

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
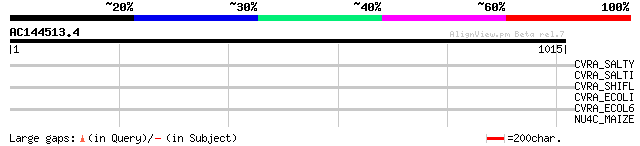
Query= AC144513.4 - phase: 0 /pseudo
(1015 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CVRA_SALTY (P65524) Cell volume regulation protein A 35 1.3
CVRA_SALTI (P65525) Cell volume regulation protein A 35 1.3
CVRA_SHIFL (Q83RQ1) Cell volume regulation protein A 33 3.0
CVRA_ECOLI (P76007) Cell volume regulation protein A 33 3.0
CVRA_ECOL6 (P59239) Cell volume regulation protein A 33 3.0
NU4C_MAIZE (P11647) NAD(P)H-quinone oxidoreductase chain 4, chlo... 33 5.1
>CVRA_SALTY (P65524) Cell volume regulation protein A
Length = 577
Score = 34.7 bits (78), Expect = 1.3
Identities = 30/106 (28%), Positives = 47/106 (44%), Gaps = 3/106 (2%)
Query: 647 FMVSKLLLMPPALLTFFMQMAIFSFVRLTLMRPLLYPTFFLNTKLFLARKLILISQIWSL 706
F+V LL+ P L + I S + RPL T L + F R+ I IS +
Sbjct: 283 FLVLGLLVTPSDLWPIAVPALILSIWMIFFARPLSVFTGLLPFRGFNLRERIFISWVGLR 342
Query: 707 AQIPVLVLKIYSELISLSTSRIIFLSIWGFLLSLEDLKLKILSLLW 752
+P+ +L ++ + L +R+ F F + L L L+ SL W
Sbjct: 343 GAVPI-ILAVFPMMAGLENARLFFNV--AFFVVLVSLLLQGTSLSW 385
>CVRA_SALTI (P65525) Cell volume regulation protein A
Length = 577
Score = 34.7 bits (78), Expect = 1.3
Identities = 30/106 (28%), Positives = 47/106 (44%), Gaps = 3/106 (2%)
Query: 647 FMVSKLLLMPPALLTFFMQMAIFSFVRLTLMRPLLYPTFFLNTKLFLARKLILISQIWSL 706
F+V LL+ P L + I S + RPL T L + F R+ I IS +
Sbjct: 283 FLVLGLLVTPSDLWPIAVPALILSIWMIFFARPLSVFTGLLPFRGFNLRERIFISWVGLR 342
Query: 707 AQIPVLVLKIYSELISLSTSRIIFLSIWGFLLSLEDLKLKILSLLW 752
+P+ +L ++ + L +R+ F F + L L L+ SL W
Sbjct: 343 GAVPI-ILAVFPMMAGLENARLFFNV--AFFVVLVSLLLQGTSLSW 385
>CVRA_SHIFL (Q83RQ1) Cell volume regulation protein A
Length = 578
Score = 33.5 bits (75), Expect = 3.0
Identities = 29/106 (27%), Positives = 47/106 (43%), Gaps = 3/106 (2%)
Query: 647 FMVSKLLLMPPALLTFFMQMAIFSFVRLTLMRPLLYPTFFLNTKLFLARKLILISQIWSL 706
F+V LL+ P LL + I S + RPL L + F R+ + IS +
Sbjct: 283 FLVLGLLVNPSDLLPIAIPALILSAWMIFFARPLSVFAGLLPFRGFNLRERVFISWVGLR 342
Query: 707 AQIPVLVLKIYSELISLSTSRIIFLSIWGFLLSLEDLKLKILSLLW 752
+P+ +L ++ + L +R+ F F + L L L+ SL W
Sbjct: 343 GAVPI-ILAVFPMMAGLENARLFFNV--AFFVVLVSLLLQGTSLSW 385
>CVRA_ECOLI (P76007) Cell volume regulation protein A
Length = 578
Score = 33.5 bits (75), Expect = 3.0
Identities = 29/106 (27%), Positives = 47/106 (43%), Gaps = 3/106 (2%)
Query: 647 FMVSKLLLMPPALLTFFMQMAIFSFVRLTLMRPLLYPTFFLNTKLFLARKLILISQIWSL 706
F+V LL+ P LL + I S + RPL L + F R+ + IS +
Sbjct: 283 FLVLGLLVNPSDLLPIAIPALILSAWMIFFARPLSVFAGLLPFRGFNLRERVFISWVGLR 342
Query: 707 AQIPVLVLKIYSELISLSTSRIIFLSIWGFLLSLEDLKLKILSLLW 752
+P+ +L ++ + L +R+ F F + L L L+ SL W
Sbjct: 343 GAVPI-ILAVFPMMAGLENARLFFNV--AFFVVLVSLLLQGTSLSW 385
>CVRA_ECOL6 (P59239) Cell volume regulation protein A
Length = 578
Score = 33.5 bits (75), Expect = 3.0
Identities = 29/106 (27%), Positives = 47/106 (43%), Gaps = 3/106 (2%)
Query: 647 FMVSKLLLMPPALLTFFMQMAIFSFVRLTLMRPLLYPTFFLNTKLFLARKLILISQIWSL 706
F+V LL+ P LL + I S + RPL L + F R+ + IS +
Sbjct: 283 FLVLGLLVNPSDLLPIAIPALILSAWMIFFARPLSVFAGLLPFRGFNLRERVFISWVGLR 342
Query: 707 AQIPVLVLKIYSELISLSTSRIIFLSIWGFLLSLEDLKLKILSLLW 752
+P+ +L ++ + L +R+ F F + L L L+ SL W
Sbjct: 343 GAVPI-ILAVFPMMAGLENARLFFNV--AFFVVLVSLLLQGTSLSW 385
>NU4C_MAIZE (P11647) NAD(P)H-quinone oxidoreductase chain 4,
chloroplast (EC 1.6.5.-) (NAD(P)H dehydrogenase, chain
4) (NADH-plastoquinone oxidoreductase chain 4)
Length = 500
Score = 32.7 bits (73), Expect = 5.1
Identities = 28/86 (32%), Positives = 46/86 (52%), Gaps = 12/86 (13%)
Query: 651 KLLLMPPALLTFFMQ--MAIFSFVRLTLMRPLL--YPTFFLNTKLFL---ARKLILISQI 703
K +LMP L+TF M M + L+++R + Y F + K F+ R+L L+ I
Sbjct: 409 KFMLMPKMLITFVMAIGMILTPIYLLSMLRQMFYGYKLFHVPNKNFVDSGPRELFLLICI 468
Query: 704 WSLAQIPVLVLKIYSELI-SLSTSRI 728
+ +PV+ + IY +L+ SLS R+
Sbjct: 469 F----LPVIGIGIYPDLVLSLSVDRV 490
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.368 0.167 0.619
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 94,866,750
Number of Sequences: 164201
Number of extensions: 3313756
Number of successful extensions: 18458
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 18453
Number of HSP's gapped (non-prelim): 12
length of query: 1015
length of database: 59,974,054
effective HSP length: 120
effective length of query: 895
effective length of database: 40,269,934
effective search space: 36041590930
effective search space used: 36041590930
T: 11
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 71 (32.0 bits)
Medicago: description of AC144513.4


