912
Query: 114 KETRIDIGVRKFELFEMKETETIDEMYGRFTI----IMNELRSLEKDFTIHERVRKILRC 169
+T+I + + E+ E E E +E+ T + +E L+KD E +
Sbjct: 913 IKTKIQLEAKIKEVTERAEDE--EEINAELTAKKRKLEDECSELKKDIDDLELTLAKVEK 970
Query: 170 LPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQ 229
+ + V +TE + L++ I L + LQE +++ + L+ ++ +N
Sbjct: 971 EKHATENKVKNLTE----EMAGLDETIAKLTKEKKALQE---AHQQTLDDLQMEEDKVNT 1023
Query: 230 ELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVT 289
+ +QQ D E +Q L L+R ++ + + + A++ TE DK Q+
Sbjct: 1024 LTKAKTKLEQQVDDLEGSLEQEKKLCMDLERAKRKLEGDLKL--AQESTMDTENDKQQLN 1081
Query: 290 ---CYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSD 346
+++ + + + + + LQ K +E EE+ +A+ AK++
Sbjct: 1082 EKLKKKEFEMSNLQGKIEDEQALAMQLQKKIKELQARIEELEEEI----EAERASRAKAE 1137
Query: 347 NEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRK-----EKEILEKENK 401
+ +L + E + + L + + E + E +K E+ L+ E
Sbjct: 1138 KQRSDLS-----RELEEISERLEEAGGATSAQIELNKKREAEFQKMRRDLEESTLQHEAT 1192
Query: 402 DLIKNIQHSES--------KEVSENKNNSQKENIILKENVLKLKNDISNFVKSTETFQKI 453
+H++S + K +KE LK + L +++ K+ F+K+
Sbjct: 1193 AAALRKKHADSVAELGKQIDSLQRVKQKLEKEKSELKMEINDLASNMETVSKAKANFEKM 1252
Query: 454 MGSQVSVFDKACIGFKTNQKQKLYENFFIPKKNKNT 489
+ + I K ++Q+L K +T
Sbjct: 1253 CRTLEDQLSE--IKTKEEEQQRLINELSAQKARLHT 1286
Score = 33.9 bits (76), Expect = 3.6
Identities = 59/300 (19%), Positives = 117/300 (38%), Gaps = 34/300 (11%)
Query: 154 EKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSN 213
E F I +R + W + I LK E + ++K +E+ +
Sbjct: 813 ESIFCIQYNIRAFMNVKHWPWMKLYFKIKPL--LKSAETEKEMANMKEEFEKTKEELAKT 870
Query: 214 KSKMIALKTNQESLNQE---LEMNFDNQQQEL-DEEDHQDQIILLTRKLQRMIQRRDQNK 269
++K L+ +L QE L++ + L D E+ DQ+I +L+ I
Sbjct: 871 EAKRKELEEKMVTLMQEKNDLQLQVQAEADALADAEERCDQLIKTKIQLEAKI------- 923
Query: 270 RNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTN----QKSFMTTWD 325
KE + D+ ++ K ++EC K+ +L+ +K T +
Sbjct: 924 ------KEVTERAEDEEEINAELTAKKRKLEDECSELKKDIDDLELTLAKVEKEKHATEN 977
Query: 326 EPEEQTNE----DEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECER 381
+ + T E DE K +E + + E + L + LEQ+ +
Sbjct: 978 KVKNLTEEMAGLDETIAKLTKEKKALQEAHQQTLDDLQMEEDKVNTLTKAKTKLEQQVDD 1037
Query: 382 LRNENLELRKEKEI---LEKENKDLIKNIQHSESKEVSENKNNSQKENIILKENVLKLKN 438
L L +EK++ LE+ + L +++ ++ + + +N+ Q+ N LK+ ++ N
Sbjct: 1038 LEG---SLEQEKKLCMDLERAKRKLEGDLKLAQ-ESTMDTENDKQQLNEKLKKKEFEMSN 1093
>C5P2_MACFA (Q9BE52) CDK5 regulatory subunit associated protein 2
(CDK5 activator-binding protein C48) (QflA-15432)
Length = 862
Score = 50.1 bits (118), Expect = 5e-05
Identities = 87/436 (19%), Positives = 179/436 (40%), Gaps = 55/436 (12%)
Query: 49 DEDLNEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHE 108
+EDL EK++E +E+ K L KA LTMAL +E K +E+ ++
Sbjct: 296 EEDLREKEREIATEK--KNSLKRDKAIQGLTMALKSKE-------KKVEELNSEIEKLSA 346
Query: 109 GTSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRS--LEKDFTIHERVRKI 166
+ +E ++K + E + +E + ++ ELRS L K H R+R+
Sbjct: 347 AFAKAREA-----LQKAQTQEFQGSENYEAALSGKEALLTELRSQNLTKSAENH-RLRRS 400
Query: 167 LRCLPRSWRHIVTAITEA-KDLKKLRLEDLIGSLKAHEVLLQEDKSSNK--SKMIALKTN 223
++ + + + KDL++ E G ++ + +K N+ + A++
Sbjct: 401 IKKITQELSDLQQERERLEKDLEEAHREKSRGDCTIRDLRNEVEKLRNEVNERKKAMENR 460
Query: 224 QESLNQELEMNFDNQQQEL----DEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENA 279
++L E NQ+Q + + +H+D +LL + ++ ++ QN A
Sbjct: 461 YKNLLSESSKKLHNQEQVIKHLTERTNHKD--MLLQKFNEKDLEVIQQNHYLMTAEDLEL 518
Query: 280 KTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADL 339
++E G +CP + S T + + E+Q+++ ++
Sbjct: 519 RSE--------------GLITEKCPSQQSPGSK---------TIFSKEEKQSSDYQELIQ 555
Query: 340 CLMAKSD---NEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKE-KEI 395
L + D + +L S + +N+F + LE++ +N L ++ EI
Sbjct: 556 VLKKEQDIYTHLVKSLQESDSINNLQAELNNIFALRKQLERDVLSYQNLRRTLEEQISEI 615
Query: 396 LEKENKDLIKNIQHSESKEVSENKNNSQKENIILKENVLKLKNDISNFVKSTETFQKIMG 455
+E + + + +NN + +E + K +D+ VK T + +
Sbjct: 616 RRREEESFSFYSDQTSYLSICLEENNRFQVEHFSQEELKKKVSDLIQLVKELYTDDQHL- 674
Query: 456 SQVSVFDKACIGFKTN 471
+ ++FD +C+GF+ N
Sbjct: 675 -KKTIFDLSCMGFQGN 689
>MYH6_MOUSE (Q02566) Myosin heavy chain, cardiac muscle alpha isoform
(MyHC-alpha)
Length = 1938
Score = 49.7 bits (117), Expect = 6e-05
Identities = 96/450 (21%), Positives = 177/450 (39%), Gaps = 73/450 (16%)
Query: 50 EDLNEKKKEDWSEEEGK-----RMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLK 104
E+ +KK+ D S++ K + L + KL +E R++E + + L+
Sbjct: 1079 EEKLKKKEFDISQQNSKIEDEQALALQLQKKL-------KENQARIEELE------EELE 1125
Query: 105 VHHEGTSHVKETRIDIGVRKFELFEMKE------TETIDEMYGRFTIIMNELRSLEKDFT 158
+ V++ R D+ E+ E E + I+ R R LE+
Sbjct: 1126 AERTARAKVEKLRSDLSRELEEISERLEEAGGATSVQIEMNKKREAEFQKMRRDLEEATL 1185
Query: 159 IHERVRKILRCLPRSWRHIVTAITEAKDLKKL-RLEDLIGSLKAHEVLLQEDKSSNKSKM 217
HE LR +H + + + L R++ + K+ L +D +SN ++
Sbjct: 1186 QHEATAAALRK-----KHADSVAELGEQIDNLQRVKQKLEKEKSEFKLELDDVTSNMEQI 1240
Query: 218 IALKTNQESLNQELEMNFDNQQQELDEEDH------------QDQIILLTRKLQR---MI 262
I K N E +++ LE + + +L+E Q + L R+L+ +I
Sbjct: 1241 IKAKANLEKVSRTLEDQANEYRVKLEEAQRSLNDFTTQRAKLQTENGELARQLEEKEALI 1300
Query: 263 QRRDQNKRNFPARKENAKTELDKS-----------QVTCYGCYKLGHYKNECPLNKRKSS 311
+ + K ++ + E+ K +L++ Q + + C L E K +
Sbjct: 1301 SQLTRGKLSYTQQMEDLKRQLEEEGKAKNALAHALQSSRHDCDLLREQYEEEMEAKAELQ 1360
Query: 312 NLQTNQKSFMTTWDEPEE-----QTNEDEQADLCLMAKSDNEEVNLDP----CSSCEKTE 362
+ + S + W E +T E E+A L + + E ++ CSS EKT+
Sbjct: 1361 RVLSKANSEVAQWRTKYETDAIQRTEELEEAKKKLAQRLQDAEEAVEAVNAKCSSLEKTK 1420
Query: 363 HLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNS 422
H N + L + ER L K++ +K + + + S+S+ S S
Sbjct: 1421 HRLQN---EIEDLMVDVERSNAAAAALDKKQRNFDKILAEWKQKYEESQSELES-----S 1472
Query: 423 QKENIILKENVLKLKNDISNFVKSTETFQK 452
QKE L + KLKN ++ ETF++
Sbjct: 1473 QKEARSLSTELFKLKNAYEESLEHLETFKR 1502
Score = 36.6 bits (83), Expect = 0.56
Identities = 57/268 (21%), Positives = 107/268 (39%), Gaps = 37/268 (13%)
Query: 178 VTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESL----NQELEM 233
V +T++K + +++DL GSL+ + + + + + + LK QES+ N +L++
Sbjct: 1019 VNTLTKSKVKLEQQVDDLEGSLEQEKKVRMDLERAKRKLEGDLKLTQESIMDLENDKLQL 1078
Query: 234 -------NFDNQQQELDEEDHQDQIILLTRKLQ----RMIQRRDQNKRNFPARKENAKTE 282
FD QQ ED Q + L +KL+ R+ + ++ + AR + K
Sbjct: 1079 EEKLKKKEFDISQQNSKIEDEQALALQLQKKLKENQARIEELEEELEAERTARAKVEKLR 1138
Query: 283 LDKSQVTCYGCYKL----GHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQAD 338
D S+ +L G + +NK++ + Q ++ EE T + E
Sbjct: 1139 SDLSRELEEISERLEEAGGATSVQIEMNKKREAEFQKMRRDL-------EEATLQHEATA 1191
Query: 339 LCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECE--RLRNENLELRKEKEIL 396
L K S + DNL Q LE+E +L +++ E+ I
Sbjct: 1192 AALRKKH---------ADSVAELGEQIDNLQRVKQKLEKEKSEFKLELDDVTSNMEQIIK 1242
Query: 397 EKENKDLIKNIQHSESKEVSENKNNSQK 424
K N + + ++ E +Q+
Sbjct: 1243 AKANLEKVSRTLEDQANEYRVKLEEAQR 1270
Score = 35.0 bits (79), Expect = 1.6
Identities = 57/256 (22%), Positives = 100/256 (38%), Gaps = 30/256 (11%)
Query: 191 RLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDHQDQ 250
R++D + +A L+E KM++L + L +++ DN D E+ DQ
Sbjct: 860 RVKDALEKSEARRKELEE-------KMVSLLQEKNDLQLQVQAEQDNLN---DAEERCDQ 909
Query: 251 IIL----LTRKLQRMIQR-RDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPL 305
+I L K++ M +R D+ + N + K E + S++ K E L
Sbjct: 910 LIKNKIQLEAKVKEMTERLEDEEEMNAELTAKKRKLEDECSEL------KKDIDDLELTL 963
Query: 306 NKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLF 365
K + T K T EE DE K +E + + E
Sbjct: 964 AKVEKEKHATENKVKNLT----EEMAGLDEIIAKLTKEKKALQEAHQQALDDLQAEEDKV 1019
Query: 366 DNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKE 425
+ L S LEQ+ + L + +K + LE+ + L + K E+ + + +
Sbjct: 1020 NTLTKSKVKLEQQVDDLEGSLEQEKKVRMDLERAKRKL-----EGDLKLTQESIMDLEND 1074
Query: 426 NIILKENVLKLKNDIS 441
+ L+E + K + DIS
Sbjct: 1075 KLQLEEKLKKKEFDIS 1090
Score = 33.1 bits (74), Expect = 6.2
Identities = 48/204 (23%), Positives = 87/204 (42%), Gaps = 35/204 (17%)
Query: 69 LLNFKAKLFLTMA-LSREEYDRVQECKNAKE-----------IWDTLKVHHEGTSHVKET 116
L+N K K+ + L E + VQEC+NA+E + + LK + ++H++
Sbjct: 1725 LINQKKKMESDLTQLQTEVEEAVQECRNAEEKAKKAITDAAMMAEELKKEQDTSAHLERM 1784
Query: 117 RIDIGVRKFEL-FEMKETETIDEMYGRFTI--IMNELRSLEKDFTIHE-RVRKILRCLPR 172
+ ++ +L + E E I G+ + + +R LE + + R + ++ + +
Sbjct: 1785 KKNMEQTIKDLQHRLDEAEQIALKGGKKQLQKLEARVRELENELEAEQKRNAESVKGMRK 1844
Query: 173 SWRHI--VTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQE 230
S R I +T TE +RL+DL+ L+ K+ A K E ++
Sbjct: 1845 SERRIKELTYQTEEDKKNLMRLQDLVDKLQL--------------KVKAYKRQAEEAEEQ 1890
Query: 231 LEMN---FDNQQQELDEEDHQDQI 251
N F Q ELDE + + I
Sbjct: 1891 ANTNLSKFRKVQHELDEAEERADI 1914
>CCD2_HUMAN (Q96LB3) Coiled-coil domain containing protein 2
(Capillary morphogenesis protein-1) (CMG-1)
Length = 600
Score = 49.7 bits (117), Expect = 6e-05
Identities = 52/275 (18%), Positives = 120/275 (42%), Gaps = 42/275 (15%)
Query: 211 SSNKSKMIALKTNQESLNQELE--------MNFDNQQQELDEEDHQDQIILLTRKLQRMI 262
+ ++K +++ +E + QE + M+F+NQ + L+ + ++++ LQ+ +
Sbjct: 192 TERQAKEKQIRSVEEEIEQEKQATDDIIKNMSFENQVKYLEMKTTNEKLLQELDTLQQQL 251
Query: 263 QRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMT 322
++ +KE+ + E+ SQV K E L K L++++ +
Sbjct: 252 DSQNM-------KKESLEAEIAHSQV-----------KQEAVLLHEKLYELESHRDQMIA 293
Query: 323 TWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERL 382
++ + +E+ L K DN+E+ +S E+ L + + + Q E +
Sbjct: 294 --EDKSIGSPMEEREKLLKQIKDDNQEI-----ASMER------QLTDTKEKINQFIEEI 340
Query: 383 RNENLELRKEKEILEKENKDLIKNIQHSES--KEVSENKNNSQKENIILKENVLKLKNDI 440
R +++L + + + ++ K+L K +H ++ + E KN K ++ N++ L
Sbjct: 341 RQLDMDLEEHQGEMNQKYKELKKREEHMDTFIETFEETKNQELKRKAQIEANIVALLEHC 400
Query: 441 SNFVKSTETFQKIMGSQVSVFDKACIGFKTNQKQK 475
S + E I ++ + + FK+ + QK
Sbjct: 401 SRNINRIEQISSITNQELKMMQDD-LNFKSTEVQK 434
Score = 34.7 bits (78), Expect = 2.1
Identities = 69/345 (20%), Positives = 139/345 (40%), Gaps = 49/345 (14%)
Query: 86 EYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTI 145
E R Q K I ++ + +K+ +I + +L + KE +
Sbjct: 285 ESHRDQMIAEDKSIGSPMEEREKLLKQIKDDNQEIASMERQLTDTKE---------KINQ 335
Query: 146 IMNELRSLEKDFTIHE-RVRKILRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEV 204
+ E+R L+ D H+ + + + L + H+ T I ++ K L+ ++A+ V
Sbjct: 336 FIEEIRQLDMDLEEHQGEMNQKYKELKKREEHMDTFIETFEETKNQELKRK-AQIEANIV 394
Query: 205 LLQE--DKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRM- 261
L E ++ N+ + I+ TNQE + ++NF + + + + Q+ LT +QR+
Sbjct: 395 ALLEHCSRNINRIEQISSITNQELKMMQDDLNFKSTEVQKSQSTAQN----LTSDIQRLQ 450
Query: 262 --IQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSN------- 312
+Q+ + + + + K+++ K T Y N+ P K
Sbjct: 451 LDLQKMELLESKMTEEQHSLKSKI-KQMTTDLEIY------NDLPALKSSGEEKIKKLHQ 503
Query: 313 ----LQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEV-NLDPCSSCEKTEHLFDN 367
L T++ +F E+Q E E L + ++ NL+ K +HL N
Sbjct: 504 ERMILSTHRNAFKKIM---EKQNIEYEALKTQLQENETHSQLTNLE-----RKWQHLEQN 555
Query: 368 LFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSES 412
F + + + + + ++ K+I E NK ++ + HS S
Sbjct: 556 NFAMKEFIATKSQESDYQPIKKNVTKQIAE-YNKTIVDAL-HSTS 598
>SPOF_SCHPO (Q10411) Sporulation-specific protein 15
Length = 1957
Score = 49.3 bits (116), Expect = 8e-05
Identities = 97/447 (21%), Positives = 187/447 (41%), Gaps = 64/447 (14%)
Query: 92 ECKNAKEIWDTLKVHHEGT--------SHVKETRIDIGVRKFELFEMKE----------- 132
E KN +E+ D LK HE S + + + + EL + E
Sbjct: 711 EAKNLREVIDNLKGKHETLEAQRNDLHSSLSDAKNTNAILSSELTKSSEDVKRLTANVET 770
Query: 133 -TETIDEMYGRFTIIMNELRSL---------------EKDFTIHERVRKILRCLPRSWRH 176
T+ M FT ++N +S+ ++ T+ E K+ +
Sbjct: 771 LTQDSKAMKQSFTSLVNSYQSISNLYHELRDDHVNMQSQNNTLLESESKLKTDCENLTQQ 830
Query: 177 IVTAITEAKDL--KKLRLEDLIGSLKAHEVLLQED-KSSNKSKMIALKTNQESLNQ--EL 231
+T I + L K + E + LK L D K+ S +A+ N + L Q EL
Sbjct: 831 NMTLIDNVQKLMHKHVNQESKVSELKEVNGKLSLDLKNLRSSLNVAISDNDQILTQLAEL 890
Query: 232 EMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCY 291
N+D+ L++E Q L + + ++ + + + + K K ++++S+ +
Sbjct: 891 SKNYDS----LEQESAQLNSGLKSLEAEKQLLHTENEELHIRLDKLTGKLKIEESKSSDL 946
Query: 292 GCYKLGHYKNECPLNKRKSSNLQTNQ--KSFMTTWDEPEEQTNEDEQADLCLMAKSDNEE 349
G KL + E ++ K N+ +Q S + DE ++++ E AD+ + K+ E
Sbjct: 947 G-KKLTARQEE--ISNLKEENMSQSQAITSVKSKLDETLSKSSKLE-ADIEHL-KNKVSE 1001
Query: 350 VNLDPCSSCEKTEHLFDNLFYSSQ---MLEQECERLRNENLELRKEKEILEKENKDLIKN 406
V ++ + E L D+L + + L+ E E+ R EN +L+ + ++ E ++L+
Sbjct: 1002 VEVERNALLASNERLMDDLKNNGENIASLQTEIEKKRAENDDLQSKLSVVSSEYENLL-- 1059
Query: 407 IQHSESKEVSENKNNSQKENIILKENVLKLKNDIS----NFVKSTETFQKIMGSQVSVFD 462
+ S++ + E+K N K +++NV KL ++ + T + K+ + D
Sbjct: 1060 LISSQTNKSLEDKTNQLK---YIEKNVQKLLDEKDQRNVELEELTSKYGKLGEENAQIKD 1116
Query: 463 KACIGFKTNQKQ-KLYENFFIPKKNKN 488
+ K ++KQ L NF K K+
Sbjct: 1117 ELLALRKKSKKQHDLCANFVDDLKEKS 1143
Score = 44.3 bits (103), Expect = 0.003
Identities = 95/430 (22%), Positives = 164/430 (38%), Gaps = 70/430 (16%)
Query: 63 EEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHV--------K 114
EE K + N+ + L + R+ E N K DTL + +E + K
Sbjct: 301 EELKHNVANYSDAIVHKDKLIEDLSTRISEFDNLKSERDTLSIKNEKLEKLLRNTIGSLK 360
Query: 115 ETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSW 174
++R + E+ E+KE+ I ++L E + E+ K L+ +
Sbjct: 361 DSRTSNSQLEEEMVELKESNRT---------IHSQLTDAESKLSSFEQENKSLKGSIDEY 411
Query: 175 RHIVTAITEAKDLKKLRLEDLIGSL-----KAHEVLLQEDKSSNKSKMIALKTNQESLNQ 229
++ +++ + +LE+ SL K E+ + D + K K + E + Q
Sbjct: 412 QNNLSSKDKMVKQVSSQLEEARSSLAHATGKLAEINSERDFQNKKIK------DFEKIEQ 465
Query: 230 ELEMNFDNQQQELDEE----DHQDQIILLTRK------------------LQRMIQRRDQ 267
+L ++ EL E+ D +DQ + R+ LQR I +
Sbjct: 466 DLRACLNSSSNELKEKSALIDKKDQELNNLREQIKEQKKVSESTQSSLQSLQRDILNEKK 525
Query: 268 NKRNFPARKENAKTELD---------KSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQK 318
+ ++ K EL SQ++ K L++ K+S LQT
Sbjct: 526 KHEVYESQLNELKGELQTEISNSEHLSSQLSTLAAEKEAAVATNNELSESKNS-LQTLCN 584
Query: 319 SFMTTWDEPEEQTNEDEQ--ADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQM-- 374
+F + Q E+EQ + L K NE + T+ L D Q+
Sbjct: 585 AFQEKLAKSVMQLKENEQNFSSLDTSFKKLNESHQELENNHQTITKQLKDTSSKLQQLQL 644
Query: 375 ----LEQECERLRNENLELRKEKEILEKENKDLIKNIQHSES--KEVSENKNNSQKENII 428
EQ+ L +EN +LR + LE+ NK LIK + +S K + K + +K
Sbjct: 645 ERANFEQKESTLSDENNDLRTKLLKLEESNKSLIKKQEDVDSLEKNIQTLKEDLRKSEEA 704
Query: 429 LKENVLKLKN 438
L+ + L+ KN
Sbjct: 705 LRFSKLEAKN 714
Score = 37.4 bits (85), Expect = 0.33
Identities = 85/404 (21%), Positives = 160/404 (39%), Gaps = 52/404 (12%)
Query: 62 EEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIG 121
E+E LL K T L + E ECK +E + ++ E + ++E + ++
Sbjct: 251 EQERLEKLLVSSNKTVST--LRQTENSLRAECKTLQEKLEKCAINEEDSKLLEELKHNVA 308
Query: 122 VRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTI-HERVRKILRCLPRSWRHIVTA 180
+ + + + I+++ R + N L+S +I +E++ K+LR
Sbjct: 309 --NYSDAIVHKDKLIEDLSTRISEFDN-LKSERDTLSIKNEKLEKLLR----------NT 355
Query: 181 ITEAKDLK--KLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLN---QELEMNF 235
I KD + +LE+ + LK + + +SK+ + + +SL E + N
Sbjct: 356 IGSLKDSRTSNSQLEEEMVELKESNRTIHSQLTDAESKLSSFEQENKSLKGSIDEYQNNL 415
Query: 236 DNQQQELDE-----EDHQDQIILLTRKLQRMIQRRD-QNKRNFPARKENAKTELDKSQVT 289
++ + + + E+ + + T KL + RD QNK+ + +K +
Sbjct: 416 SSKDKMVKQVSSQLEEARSSLAHATGKLAEINSERDFQNKKI---------KDFEKIEQD 466
Query: 290 CYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEE 349
C L NE K KS+ + + ++ +EQ E L + +
Sbjct: 467 LRAC--LNSSSNEL---KEKSALIDKKDQELNNLREQIKEQKKVSESTQSSLQSLQRDI- 520
Query: 350 VNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQH 409
L+ E E + L Q E L ++ L EKE N +L
Sbjct: 521 --LNEKKKHEVYESQLNELKGELQTEISNSEHLSSQLSTLAAEKEAAVATNNEL------ 572
Query: 410 SESKEVSENKNNSQKENIILKENVLKLKNDISNFVKSTETFQKI 453
SESK + N+ +E L ++V++LK + NF +F+K+
Sbjct: 573 SESKNSLQTLCNAFQEK--LAKSVMQLKENEQNFSSLDTSFKKL 614
Score = 37.0 bits (84), Expect = 0.43
Identities = 78/372 (20%), Positives = 142/372 (37%), Gaps = 37/372 (9%)
Query: 106 HHEGTSHVKETRIDIGVRKFEL--FEMKETETIDEMYGRFTIIMN---ELRSLEKDFTIH 160
H T +K+T + + E FE KE+ DE T ++ +SL K
Sbjct: 625 HQTITKQLKDTSSKLQQLQLERANFEQKESTLSDENNDLRTKLLKLEESNKSLIKKQEDV 684
Query: 161 ERVRKILRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIAL 220
+ + K ++ L R A+ +K L+ L ++I +LK L+ ++ S +
Sbjct: 685 DSLEKNIQTLKEDLRKSEEALRFSK-LEAKNLREVIDNLKGKHETLEAQRNDLHSSLSDA 743
Query: 221 KTNQESLNQELEMNFDNQQQ--------ELDEEDHQDQIILLTRKLQRMIQRRDQNKRNF 272
K L+ EL + ++ ++ D + + L Q + + + +
Sbjct: 744 KNTNAILSSELTKSSEDVKRLTANVETLTQDSKAMKQSFTSLVNSYQSISNLYHELRDDH 803
Query: 273 PARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTN 332
+ T L+ C L N +K + NQ+S ++ E + +
Sbjct: 804 VNMQSQNNTLLESESKLKTDCENLTQQNMTLIDNVQKLMHKHVNQESKVSELKEVNGKLS 863
Query: 333 EDEQ--ADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELR 390
D + +A SDN+++ L + K Y S LEQE +L + L
Sbjct: 864 LDLKNLRSSLNVAISDNDQI-LTQLAELSKN--------YDS--LEQESAQLNSGLKSLE 912
Query: 391 KEKEILEKENKDL---------IKNIQHSESKEVSENKNNSQKENIILKENVLKLKNDIS 441
EK++L EN++L I+ S+S ++ + Q+E LKE + I+
Sbjct: 913 AEKQLLHTENEELHIRLDKLTGKLKIEESKSSDLGKKLTARQEEISNLKEENMSQSQAIT 972
Query: 442 NF-VKSTETFQK 452
+ K ET K
Sbjct: 973 SVKSKLDETLSK 984
>MYH4_RABIT (Q28641) Myosin heavy chain, skeletal muscle, juvenile
Length = 1938
Score = 49.3 bits (116), Expect = 8e-05
Identities = 86/443 (19%), Positives = 186/443 (41%), Gaps = 56/443 (12%)
Query: 53 NEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECK-NAKEIWDTLKVHHEGTS 111
N+K++ D ++ + + N ++K+ AL+ + +++E + +E+ + ++ +
Sbjct: 1074 NDKQQLDEKLKKKEFEMSNLQSKIEDEQALAMQLQKKIKELQARIEELEEEIEAERASRA 1133
Query: 112 HVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLP 171
++ R D+ E+ +E ++E G + + + E +F +++R+ L
Sbjct: 1134 KAEKQRSDLSRELEEI-----SERLEEAGGATSAQIEMNKKREAEF---QKMRRDLE--E 1183
Query: 172 RSWRHIVTAITEAKDLKK--LRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLN- 228
+ +H TA T K L + I +L+ + L+++KS K ++ L +N E+++
Sbjct: 1184 ATLQHEATAATLRKKHADSVAELGEQIDNLQRVKQKLEKEKSELKMEIDDLASNMETVSK 1243
Query: 229 -----QELEMNFDNQQQELD--EEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKT 281
+++ ++Q EL EE+HQ I L+ + R+ + R K++ +
Sbjct: 1244 AKGNLEKMCRTLEDQVSELKTKEEEHQRLINDLSAQRARLQTESGEFSRQLD-EKDSLVS 1302
Query: 282 ELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCL 341
+L + + ++ K + + S L +S D EQ E+++A L
Sbjct: 1303 QLSRGKQAF--TQQIEELKRQLEEEIKAKSALAHALQSARHDCDLLREQYEEEQEAKAEL 1360
Query: 342 ---MAKSDNE------EVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKE 392
M+K+++E + D E+ E L Q E+ E + + L K
Sbjct: 1361 QRAMSKANSEVAQWRTKYETDAIQRTEELEEAKKKLAQRLQDAEEHVEAVNAKCASLEKT 1420
Query: 393 KEILEKENKDLIKNIQHSES-----------------------KEVSENKNNSQKENIIL 429
K+ L+ E +DL+ +++ + + +E SQKE+ L
Sbjct: 1421 KQRLQNEVEDLMIDVERTNAACAALDKKQRNFDKILAEWKHKYEETHAELEASQKESRSL 1480
Query: 430 KENVLKLKNDISNFVKSTETFQK 452
V K+KN + ET ++
Sbjct: 1481 STEVFKVKNAYEESLDQLETLKR 1503
Score = 45.4 bits (106), Expect = 0.001
Identities = 62/271 (22%), Positives = 111/271 (40%), Gaps = 32/271 (11%)
Query: 178 VTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDN 237
V +T+AK + +++DL GSL+ E ++ D K K+ L QE M+ +N
Sbjct: 1020 VNTLTKAKTKLEQQVDDLEGSLE-QEKKIRMDLERAKRKL----EGDLKLAQESTMDIEN 1074
Query: 238 QQQELDE----------------EDHQDQIILLTRKLQRMIQRRDQNKRNFPA-RKENAK 280
+Q+LDE ED Q + L +K++ + R ++ + A R AK
Sbjct: 1075 DKQQLDEKLKKKEFEMSNLQSKIEDEQALAMQLQKKIKELQARIEELEEEIEAERASRAK 1134
Query: 281 TELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLC 340
E +S ++ ++ E S+ ++ N+K E E Q + +
Sbjct: 1135 AEKQRSDLS-RELEEISERLEEA--GGATSAQIEMNKKR------EAEFQKMRRDLEEAT 1185
Query: 341 LMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKEN 400
L ++ + S + DNL Q LE+E L+ E +L E + K
Sbjct: 1186 LQHEATAATLRKKHADSVAELGEQIDNLQRVKQKLEKEKSELKMEIDDLASNMETVSKAK 1245
Query: 401 KDLIKNIQHSESKEVSENKNNSQKENIILKE 431
+L K + E +VSE K ++ ++ +
Sbjct: 1246 GNLEKMCRTLED-QVSELKTKEEEHQRLIND 1275
Score = 33.5 bits (75), Expect = 4.7
Identities = 44/201 (21%), Positives = 84/201 (40%), Gaps = 21/201 (10%)
Query: 133 TETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKLRL 192
+E + ++ + T ++N + LE D + + IV A++ K +
Sbjct: 1713 SERVQLLHTQNTSLINTKKKLETDISQ----------IQGEMEDIVQEARNAEEKAKKAI 1762
Query: 193 EDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDHQDQII 252
D ++ A E+ ++D S++ +M K N E ++L+ D +Q L + + QI
Sbjct: 1763 TD--AAMMAEELKKEQDTSAHLERM---KKNMEQTVKDLQHRLDEAEQ-LALKGGKKQIQ 1816
Query: 253 LLTRKLQRM-IQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSS 311
L +++ + + + KRN A K K E ++T Y+ + +
Sbjct: 1817 KLEARVRELEAEVESEQKRNVEAVKGLRKHERRVKELT----YQTEEDRKNVLRLQDLVD 1872
Query: 312 NLQTNQKSFMTTWDEPEEQTN 332
LQ KS+ +E EEQ N
Sbjct: 1873 KLQAKVKSYKRQAEEAEEQCN 1893
>MYH8_HUMAN (P13535) Myosin heavy chain, skeletal muscle, perinatal
(MyHC-perinatal)
Length = 1937
Score = 48.9 bits (115), Expect = 1e-04
Identities = 85/441 (19%), Positives = 186/441 (41%), Gaps = 52/441 (11%)
Query: 53 NEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECK-NAKEIWDTLKVHHEGTS 111
N+K++ D E+ + + N +K+ A+ + +++E + +E+ + ++ +
Sbjct: 1074 NDKQQLDEKLEKKEFEISNLISKIEDEQAVEIQLQKKIKELQARIEELGEEIEAERASRA 1133
Query: 112 HVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLP 171
++ R D+ E+ +E ++E G + + + E +F +++R+ L
Sbjct: 1134 KAEKQRSDLSRELEEI-----SERLEEAGGATSAQVELNKKREAEF---QKLRRDLEEAT 1185
Query: 172 RSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLN--- 228
+V A+ + L + I +L+ + L+++KS K + L +N E+++
Sbjct: 1186 LQHEAMVAALRKKHADSMAELGEQIDNLQRVKQKLEKEKSELKMETDDLSSNAEAISKAK 1245
Query: 229 ---QELEMNFDNQQQELDEEDHQDQIIL--LTRKLQRMIQRRDQNKRNFPARKENAKTEL 283
+++ + ++Q EL ++ + Q ++ LT + R+ + R K+ ++L
Sbjct: 1246 GNLEKMCRSLEDQVSELKTKEEEQQRLINDLTAQRARLQTEAGEYSRQLD-EKDALVSQL 1304
Query: 284 DKS-QVTCYGCYKLGHYKNECPLNKRKSSN-LQTNQKSFMTTWDEPEEQTNEDEQADLCL 341
+S Q + +L H E K ++ LQ+++ ++ EE+ ++ +A+L
Sbjct: 1305 SRSKQASTQQIEELKHQLEEETKAKNALAHALQSSRHDCDLLREQYEEE--QEGKAELQR 1362
Query: 342 MAKSDNEEV-------NLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKE 394
N EV D E+ E L Q E+ E + + L K K+
Sbjct: 1363 ALSKANSEVAQWRTKYETDAIQRTEELEEAKKKLAQRLQEAEEHVEAVNAKCASLEKTKQ 1422
Query: 395 ILEKENKDLIKNIQHSES-------------KEVSENKNN----------SQKENIILKE 431
L+ E +DL+ +++ S + K +SE K SQKE+ L
Sbjct: 1423 RLQNEVEDLMLDVERSNAACAALDKKQRNFDKVLSEWKQKYEETQAELEASQKESRSLST 1482
Query: 432 NVLKLKNDISNFVKSTETFQK 452
+ K+KN + ET ++
Sbjct: 1483 ELFKVKNVYEESLDQLETLRR 1503
Score = 47.8 bits (112), Expect = 2e-04
Identities = 68/282 (24%), Positives = 116/282 (41%), Gaps = 54/282 (19%)
Query: 178 VTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDN 237
V +T+AK + +++DL GSL+ + L + + + + LK QES M+ +N
Sbjct: 1020 VNILTKAKTKLEQQVDDLEGSLEQEKKLRMDLERAKRKLEGDLKLAQEST-----MDMEN 1074
Query: 238 QQQELDE----------------EDHQDQIILLTRKLQRMIQRRDQNKRNFPA-RKENAK 280
+Q+LDE ED Q I L +K++ + R ++ A R AK
Sbjct: 1075 DKQQLDEKLEKKEFEISNLISKIEDEQAVEIQLQKKIKELQARIEELGEEIEAERASRAK 1134
Query: 281 TELDKSQVTCYGCYKL-----------GHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEE 329
E +S ++ +L G + LNK++ + Q ++ EE
Sbjct: 1135 AEKQRSDLS----RELEEISERLEEAGGATSAQVELNKKREAEFQKLRRDL-------EE 1183
Query: 330 QTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLEL 389
T + E L K + S E E + DNL Q LE+E L+ E +L
Sbjct: 1184 ATLQHEAMVAALRKKHAD--------SMAELGEQI-DNLQRVKQKLEKEKSELKMETDDL 1234
Query: 390 RKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENIILKE 431
E + K +L K + E +VSE K +++ ++ +
Sbjct: 1235 SSNAEAISKAKGNLEKMCRSLED-QVSELKTKEEEQQRLIND 1275
Score = 35.4 bits (80), Expect = 1.2
Identities = 49/238 (20%), Positives = 100/238 (41%), Gaps = 35/238 (14%)
Query: 97 KEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKD 156
+E+W TL+ ++ +D +E + ++ + T ++N + LE D
Sbjct: 1689 EELWATLEQTERSRKIAEQELLDA------------SERVQLLHTQNTSLINTKKKLEND 1736
Query: 157 FT-IHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKS 215
+ + V ++++ + AIT+A ++ A E+ ++D S++
Sbjct: 1737 VSQLQSEVEEVIQESRNAEEKAKKAITDA-------------AMMAEELKKEQDTSAHLE 1783
Query: 216 KMIALKTNQESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMI-QRRDQNKRNFPA 274
+M K N E ++L+ D +Q L + + QI L +++ + + ++ KRN A
Sbjct: 1784 RM---KKNLEQTVKDLQHRLDEAEQ-LALKGGKKQIQKLEARVRELEGEVENEQKRNAEA 1839
Query: 275 RKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTN 332
K K E ++T Y+ + + LQ KS+ +E EEQ+N
Sbjct: 1840 VKGLRKHERRVKELT----YQTEEDRKNVLRLQDLVDKLQAKVKSYKRQAEEAEEQSN 1893
Score = 32.7 bits (73), Expect = 8.1
Identities = 96/462 (20%), Positives = 199/462 (42%), Gaps = 64/462 (13%)
Query: 48 YDEDLNE---KKKEDWSE-EEGKRMLLNFKAKLFLTMALSREEYDRVQ----ECKNAK-E 98
+D+ L+E K +E +E E ++ + +LF + E D+++ E KN + E
Sbjct: 1452 FDKVLSEWKQKYEETQAELEASQKESRSLSTELFKVKNVYEESLDQLETLRRENKNLQQE 1511
Query: 99 IWDTLKVHHEGTSHVKET---RIDIGVRKFEL-FEMKETE-TIDEMYGRFTIIMNELRSL 153
I D + EG + E + + K E+ ++E E +++ G+ I EL +
Sbjct: 1512 ISDLTEQIAEGGKQIHELEKIKKQVEQEKCEIQAALEEAEASLEHEEGKILRIQLELNQV 1571
Query: 154 --EKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQE-DK 210
E D I E+ +I + L R+ +V + D + D + K E L E +
Sbjct: 1572 KSEVDRKIAEKDEEIDQ-LKRNHTRVVETMQSTLDAEIRSRNDALRVKKKMEGDLNEMEI 1630
Query: 211 SSNKSKMIALKT-----NQESLNQELEMNFDNQQQELDEEDHQDQIILLTRK-------L 258
N + +A ++ N + + +E +++ D+ + +ED ++Q+ ++ R+ +
Sbjct: 1631 QLNHANRLAAESLRNYRNTQGILKETQLHLDDALR--GQEDLKEQLAIVERRANLLQAEI 1688
Query: 259 QRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQK 318
+ + +Q +R+ RK + LD S+ +L H +N +N +K L+ +
Sbjct: 1689 EELWATLEQTERS---RKIAEQELLDASERV-----QLLHTQNTSLINTKKK--LENDVS 1738
Query: 319 SFMTTWDEP-EEQTNEDEQA-----DLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSS 372
+ +E +E N +E+A D +MA+ +E D + E+ +
Sbjct: 1739 QLQSEVEEVIQESRNAEEKAKKAITDAAMMAEELKKEQ--DTSAHLERMK---------- 1786
Query: 373 QMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENIILKEN 432
+ LEQ + L++ L + +++ K K I+ ++ + +E+ N QK N +
Sbjct: 1787 KNLEQTVKDLQHR---LDEAEQLALKGGKKQIQKLE-ARVRELEGEVENEQKRNAEAVKG 1842
Query: 433 VLKLKNDISNFVKSTETFQKIMGSQVSVFDKACIGFKTNQKQ 474
+ K + + TE +K + + DK K+ ++Q
Sbjct: 1843 LRKHERRVKELTYQTEEDRKNVLRLQDLVDKLQAKVKSYKRQ 1884
Score = 32.7 bits (73), Expect = 8.1
Identities = 61/322 (18%), Positives = 118/322 (35%), Gaps = 44/322 (13%)
Query: 150 LRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQED 209
L+ E F I VR + W + I LK E + ++K +++
Sbjct: 808 LQRREALFCIQYNVRAFMNVKHWPWMKLFFKIKPL--LKSAETEKEMATMKEEFQKTKDE 865
Query: 210 KSSNKSKMIALKTNQESLNQE---LEMNFDNQQQEL-DEEDHQDQIIL----LTRKLQRM 261
+ +++K L+ +L +E L++ ++ L D E+ +Q+I L K++ +
Sbjct: 866 LAKSEAKRKELEEKMVTLLKEKNDLQLQVQSEADSLADAEERCEQLIKNKIQLEAKIKEV 925
Query: 262 IQRRDQN----------KRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSS 311
+R ++ KR K ++D ++T L + E + K
Sbjct: 926 TERAEEEEEINAELTAKKRKLEDECSELKKDIDDLELT------LAKVEKEKHATENKVK 979
Query: 312 NLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYS 371
NL EE DE K +E + + E + L +
Sbjct: 980 NLT-------------EEMAGLDETIAKLSKEKKALQETHQQTLDDLQAEEDKVNILTKA 1026
Query: 372 SQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENIILKE 431
LEQ+ + L + +K + LE+ + L + K E+ + + + L E
Sbjct: 1027 KTKLEQQVDDLEGSLEQEKKLRMDLERAKRKL-----EGDLKLAQESTMDMENDKQQLDE 1081
Query: 432 NVLKLKNDISNFVKSTETFQKI 453
+ K + +ISN + E Q +
Sbjct: 1082 KLEKKEFEISNLISKIEDEQAV 1103
>MYH6_MESAU (P13539) Myosin heavy chain, cardiac muscle alpha isoform
(MyHC-alpha)
Length = 1939
Score = 48.9 bits (115), Expect = 1e-04
Identities = 96/450 (21%), Positives = 176/450 (38%), Gaps = 73/450 (16%)
Query: 50 EDLNEKKKEDWSEEEGK-----RMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLK 104
E+ +KK+ D S++ K + L + KL +E R++E + + L+
Sbjct: 1079 EEKLKKKEFDISQQNSKIEDEQALALQLQKKL-------KENQARIEELE------EELE 1125
Query: 105 VHHEGTSHVKETRIDIGVRKFELFEMKE------TETIDEMYGRFTIIMNELRSLEKDFT 158
+ V++ R D+ E+ E E + I+ R R LE+
Sbjct: 1126 AERTARAKVEKLRSDLTRELEEISERLEEAGGATSVQIEMNKKREAEFQKMRRDLEEATL 1185
Query: 159 IHERVRKILRCLPRSWRHIVTAITEAKDLKKL-RLEDLIGSLKAHEVLLQEDKSSNKSKM 217
HE LR +H + + + L R++ + K+ L +D +SN ++
Sbjct: 1186 QHEATAAALRK-----KHADSVAELGEQIDNLQRVKQKLEKEKSEFKLELDDVTSNMEQI 1240
Query: 218 IALKTNQESLNQELEMNFDNQQQELDEEDH------------QDQIILLTRKLQR---MI 262
I K N E +++ LE + + +L+E Q + L R+L+ +I
Sbjct: 1241 IKAKANLEKVSRTLEDQANEYRVKLEESQRSLNDFTTQRAKLQTENGELARQLEEKEALI 1300
Query: 263 QRRDQNKRNFPARKENAKTELDKS-----------QVTCYGCYKLGHYKNECPLNKRKSS 311
+ + K ++ + E+ K +L++ Q + C L E K +
Sbjct: 1301 SQLTRGKLSYTQQMEDLKRQLEEEGKAKNALAHALQSARHDCDLLREQYEEEMEAKAELQ 1360
Query: 312 NLQTNQKSFMTTWDEPEE-----QTNEDEQADLCLMAKSDNEEVNLDP----CSSCEKTE 362
+ + S + W E +T E E+A L + + E ++ CSS EKT+
Sbjct: 1361 RVLSKANSEVAQWRTKYETDAIQRTEELEEAKKKLAQRLQDAEEAVEAVNAKCSSLEKTK 1420
Query: 363 HLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNS 422
H N + L + ER L K++ +K + + + S+S+ S S
Sbjct: 1421 HRLQN---EIEDLMVDVERSNAAAAALDKKQRNFDKILAEWKQKYEESQSELES-----S 1472
Query: 423 QKENIILKENVLKLKNDISNFVKSTETFQK 452
QKE L + KLKN ++ ETF++
Sbjct: 1473 QKEARSLSTELFKLKNAYEESLEHLETFKR 1502
Score = 37.0 bits (84), Expect = 0.43
Identities = 59/272 (21%), Positives = 106/272 (38%), Gaps = 45/272 (16%)
Query: 178 VTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESL----NQELEM 233
V +T++K + +++DL GSL+ + + + + + + L QES+ N +L++
Sbjct: 1019 VNTLTKSKVKLEQQVDDLEGSLEQEKKVRMDLERAKRKLEGDLNVTQESIMDLENDKLQL 1078
Query: 234 -------NFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPA-RKENAKTELDK 285
FD QQ ED Q + L +KL+ R ++ + A R AK E +
Sbjct: 1079 EEKLKKKEFDISQQNSKIEDEQALALQLQKKLKENQARIEELEEELEAERTARAKVEKLR 1138
Query: 286 SQVTCYGCYKL-----------GHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNED 334
S +T +L G + +NK++ + Q ++ EE T +
Sbjct: 1139 SDLT----RELEEISERLEEAGGATSVQIEMNKKREAEFQKMRRDL-------EEATLQH 1187
Query: 335 EQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECE--RLRNENLELRKE 392
E L K S + DNL Q LE+E +L +++ E
Sbjct: 1188 EATAAALRKKH---------ADSVAELGEQIDNLQRVKQKLEKEKSEFKLELDDVTSNME 1238
Query: 393 KEILEKENKDLIKNIQHSESKEVSENKNNSQK 424
+ I K N + + ++ E SQ+
Sbjct: 1239 QIIKAKANLEKVSRTLEDQANEYRVKLEESQR 1270
Score = 33.9 bits (76), Expect = 3.6
Identities = 58/264 (21%), Positives = 102/264 (37%), Gaps = 24/264 (9%)
Query: 187 LKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQE---LEMNFDNQQQEL- 242
LK E + ++K ++E ++++ L+ SL QE L+ +Q L
Sbjct: 842 LKSAETEKEMANMKEEFGRVKESLEKSEARRKELEEKMVSLLQEKNDLQFQVQAEQDNLN 901
Query: 243 DEEDHQDQIIL----LTRKLQRMIQR-RDQNKRNFPARKENAKTELDKSQVTCYGCYKLG 297
D E+ DQ+I L K++ M +R D+ + N + K E + S++ K
Sbjct: 902 DAEERCDQLIKNKIQLEAKVKEMTERLEDEEEMNAELTSKKRKLEDECSEL------KKD 955
Query: 298 HYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSS 357
E L K + T K T EE DE K +E +
Sbjct: 956 IDDLELTLAKVEKEKHATENKVKNLT----EEMAGLDEIIAKLTKEKKALQEAHQQALDD 1011
Query: 358 CEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEVSE 417
+ E + L S LEQ+ + L + +K + LE+ + L + E
Sbjct: 1012 LQAEEDKVNTLTKSKVKLEQQVDDLEGSLEQEKKVRMDLERAKRKL-----EGDLNVTQE 1066
Query: 418 NKNNSQKENIILKENVLKLKNDIS 441
+ + + + + L+E + K + DIS
Sbjct: 1067 SIMDLENDKLQLEEKLKKKEFDIS 1090
Score = 33.1 bits (74), Expect = 6.2
Identities = 48/204 (23%), Positives = 87/204 (42%), Gaps = 35/204 (17%)
Query: 69 LLNFKAKLFLTMA-LSREEYDRVQECKNAKE-----------IWDTLKVHHEGTSHVKET 116
L+N K K+ + L E + VQEC+NA+E + + LK + ++H++
Sbjct: 1725 LINQKKKMEADLTQLQTEVEEAVQECRNAEEKAKKAITDAAMMAEELKKEQDTSAHLERM 1784
Query: 117 RIDIGVRKFEL-FEMKETETIDEMYGRFTI--IMNELRSLEKDFTIHE-RVRKILRCLPR 172
+ ++ +L + E E I G+ + + +R LE + + R + ++ + +
Sbjct: 1785 KKNMEQTIKDLQHRLDEAEQIALKGGKKQLQKLEARVRELENELEAEQKRNAESVKGMRK 1844
Query: 173 SWRHI--VTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQE 230
S R I +T TE +RL+DL+ L+ K+ A K E ++
Sbjct: 1845 SERRIKELTYQTEEDKKNLVRLQDLVDKLQL--------------KVKAYKRQAEEAEEQ 1890
Query: 231 LEMN---FDNQQQELDEEDHQDQI 251
N F Q ELDE + + I
Sbjct: 1891 ANTNLSKFRKVQHELDEAEERADI 1914
>MYH6_HUMAN (P13533) Myosin heavy chain, cardiac muscle alpha isoform
(MyHC-alpha)
Length = 1939
Score = 48.5 bits (114), Expect = 1e-04
Identities = 85/398 (21%), Positives = 156/398 (38%), Gaps = 55/398 (13%)
Query: 97 KEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEMKE------TETIDEMYGRFTIIMNEL 150
+E+ + L+ + V++ R D+ E+ E E + I+ R
Sbjct: 1118 EELEEELEAERTARAKVEKLRSDLSRELEEISERLEEAGGATSVQIEMNKKREAEFQKMR 1177
Query: 151 RSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKL-RLEDLIGSLKAHEVLLQED 209
R LE+ HE LR +H + + + L R++ + K+ L +D
Sbjct: 1178 RDLEEATLQHEATAAALRK-----KHADSVAELGEQIDNLQRVKQKLEKEKSEFKLELDD 1232
Query: 210 KSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDH------------QDQIILLTRK 257
+SN ++I K N E +++ LE + + +L+E Q + L R+
Sbjct: 1233 VTSNMEQIIKAKANLEKVSRTLEDQANEYRVKLEEAQRSLNDFTTQRAKLQTENGELARQ 1292
Query: 258 LQR---MIQRRDQNKRNFPARKENAKTELDKS-----------QVTCYGCYKLGHYKNEC 303
L+ +I + + K ++ + E+ K +L++ Q + C L E
Sbjct: 1293 LEEKEALISQLTRGKLSYTQQMEDLKRQLEEEGKAKNALAHALQSARHDCDLLREQYEEE 1352
Query: 304 PLNKRKSSNLQTNQKSFMTTWDEPEE-----QTNEDEQADLCLMAKSDNEEVNLDP---- 354
K + + + S + W E +T E E+A L + + E ++
Sbjct: 1353 TEAKAELQRVLSKANSEVAQWRTKYETDAIQRTEELEEAKKKLAQRLQDAEEAVEAVNAK 1412
Query: 355 CSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKE 414
CSS EKT+H N + L + ER L K++ +K + + + S+S+
Sbjct: 1413 CSSLEKTKHRLQN---EIEDLMVDVERSNAAAAALDKKQRNFDKILAEWKQKYEESQSEL 1469
Query: 415 VSENKNNSQKENIILKENVLKLKNDISNFVKSTETFQK 452
S SQKE L + KLKN ++ ETF++
Sbjct: 1470 ES-----SQKEARSLSTELFKLKNAYEESLEHLETFKR 1502
Score = 35.8 bits (81), Expect = 0.95
Identities = 55/267 (20%), Positives = 110/267 (40%), Gaps = 30/267 (11%)
Query: 187 LKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQE---LEMNFDNQQQEL- 242
LK E + ++K ++E ++++ L+ SL QE L++ +Q L
Sbjct: 842 LKSAETEKEMATMKEEFGRIKETLEKSEARRKELEEKMVSLLQEKNDLQLQVQAEQDNLN 901
Query: 243 DEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNE 302
D E+ DQ+I K+Q + ++ N+R + NA+ K ++ ++E
Sbjct: 902 DAEERCDQLI--KNKIQLEAKVKEMNERLEDEEEMNAELTAKKRKL-----------EDE 948
Query: 303 CPLNKRKSSNLQTN----QKSFMTTWDEPEEQTNE----DEQADLCLMAKSDNEEVNLDP 354
C K+ +L+ +K T ++ + T E DE K +E +
Sbjct: 949 CSELKKDIDDLELTLAKVEKEKHATENKVKNLTEEMAGLDEIIAKLTKEKKALQEAHQQA 1008
Query: 355 CSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKE 414
+ E ++L S LEQ+ + L + +K + LE+ + L + K
Sbjct: 1009 LDDLQVEEDKVNSLSKSKVKLEQQVDDLEGSLEQEKKVRMDLERAKRKL-----EGDLKL 1063
Query: 415 VSENKNNSQKENIILKENVLKLKNDIS 441
E+ + + + + L+E + K + DI+
Sbjct: 1064 TQESIMDLENDKLQLEEKLKKKEFDIN 1090
Score = 35.4 bits (80), Expect = 1.2
Identities = 56/268 (20%), Positives = 108/268 (39%), Gaps = 37/268 (13%)
Query: 178 VTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESL----NQELEM 233
V +++++K + +++DL GSL+ + + + + + + LK QES+ N +L++
Sbjct: 1019 VNSLSKSKVKLEQQVDDLEGSLEQEKKVRMDLERAKRKLEGDLKLTQESIMDLENDKLQL 1078
Query: 234 -------NFDNQQQELDEEDHQDQIILLTRKLQ----RMIQRRDQNKRNFPARKENAKTE 282
FD QQ ED Q + L +KL+ R+ + ++ + AR + K
Sbjct: 1079 EEKLKKKEFDINQQNSKIEDEQALALQLQKKLKENQARIEELEEELEAERTARAKVEKLR 1138
Query: 283 LDKSQVTCYGCYKL----GHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQAD 338
D S+ +L G + +NK++ + Q ++ EE T + E
Sbjct: 1139 SDLSRELEEISERLEEAGGATSVQIEMNKKREAEFQKMRRDL-------EEATLQHEATA 1191
Query: 339 LCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECE--RLRNENLELRKEKEIL 396
L K S + DNL Q LE+E +L +++ E+ I
Sbjct: 1192 AALRKKH---------ADSVAELGEQIDNLQRVKQKLEKEKSEFKLELDDVTSNMEQIIK 1242
Query: 397 EKENKDLIKNIQHSESKEVSENKNNSQK 424
K N + + ++ E +Q+
Sbjct: 1243 AKANLEKVSRTLEDQANEYRVKLEEAQR 1270
Score = 33.5 bits (75), Expect = 4.7
Identities = 49/204 (24%), Positives = 86/204 (42%), Gaps = 35/204 (17%)
Query: 69 LLNFKAKLFLTMA-LSREEYDRVQECKNAKE-----------IWDTLKVHHEGTSHVKET 116
L+N K K+ + L E + VQEC+NA+E + + LK + ++H++
Sbjct: 1725 LINQKKKMEADLTQLQSEVEEAVQECRNAEEKAKKAITDAAMMAEELKKEQDTSAHLERM 1784
Query: 117 RIDIGVRKFEL-FEMKETETIDEMYGRFTIIMNELRSLEKDFTI---HERVRKILRCLPR 172
+ ++ +L + E E I G+ + E R E + + +R + ++ + +
Sbjct: 1785 KKNMEQTIKDLQHRLDEAEQIALKGGKKQLQKLEARVRELEGELEAEQKRNAESVKGMRK 1844
Query: 173 SWRHI--VTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQE 230
S R I +T TE LRL+DL+ L+ K+ A K E ++
Sbjct: 1845 SERRIKELTYQTEEDKKNLLRLQDLVDKLQL--------------KVKAYKRQAEEAEEQ 1890
Query: 231 LEMN---FDNQQQELDEEDHQDQI 251
N F Q ELDE + + I
Sbjct: 1891 ANTNLSKFRKVQHELDEAEERADI 1914
>RA50_ARCFU (O29230) DNA double-strand break repair rad50 ATPase
Length = 886
Score = 48.1 bits (113), Expect = 2e-04
Identities = 101/527 (19%), Positives = 212/527 (40%), Gaps = 91/527 (17%)
Query: 89 RVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFE-----MKETETIDEMYGRF 143
+ +E K K + + + S + + D+ R+ +L + + +E +
Sbjct: 275 KAKEVKELKPKAERYSILEKLLSEINQALRDVEKREGDLTREAAGIQAQLKKAEEDNSKL 334
Query: 144 TIIMNELRSLEKDFTIHERVRKILRCLPRSWRHI--VTAITEAKDLKKLRLE---DLIGS 198
I + LE++ E+ ++L L + + A E K+L ++E DL+
Sbjct: 335 EEITKRIEELERELERFEKSHRLLETLKPKMDRMQGIKAKLEEKNLTPDKVEKMYDLLSK 394
Query: 199 LKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQ------QELDEEDHQDQII 252
K E + E +K +LKT L + +E ++ +ELDEE ++ +
Sbjct: 395 AKEEEKEITEKLKKLIAKKSSLKTRGAQLKKAVEELKSAERTCPVCGRELDEEHRKNIMA 454
Query: 253 LLTRKLQRM---IQRRDQNKRNFPARKENAKTELDK------------------SQVTCY 291
TR+++R+ + + D+ ++ R E + L+K ++++ +
Sbjct: 455 EYTREMKRIAEELAKADEIEKKLKERLEKVEKALEKQETVLKYRQMVDELKALENELSSH 514
Query: 292 GCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLM-AKSDNEEV 350
KL E K + L+ QK +++ +E + + + L +S+ E+
Sbjct: 515 DAEKLSAESEEYRKVKERLDGLRGQQKILLSSASRIKELKSSLREIEEALKNVESERGEL 574
Query: 351 NLDPCSSCEKTEHLFDNLFYSSQMLEQECERLR---NENLELR----------KEKEILE 397
+ + + F S + LE+E + LR N+ LEL+ K +E LE
Sbjct: 575 H----------RKIREEGFESLEELEREVQSLRPFYNKWLELKDAESRLESELKRREKLE 624
Query: 398 KENKDLIKNIQHSESK------EVSE-NKNNSQKENIILKENVLKLKNDISNFVKSTETF 450
E + I ++ + K ++ E + S++E+ L + L+ +++ ET
Sbjct: 625 DEISEAIAKLEEANGKAEEIRGQIDELLRIYSEEEHRRLSDEHLRKSKELAGLKSRLETL 684
Query: 451 Q----------KIMGSQVSVFDKACIGFKTNQKQKLYENFFIPKKNKNTCERYKCTYCEK 500
+ K + Q++ D + +K +++E IP+ + E+++ K
Sbjct: 685 RESLQSAEKDLKFLEEQLAKMD------EYRKKVEVFEKIAIPELTR-IREKFR-----K 732
Query: 501 YGHL-EPFCFRKRKRLASQKFKNNYTNFHKKETLNNTNHQGPKKIWV 546
Y +L R+ +R ASQ F+ + L T +G +K+ V
Sbjct: 733 YRNLVAENSMREVERYASQIFEELTEGKYSGVRLKKTTERGKEKLKV 779
>MYH7_RAT (P02564) Myosin heavy chain, cardiac muscle beta isoform
(MyHC-beta)
Length = 1935
Score = 48.1 bits (113), Expect = 2e-04
Identities = 89/440 (20%), Positives = 173/440 (39%), Gaps = 50/440 (11%)
Query: 53 NEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECK-NAKEIWDTLKVHHEGTS 111
N+K++ D ++ L A++ AL + +++E + +E+ + L+ +
Sbjct: 1071 NDKQQLDERLKKKDFELNALNARIEDEQALGSQLQKKLKELQARIEELEEELEAERTARA 1130
Query: 112 HVKETRIDIGVRKFELFEMKE------TETIDEMYGRFTIIMNELRSLEKDFTIHERVRK 165
V++ R D+ E+ E E + I+ R R LE+ HE
Sbjct: 1131 KVEKLRSDLSRELEEISERLEEAGGATSVQIEMNKKREAEFQKMRRDLEEATLQHEATAA 1190
Query: 166 ILRCLPRSWRHIVTAITEAKDLKKL-RLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQ 224
LR +H + + + L R++ + K+ L +D +SN ++I K N
Sbjct: 1191 ALRK-----KHADSVAELGEQIDNLQRVKQKLEKEKSEFKLELDDVTSNMEQIIKAKANL 1245
Query: 225 ESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELD 284
E + + LE + + + +E Q + LTR+ ++ + R KE ++L
Sbjct: 1246 EKMCRTLEDQMNEHRSKAEET--QRSVNDLTRQRAKLQTENGELSRQLD-EKEALISQLT 1302
Query: 285 KSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCL--- 341
+ ++T +L K + + + L +S D EQ E+ +A L
Sbjct: 1303 RGKLTY--TQQLEDLKRQLEEEVKAKNALAHALQSARHDCDLLREQYEEETEAKAELQRV 1360
Query: 342 MAKSDNE------EVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEI 395
++K+++E + D E+ E L Q E+ E + + L K K
Sbjct: 1361 LSKANSEVAQWRTKYETDAIQRTEELEEAKKKLAQRLQDAEEAVEAVNAKCSSLEKTKHR 1420
Query: 396 LEKENKDLIKNIQHSESKEVSENKN-----------------------NSQKENIILKEN 432
L+ E +DL+ +++ S + + +K +SQKE L
Sbjct: 1421 LQNEIEDLMVDVERSNAAAAALDKKQRNFDKILVEWKQKYEESQSELESSQKEARSLSTE 1480
Query: 433 VLKLKNDISNFVKSTETFQK 452
+ KLKN ++ ETF++
Sbjct: 1481 LFKLKNAYEESLEHLETFKR 1500
Score = 35.0 bits (79), Expect = 1.6
Identities = 56/271 (20%), Positives = 113/271 (41%), Gaps = 46/271 (16%)
Query: 178 VTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDN 237
V +T+AK + +++DL GSL + + + + + + LK QES+ M+ +N
Sbjct: 1017 VNTLTKAKVKLEQQVDDLEGSLDQDKKVRMDLERAKRKLEGDLKLTQESI-----MDLEN 1071
Query: 238 QQQELDE----------------EDHQ---DQIILLTRKLQRMIQRRDQN---KRNFPAR 275
+Q+LDE ED Q Q+ ++LQ I+ ++ +R A+
Sbjct: 1072 DKQQLDERLKKKDFELNALNARIEDEQALGSQLQKKLKELQARIEELEEELEAERTARAK 1131
Query: 276 KENAKTELDK--SQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNE 333
E +++L + +++ G + +NK++ + Q ++ EE T +
Sbjct: 1132 VEKLRSDLSRELEEISERLEEAGGATSVQIEMNKKREAEFQKMRRDL-------EEATLQ 1184
Query: 334 DEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEK 393
E L K S + DNL Q LE+E + E ++
Sbjct: 1185 HEATAAALRKKH---------ADSVAELGEQIDNLQRVKQKLEKEKSEFKLELDDVTSNM 1235
Query: 394 EILEKENKDLIKNIQHSESKEVSENKNNSQK 424
E + K +L K + E +++E+++ +++
Sbjct: 1236 EQIIKAKANLEKMCRTLED-QMNEHRSKAEE 1265
Score = 34.3 bits (77), Expect = 2.8
Identities = 49/204 (24%), Positives = 88/204 (43%), Gaps = 35/204 (17%)
Query: 69 LLNFKAKLFLTMA-LSREEYDRVQECKNAKE-----------IWDTLKVHHEGTSHVKET 116
L+N K K+ ++ L E + VQEC+NA+E + + LK + ++H++
Sbjct: 1723 LINQKKKMDADLSQLQTEVEEAVQECRNAEEKAKKAITDAAMMAEELKKEQDTSAHLERM 1782
Query: 117 RIDIGVRKFEL-FEMKETETIDEMYGRFTI--IMNELRSLEKDFTIHE-RVRKILRCLPR 172
+ ++ +L + E E I G+ + + +R LE + + R + ++ + +
Sbjct: 1783 KNNMEQTIKDLQHRLDEAEQIALKGGKKQLQKLEARVRELENELEAEQKRNAESVKGMRK 1842
Query: 173 SWRHI--VTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQE 230
S R I +T TE LRL+DL+ L+ K+ A K E ++
Sbjct: 1843 SERRIKELTYQTEEDRKNLLRLQDLVDKLQL--------------KVKAYKRQAEEAEEQ 1888
Query: 231 LEMN---FDNQQQELDEEDHQDQI 251
N F Q ELDE + + I
Sbjct: 1889 ANTNLSKFRKVQHELDEAEERADI 1912
>NIN_MOUSE (Q61043) Ninein
Length = 2035
Score = 47.8 bits (112), Expect = 2e-04
Identities = 85/458 (18%), Positives = 182/458 (39%), Gaps = 97/458 (21%)
Query: 62 EEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIG 121
+EE + + ++ L L+ E Y E K +E+ +T + VK +
Sbjct: 1251 QEELRLVETRYEESLDSNKELTAEVYRLQDEMKKMEEVMETFLSLEKSYDEVK-----VE 1305
Query: 122 VRKFELFEMKETETIDEMYGRFTIIMN------------ELRSLEKDFTIHERVRKILRC 169
+ ++ ++++ GR + + E+ S EK + + ++ C
Sbjct: 1306 NEELRALVLRLQGKMEKVLGRAALQGDSYALWEAPSENLEVASDEKMLELRQTPKE---C 1362
Query: 170 LPR--SWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVL-------LQEDKSSNKSKMI-- 218
P+ S HI+ T+ + L+ +KAHE+ +K S ++++I
Sbjct: 1363 TPKVVSMHHIIEECTQETQCCEQGSTKLLARIKAHEIAWFHRAIKTHPEKPSAQNRVIPE 1422
Query: 219 ----ALKTNQESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPA 274
L + L QE + + EL+++ Q+ L ++ +++++D +
Sbjct: 1423 GSAALLGLQDKHLQQEATI----AELELEKQKLQELTRNLRERVTALVRQKDAPSQG--Q 1476
Query: 275 RKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQ---KSFMTTWDEPEE-- 329
++E K + Q+TC G + + L + +S LQ ++ +TT +E +
Sbjct: 1477 KEEELKAMMQDLQITC------GEMQRKVELLRYESEKLQEENSILRNEITTLNEEDSIS 1530
Query: 330 ------------------QTNEDEQADLCLMAKS---------------DNEEVNLDPCS 356
+T E E+A + M + D+E + L +
Sbjct: 1531 NLKLEELNGSQEELWQKIETIEQEKASIQTMVEKLKKQVSDLKIKNQQLDSENIELSQKN 1590
Query: 357 SCEKTEHLFDNLFYSSQMLEQE------CERLRNENLELRKEKEILEKENKDLIKNIQHS 410
S K E N + + ++E E+ EN L++E + + + L+ +++ +
Sbjct: 1591 SQNKEELKTLNQRLAEMLCQREEPGACTSEKWEQENASLKEELDHYKVQTSTLVSSLE-A 1649
Query: 411 ESKEVSENKNNSQKENIILKENVLKLKN-----DISNF 443
E EV + ++EN++LK+ + +LK D+S+F
Sbjct: 1650 ELSEVKLQTHVMEQENLLLKDELERLKQLHRCPDLSDF 1687
Score = 41.2 bits (95), Expect = 0.023
Identities = 61/275 (22%), Positives = 118/275 (42%), Gaps = 48/275 (17%)
Query: 175 RHIVTAITEAKDLKKLRLEDLIGS---LKAHEVLLQEDKSSNKSKMIALKTNQESL---N 228
R+ +T + E + L+LE+L GS L ++++K+S ++ + LK L N
Sbjct: 1517 RNEITTLNEEDSISNLKLEELNGSQEELWQKIETIEQEKASIQTMVEKLKKQVSDLKIKN 1576
Query: 229 QELEMNFDNQQQELDEEDHQ--DQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKS 286
Q+L D++ EL +++ Q +++ L ++L M+ +R++ P + K E + +
Sbjct: 1577 QQL----DSENIELSQKNSQNKEELKTLNQRLAEMLCQREE-----PGACTSEKWEQENA 1627
Query: 287 QVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSD 346
+ +L HYK + T S E + QT+ EQ +L L +
Sbjct: 1628 SLK----EELDHYKVQT----------STLVSSLEAELSEVKLQTHVMEQENLLLKDE-- 1671
Query: 347 NEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKN 406
L+ + L D +Q+ + + N +L KEKE+L +E K
Sbjct: 1672 -----LERLKQLHRCPDLSD--------FQQKMSSILSYNEKLLKEKEVLSEELKSCADK 1718
Query: 407 IQHSESKEVSENKNNSQKENIILKENVLKLKNDIS 441
+ +ES + ++E +E LK+ ++
Sbjct: 1719 L--AESSLLEHRIATMKQEQTAWEEQSESLKSQLA 1751
Score = 36.6 bits (83), Expect = 0.56
Identities = 57/295 (19%), Positives = 114/295 (38%), Gaps = 25/295 (8%)
Query: 192 LEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDHQDQI 251
LE + L++ L+ + + E ++L+M FD ++ +L EE Q+
Sbjct: 665 LEKRVSELRSEIADLEGQAAVLREAHHKASCRHEEEKRQLQMAFDEEKAQLQEELRQEHE 724
Query: 252 ILLTRKLQRMIQRRDQNKRNF--PARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRK 309
L +LQ+ + Q + A E L++S Y+ + + K
Sbjct: 725 RELQARLQQAAESFRQEREGLAQAAWTEEKVRGLEQS------------YQEQLLSLEEK 772
Query: 310 SSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEK-TEHLFDNL 368
+ + + ++ E Q +E C S ++ + CEK TEH L
Sbjct: 773 HALEKEELREELSEHHRRELQEGREEMETECNRRVS---QIEAQCQADCEKVTEHCEQTL 829
Query: 369 FYSSQMLEQECERLRNENLELRK----EKEILEKENKDLIKNIQHSESKEVSENKNNSQK 424
QE L +++LE R EK+ L +E D + ++ + +E + Q+
Sbjct: 830 QSLEVRHRQELRDLLDQHLEERSQWEFEKDELTQECTDAQEQLKEALQRERATAAAMKQE 889
Query: 425 ENII---LKENVLKLKNDISNFVKSTETFQKIMGSQVSVFDKACIGFKTNQKQKL 476
+ I+ K+ + L + ++ + Q SQ + + K +Q+++L
Sbjct: 890 QEILERTYKDRLNILSTEREQLLQDLKDLQNASESQHGLLSGQILELKRSQEREL 944
>MYSN_DROME (Q99323) Myosin heavy chain, non-muscle (Zipper protein)
(Myosin II) (Non-muscle MHC)
Length = 2057
Score = 47.8 bits (112), Expect = 2e-04
Identities = 67/340 (19%), Positives = 135/340 (39%), Gaps = 55/340 (16%)
Query: 132 ETETIDEMYGRFTIIMN------ELRSLEKDFTIHERVRK-ILRCLPRSWRHIVTAITEA 184
ETE +E R + + +L+ +E +H +V++ L+ + + A+ +A
Sbjct: 1686 ETELDEERKQRTAAVASKKKLEGDLKEIETTMEMHNKVKEDALKHAKKLQAQVKDALRDA 1745
Query: 185 KDLKKLRLE---------DLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNF 235
++ K + E + +L+A + L ED +S++ A +T ++ L +E+ N
Sbjct: 1746 EEAKAAKEELQALSKEADGKVKALEAEVLQLTEDLASSERARRAAETERDELAEEIANNA 1805
Query: 236 DNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYK 295
+ +DE KR AR + EL++ Q
Sbjct: 1806 NKGSLMIDE------------------------KRRLEARIATLEEELEEEQSN------ 1835
Query: 296 LGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPC 355
+E L++ + + LQ Q + ++ Q NE+ +A L K ++
Sbjct: 1836 -----SEVLLDRSRKAQLQIEQLTTELANEKSNSQKNENGRALLERQNKELKAKLAEIET 1890
Query: 356 SSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIKNIQHSESKEV 415
+ K + L LE++ E E L +K ++K+ K+L NI+ E + V
Sbjct: 1891 AQRTKVKATIATLEAKIANLEEQLENEGKERLLQQKANRKMDKKIKELTMNIE-DERRHV 1949
Query: 416 SENKNNSQKENI---ILKENVLKLKNDISNFVKSTETFQK 452
++K K N +LK N+ + + ++ +Q+
Sbjct: 1950 DQHKEQMDKLNSRIKLLKRNLDETEEELQKEKTQKRKYQR 1989
Score = 47.4 bits (111), Expect = 3e-04
Identities = 82/368 (22%), Positives = 156/368 (42%), Gaps = 25/368 (6%)
Query: 79 TMALSREEYDRV--QECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEM-KETET 135
T+A + +EY+R Q + + L+ E + +E+R + RK EL +M +E ET
Sbjct: 954 TLAKNTQEYERKYQQALVEKTTLAEQLQAEIELCAEAEESRSRLMARKQELEDMMQELET 1013
Query: 136 -IDEMYGRFTIIMNELRSLEKDFT-IHERVRKILRCLPRSWRHIVTAITEAKDLKK-LRL 192
I+E R + E + LE + + E++ + + V + K ++ L L
Sbjct: 1014 RIEEEEERVLALGGEKKKLELNIQDLEEQLEEEEAARQKLQLEKVQLDAKIKKYEEDLAL 1073
Query: 193 EDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDHQDQII 252
D E L E+++++ S+ +A + + +L+ + EL+E H+D
Sbjct: 1074 TDDQNQKLLKEKKLLEERANDLSQTLAEEEEKAKHLAKLKAKHEATITELEERLHKD--- 1130
Query: 253 LLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCY-KLGHYKNECPLNKRKSS 311
Q+ Q D++KR + K +L++ +V +L + E +
Sbjct: 1131 ------QQQRQESDRSKRKIETEVADLKEQLNERRVQVDEMQAQLAKREEELTQTLLRID 1184
Query: 312 NLQTNQKSFMTTWDEPEEQ---TNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNL 368
+ + E E Q ED +A+ AK++ +L K E L D+L
Sbjct: 1185 EESATKATAQKAQRELESQLAEIQEDLEAEKAARAKAEKVRRDLSEELEALKNE-LLDSL 1243
Query: 369 FYSSQMLEQECERLRNENL-ELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENI 427
+ +QE R + L L+K E ++ ++ +++H S+E+ N N Q EN+
Sbjct: 1244 --DTTAAQQELRSKREQELATLKKSLEEETVNHEGVLADMRHKHSQEL--NSINDQLENL 1299
Query: 428 ILKENVLK 435
+ VL+
Sbjct: 1300 RKAKTVLE 1307
Score = 40.0 bits (92), Expect = 0.051
Identities = 82/402 (20%), Positives = 149/402 (36%), Gaps = 63/402 (15%)
Query: 54 EKKKEDWSEEEG--KRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTS 111
EK + D SEE K LL+ L T A R QE K+ + V+HEG
Sbjct: 1222 EKVRRDLSEELEALKNELLD---SLDTTAAQQELRSKREQELATLKKSLEEETVNHEGV- 1277
Query: 112 HVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRC-- 169
L +M+ + + I ++L +L K T+ E+ + L
Sbjct: 1278 ---------------LADMRHKHSQE-----LNSINDQLENLRKAKTVLEKAKGTLEAEN 1317
Query: 170 --LPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESL 227
L R + ++ E D ++ + E I L+ ++ +S + K L+ E++
Sbjct: 1318 ADLATELRSVNSSRQE-NDRRRKQAESQIAELQVKLAEIERARSELQEKCTKLQQEAENI 1376
Query: 228 NQELE-------------MNFDNQ----QQELDEEDHQD--------QIILLTRKLQRMI 262
+LE N ++Q QQ L+EE Q QI LQ +
Sbjct: 1377 TNQLEEAELKASAAVKSASNMESQLTEAQQLLEEETRQKLGLSSKLRQIESEKEALQEQL 1436
Query: 263 QRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMT 322
+ D+ KRN+ + T++ + + L E K Q +
Sbjct: 1437 EEDDEAKRNYERKLAEVTTQMQEIKKKAEEDADLAKELEEGKKRLNKDIEALERQVKELI 1496
Query: 323 TWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERL 382
++ +++ + Q++L + E EK + FD + + + ++ +
Sbjct: 1497 AQNDRLDKSKKKIQSEL--EDATIELEAQRTKVLELEKKQKNFDKILAEEKAISEQIAQE 1554
Query: 383 RNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQK 424
R+ E+E EKE K L + + E+ + E+ N +K
Sbjct: 1555 RDT-----AEREAREKETKVLSVSRELDEAFDKIEDLENKRK 1591
Score = 37.7 bits (86), Expect = 0.25
Identities = 81/444 (18%), Positives = 168/444 (37%), Gaps = 48/444 (10%)
Query: 48 YDEDLNEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHH 107
Y+EDL ++ + K++L L T+A E+ + + K E T
Sbjct: 1067 YEEDLALTDDQNQKLLKEKKLLEERANDLSQTLAEEEEKAKHLAKLKAKHEATITELEER 1126
Query: 108 EGTSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKIL 167
+ D RK E E + E + ++E+++ + +R ++
Sbjct: 1127 LHKDQQQRQESDRSKRKIET----EVADLKEQLNERRVQVDEMQA-----QLAKREEELT 1177
Query: 168 RCLPRSWRHIVTAITEAKDLKKL--RLEDLIGSLKAHEVL----------LQEDKSSNKS 215
+ L R T T K ++L +L ++ L+A + L E+ + K+
Sbjct: 1178 QTLLRIDEESATKATAQKAQRELESQLAEIQEDLEAEKAARAKAEKVRRDLSEELEALKN 1237
Query: 216 KMI-ALKTN--QESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNF 272
+++ +L T Q+ L + E ++ L+EE + +L + + + N +
Sbjct: 1238 ELLDSLDTTAAQQELRSKREQELATLKKSLEEETVNHEGVLADMRHKHSQELNSINDQLE 1297
Query: 273 PARKENAKTELDKSQVTCYG-----CYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEP 327
RK AKT L+K++ T +L + N R+ ++ E
Sbjct: 1298 NLRK--AKTVLEKAKGTLEAENADLATELRSVNSSRQENDRRRKQAESQIAELQVKLAEI 1355
Query: 328 EEQTNEDEQADLCLMAKSDN-----EEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECER- 381
E +E ++ L +++N EE L ++ + ++ L + Q+LE+E +
Sbjct: 1356 ERARSELQEKCTKLQQEAENITNQLEEAELKASAAVKSASNMESQLTEAQQLLEEETRQK 1415
Query: 382 --LRNENLELRKEKEILEKENKDLIKNIQHSESKEVSENKNNSQKENIILKENVLKLKND 439
L ++ ++ EKE L++ Q E E N E + + K +
Sbjct: 1416 LGLSSKLRQIESEKEALQE---------QLEEDDEAKRNYERKLAEVTTQMQEIKKKAEE 1466
Query: 440 ISNFVKSTETFQKIMGSQVSVFDK 463
++ K E +K + + ++
Sbjct: 1467 DADLAKELEEGKKRLNKDIEALER 1490
Score = 37.4 bits (85), Expect = 0.33
Identities = 76/410 (18%), Positives = 160/410 (38%), Gaps = 47/410 (11%)
Query: 46 IPYDEDLNEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRV-QECKNAKEIWDTLK 104
I ++ L++ KK+ SE E + L + L + ++ +D++ E K E +
Sbjct: 1496 IAQNDRLDKSKKKIQSELEDATIELEAQRTKVLELEKKQKNFDKILAEEKAISEQIAQER 1555
Query: 105 VHHEGTSHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVR 164
E + KET++ + + +DE + + + N+ ++L+ + +
Sbjct: 1556 DTAEREAREKETKV-----------LSVSRELDEAFDKIEDLENKRKTLQNELDDLANTQ 1604
Query: 165 KILRCLPRSWRHIVTAITEAKDLKKLR--LEDLIGSLKAHEVLLQEDKSSNKSKMIALKT 222
TA +L+K + LE + LKA L++D + + L+
Sbjct: 1605 G-------------TADKNVHELEKAKRALESQLAELKAQNEELEDDLQLTEDAKLRLEV 1651
Query: 223 NQESLNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTE 282
N ++L + E D +E E+ + ++ R L+ + + + A K+ + +
Sbjct: 1652 NMQALRSQFER--DLLAKEEGAEEKRRGLVKQLRDLETELDEERKQRTAAVASKKKLEGD 1709
Query: 283 LDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLM 342
L + + T ++ + E L K LQ K + +E + E L +
Sbjct: 1710 LKEIETT----MEMHNKVKEDALKHAK--KLQAQVKDALRDAEEAKAAKEE-----LQAL 1758
Query: 343 AKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKD 402
+K + +V + TE L + + + E E + L E + ++ E +
Sbjct: 1759 SKEADGKVKALEAEVLQLTEDLASS-ERARRAAETERDELAEEIANNANKGSLMIDEKRR 1817
Query: 403 LIKNIQHSESKEVSENKNN------SQKENIILKENVLKLKNDISNFVKS 446
L I E + E N+ S+K + +++ +L N+ SN K+
Sbjct: 1818 LEARIATLEEELEEEQSNSEVLLDRSRKAQLQIEQLTTELANEKSNSQKN 1867
Score = 35.4 bits (80), Expect = 1.2
Identities = 25/124 (20%), Positives = 64/124 (51%), Gaps = 9/124 (7%)
Query: 170 LPRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQED-KSSNKSKMIALKTNQ--ES 226
L R + + + E + ++ +++ I +L+A L+E ++ K +++ K N+ +
Sbjct: 1874 LERQNKELKAKLAEIETAQRTKVKATIATLEAKIANLEEQLENEGKERLLQQKANRKMDK 1933
Query: 227 LNQELEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKS 286
+EL MN +++++ +D+ H++Q+ L +++ + + D+ + + KT+ K
Sbjct: 1934 KIKELTMNIEDERRHVDQ--HKEQMDKLNSRIKLLKRNLDETEEEL----QKEKTQKRKY 1987
Query: 287 QVTC 290
Q C
Sbjct: 1988 QREC 1991
>MY5A_HUMAN (Q9Y4I1) Myosin Va (Myosin 5A) (Dilute myosin heavy chain,
non-muscle) (Myosin heavy chain 12) (Myoxin)
Length = 1855
Score = 47.8 bits (112), Expect = 2e-04
Identities = 94/420 (22%), Positives = 166/420 (39%), Gaps = 73/420 (17%)
Query: 51 DLNEKKKEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGT 110
+L + K E S E K++ + + K+ M L R+ ++ ++ K E L EG
Sbjct: 904 ELKKLKIEARSVERYKKLHIGMENKI---MQLQRKVDEQNKDYKCLVEKLTNL----EGI 956
Query: 111 SHVKETRIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCL 170
+ + ++ + + +L E E GR + E+ L KD E+ R +C
Sbjct: 957 YNSETEKLRSDLERLQLSE----EEAKVATGRVLSLQEEIAKLRKDL---EQTRSEKKC- 1008
Query: 171 PRSWRHIVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQE 230
I E D K E L+ +LK LL+++K + N + Q
Sbjct: 1009 ----------IEEHADRYKQETEQLVSNLKEENTLLKQEKEA---------LNHRIVQQA 1049
Query: 231 LEMNFDNQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTC 290
EM + +++L EE Q ++ L +L R QN N +R E +L +
Sbjct: 1050 KEMT-ETMEKKLVEETKQLELDLNDERL------RYQNLLNEFSRLEERYDDLKEEMTLM 1102
Query: 291 YGCYKLGHYKNECPLNKRKSSNLQTNQKSFM----TTWDEPEEQ---------------T 331
K GH + + + +S + +++ + M + +EP E+
Sbjct: 1103 VHVPKPGHKRTDSTHSSNESEYIFSSEIAEMEDIPSRTEEPSEKKVPLDMSLFLKLQKRV 1162
Query: 332 NEDEQADLCLMAKSDN--EEVNLDPCSSCEKTEHLFDNLFYSS---QMLEQECERLRNEN 386
E EQ + + D E+V E+ + L Y S Q LE E ++L+NE
Sbjct: 1163 TELEQEKQVMQDELDRKEEQVLRSKAKEEERPQIRGAELEYESLKRQELESENKKLKNEL 1222
Query: 387 LELRKEKEILEKENKDLIK------NIQHSESKEVSENKNNSQKENIILKENVLKLKNDI 440
ELR K + EK ++ + + VSE + ++E +IL+ ++ K I
Sbjct: 1223 NELR--KALSEKSAPEVTAPGAPAYRVLMEQLTSVSEELDVRKEEVLILRSQLVSQKEAI 1280
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.355 0.157 0.539
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 150,000,480
Number of Sequences: 164201
Number of extensions: 5698057
Number of successful extensions: 46716
Number of sequences better than 10.0: 480
Number of HSP's better than 10.0 without gapping: 19
Number of HSP's successfully gapped in prelim test: 489
Number of HSP's that attempted gapping in prelim test: 43551
Number of HSP's gapped (non-prelim): 2305
length of query: 1574
length of database: 59,974,054
effective HSP length: 124
effective length of query: 1450
effective length of database: 39,613,130
effective search space: 57439038500
effective search space used: 57439038500
T: 11
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 73 (32.7 bits)
Medicago: description of AC144538.13


