

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144503.9 + phase: 0 /pseudo
(346 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
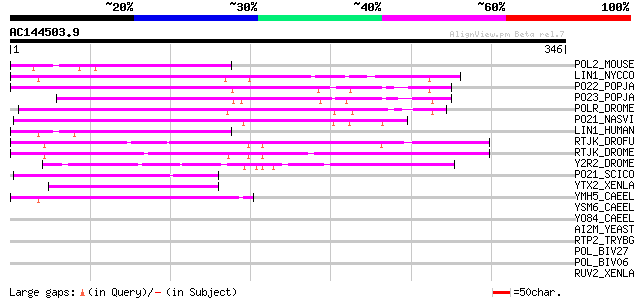
Score E
Sequences producing significant alignments: (bits) Value
POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains... 73 9e-13
LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog 72 2e-12
PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type... 69 2e-11
PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type... 68 3e-11
POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type... 64 6e-10
PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type... 64 6e-10
LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog 60 8e-09
RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile elem... 59 2e-08
RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile elem... 57 5e-08
Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I retro... 54 4e-07
PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type... 50 6e-06
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 49 1e-05
YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III 49 1e-05
YSM6_CAEEL (Q10126) Hypothetical protein F52C9.6 in chromosome III 41 0.004
YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III 40 0.009
AI2M_YEAST (P03876) Putative COX1/OXI3 intron 2 protein 35 0.36
RTP2_TRYBG (P15594) Retrotransposable element SLACS 132 kDa prot... 34 0.47
POL_BIV27 (P19561) Pol polyprotein [Contains: Protease (EC 3.4.2... 33 0.81
POL_BIV06 (P19560) Pol polyprotein [Contains: Protease (EC 3.4.2... 33 0.81
RUV2_XENLA (Q9DE27) RuvB-like 2 (EC 3.6.1.-) (Reptin) 33 1.1
>POL2_MOUSE (P11369) Retrovirus-related Pol polyprotein [Contains:
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1300
Score = 73.2 bits (178), Expect = 9e-13
Identities = 42/144 (29%), Positives = 78/144 (54%), Gaps = 9/144 (6%)
Query: 1 KLWERVIERRLRK--EARVTDNQFGFMPGRLTMEAIYLLRRVIE--RYRTDKKDLH--LV 54
K+ +++ R+++ +A + +Q GF+PG M+ + +R+ I Y KD + ++
Sbjct: 567 KILNKILANRIQEHIKAIIHPDQVGFIPG---MQGWFNIRKSINVIHYINKLKDKNHMII 623
Query: 55 FIDLEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGV 114
+D EK +D++ + K LE+ G++ Y+ IK +Y +++ E P+ G
Sbjct: 624 SLDAEKAFDKIQHPFMIKVLERSGIQGPYLNMIKAIYSKPVANIKVNGEKLEAIPLKSGT 683
Query: 115 HQGSTLSPYLFTLVLDVLTEHIQE 138
QG LSPYLF +VL+VL I++
Sbjct: 684 RQGCPLSPYLFNIVLEVLARAIRQ 707
>LIN1_NYCCO (P08548) LINE-1 reverse transcriptase homolog
Length = 1260
Score = 72.4 bits (176), Expect = 2e-12
Identities = 64/300 (21%), Positives = 133/300 (44%), Gaps = 27/300 (9%)
Query: 1 KLWERVIERRLRKEAR--VTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVF-ID 57
K+ +++ R+++ + + +Q GF+PG I VI+ K H++ ID
Sbjct: 540 KILNKILTNRIQQHIKKIIHHDQVGFIPGSQGWFNIRKSINVIQHINKLKNKDHMILSID 599
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
EK +D + + + L+K G+ +++ I+ +Y + ++ + FP+ G QG
Sbjct: 600 AEKAFDNIQHPFMIRTLKKIGIEGTFLKLIEAIYSKPTANIILNGVKLKSFPLRSGTRQG 659
Query: 118 STLSPYLFTLVLDVLT-----------EHIQELAPRCMLFAD--VVLVGESREEVNGRLE 164
LSP LF +V++VL HI + LFAD +V + +R+ LE
Sbjct: 660 CPLSPLLFNIVMEVLAIAIREEKAIKGIHIGSEEIKLSLFADDMIVYLENTRDSTTKLLE 719
Query: 165 TWRQALEAYGFRL-SRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*ND 223
++ G+++ + + ++ N ++ T++ + ++P+ + KYLG ++ D
Sbjct: 720 VIKEYSNVSGYKINTHKSVAFIYTN--NNQAEKTVKDSIPFTVVPK--KMKYLGVYLTKD 775
Query: 224 GEIEADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGK--FYRTAVRPALLYGTECWAVKS 281
++ E L+ A V K +P G+ + ++ P +Y +K+
Sbjct: 776 ----VKDLYKENYETLRKEIAEDVNKWKNIPCSWLGRINIVKMSILPKAIYNFNAIPIKA 831
>PO22_POPJA (Q03274) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 711
Score = 68.9 bits (167), Expect = 2e-11
Identities = 78/297 (26%), Positives = 124/297 (41%), Gaps = 35/297 (11%)
Query: 1 KLWERVIERRLRKEARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDLEK 60
+L R++ +RL + Q G+ T+ LL I R +K ++V +D+ K
Sbjct: 87 RLLHRILAKRLEAAVELHPAQKGYARIDGTLVNSLLLDTYISSRREQRKTYNVVSLDVRK 146
Query: 61 TYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGT-TEDFPITIGVHQGST 119
+D V + +AL++ G+ I ++T++R G+ T I GV QG
Sbjct: 147 AFDTVSHSSICRALQRLGIDEGTSNYITGSLSDSTTTIRVGPGSQTRKICIRRGVKQGDP 206
Query: 120 LSPYLFTLVLDVLTEHIQ-----------ELAPRCMLFADVVLVGESREEVNGRLETWRQ 168
LSP+LF VLD L +Q E P D++L+ ++ + L T
Sbjct: 207 LSPFLFNAVLDELLCSLQSTPGIGGTIGEEKIPVLAFADDLLLLEDNDVLLPTTLATVAN 266
Query: 169 ALEAYGFRLSRSTTEYMECNFSG----RRSRSTLEVKVGDHIIPQVT----RFKYLGSFV 220
G L+ + + SG R++ L V D+++ VT F+YLG F
Sbjct: 267 FFRLRGMSLNAKKSVSISVAASGGVCIPRTKPFLRV---DNVLLPVTDRMQTFRYLGHFF 323
Query: 221 *NDGEIEADVSHRIQAEWLKWRRASGVLCDKKVPLKLKGK--FYRTAVRPALLYGTE 275
G + V + + WLK C + PLK + K R V P LLYG +
Sbjct: 324 GLSGAAKPTVYN--LSRWLK--------CVEAAPLKPEQKLSLIREHVVPKLLYGLQ 370
>PO23_POPJA (Q05118) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 606
Score = 68.2 bits (165), Expect = 3e-11
Identities = 74/265 (27%), Positives = 111/265 (40%), Gaps = 30/265 (11%)
Query: 30 TMEAIYLLRRVIERYRTDKKDLHLVFIDLEKTYDRVPREILWKALEKKGVRIAYIRAIKD 89
T+ I +L I+ R K ++V +D+ K +D V + +A+ G+ I
Sbjct: 2 TLANIIMLEHYIKLRRLKGKTYNVVSLDIRKAFDTVSHPAILRAMRAFGIDDGMQDFIMS 61
Query: 90 MYEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLFTLVLDVLTEHIQE------LAPRC 143
A T++ TT I GV QG LSP LF +VLD L + + + P C
Sbjct: 62 TITDAYTNIVVGGRTTNKIYIRNGVKQGDPLSPVLFNIVLDELVTRLNDEQPGASMTPAC 121
Query: 144 ----MLFADVVLVGESRE-EVNGRLETWRQALEAYGFRLS-RSTTEYMECNFSGR---RS 194
+ FAD +L+ E R+ +V L T G L+ SGR RS
Sbjct: 122 KIASLAFADDLLLLEDRDIDVPNSLATTCAYFRTRGMTLNPEKCASISAATVSGRSVPRS 181
Query: 195 RSTLEVKVGDHIIP--QVTRFKYLGSFV*NDGEIEADVSHRIQAEWLKWRRASGVLCDKK 252
+ + + G +I P + FKYLG + G + V + WL+ +K
Sbjct: 182 KPSFTID-GRYIKPLGGINTFKYLGLTFSSTGVAKPTVYN--LTRWLRNL--------EK 230
Query: 253 VPLKLKGKFY--RTAVRPALLYGTE 275
PLK KFY +T + P L YG +
Sbjct: 231 APLKPNQKFYILKTHLLPRLFYGLQ 255
>POLR_DROME (P16423) Retrovirus-related Pol polyprotein from type II
retrotransposable element R2DM [Contains: Protease (EC
3.4.23.-); Reverse transcriptase (EC 2.7.7.49);
Endonuclease]
Length = 1057
Score = 63.9 bits (154), Expect = 6e-10
Identities = 66/284 (23%), Positives = 116/284 (40%), Gaps = 26/284 (9%)
Query: 6 VIERRLRKEARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDLEKTYDRV 65
++ RL Q GF+P + ++ V+ + ++ +D+ K +D +
Sbjct: 431 ILATRLNSSINWDPRQRGFLPTDGCADNATIVDLVLRHSHKHFRSCYIANLDVSKAFDSL 490
Query: 66 PREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLF 125
++ L G ++ +++ YEG TS+ ++E+F GV QG LSP LF
Sbjct: 491 SHASIYDTLRAYGAPKGFVDYVQNTYEGGGTSLNGDGWSSEEFVPARGVKQGDPLSPILF 550
Query: 126 TLVLDVLTE--------HIQELAPRCMLFA-DVVLVGESREEVNGRLETWRQALEAYGFR 176
LV+D L + FA D+VL E+R + L+ L G +
Sbjct: 551 NLVMDRLLRTLPSEIGAKVGNAITNAAAFADDLVLFAETRMGLQVLLDKTLDFLSIVGLK 610
Query: 177 LSRSTTEYMECNFSGRRSRSTLEVK---VGDHIIPQVTR---FKYLGSFV*NDGEIEADV 230
L+ + ++ + LE + VG IP + R +KYLG G + +
Sbjct: 611 LNADKCFTVGIKGQPKQKCTVLEAQSFYVGSSEIPSLKRTDEWKYLGINFTATGRVRCNP 670
Query: 231 SHRIQAEWLKWRRASGVLCDKKVPLKLKGKFY--RTAVRPALLY 272
+ I K +R + K PLK + + + RT + P L +
Sbjct: 671 AEDIGP---KLQRLT------KAPLKPQQRLFALRTVLIPQLYH 705
>PO21_NASVI (Q03278) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 1025
Score = 63.9 bits (154), Expect = 6e-10
Identities = 63/267 (23%), Positives = 111/267 (40%), Gaps = 21/267 (7%)
Query: 3 WERVIERRLRKEARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDLEKTY 62
+ R++ R+ + + Q F+ E LL +I+ R K L++ +D++K +
Sbjct: 402 FHRILANRIGEHGLLDTRQRAFIVADGVAENTSLLSAMIKEARMKIKGLYIAILDVKKAF 461
Query: 63 DRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSP 122
D V + AL +K + + I +Y + T + GV QG LSP
Sbjct: 462 DSVEHRSILDALRRKKLPLEMRNYIMWVYRNSKTRLEVVKTKGRWIRPARGVRQGDPLSP 521
Query: 123 YLFTLVLDVLTEHIQELAPRCM--------LFA-DVVLVGESREEVNGRLETWRQALEAY 173
LF V+D + + E M +FA D+VL+ E+RE + L L+
Sbjct: 522 LLFNCVMDAVLRRLPENTGFLMGAEKIGALVFADDLVLLAETREGLQASLSRIEAGLQEQ 581
Query: 174 GFRLSRSTTEYMECNFSGRRSRSTLEV----KVGDHIIPQV---TRFKYLGSFV*NDGEI 226
G + + SG+ + +E VG+ I Q+ ++KYLG + G I
Sbjct: 582 GLEMMPRKCHTLALVPSGKEKKIKVETHKPFTVGNQEITQLGHADQWKYLGVVYNSYGPI 641
Query: 227 EADVS-----HRIQAEWLKWRRASGVL 248
+ ++ R+ A LK ++ +L
Sbjct: 642 QVKINIAGDLQRVTAAPLKPQQRMAIL 668
>LIN1_HUMAN (P08547) LINE-1 reverse transcriptase homolog
Length = 1259
Score = 60.1 bits (144), Expect = 8e-09
Identities = 36/144 (25%), Positives = 75/144 (52%), Gaps = 9/144 (6%)
Query: 1 KLWERVIERRLRKEAR--VTDNQFGFMPGRLTMEAIYLLRR---VIERYRTDKKDLHLVF 55
K+ +++ ++++ + + +Q GF+P M+ + +R+ +I+ K H++
Sbjct: 540 KILNKILANQIQQHIKKLIHHDQVGFIPA---MQGWFNIRKSINIIQHINRTKDTNHMII 596
Query: 56 -IDLEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGV 114
ID EK +D++ + + K L K G+ Y++ I+ +Y+ + ++ E P+ G
Sbjct: 597 SIDAEKAFDKIQQPFMLKPLNKLGIDGTYLKIIRAIYDKPTANIILNGQKLEAPPLKTGT 656
Query: 115 HQGSTLSPYLFTLVLDVLTEHIQE 138
QG LSP L +VL+VL I++
Sbjct: 657 RQGCPLSPLLPNIVLEVLARAIRQ 680
>RTJK_DROFU (P21329) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 58.9 bits (141), Expect = 2e-08
Identities = 79/320 (24%), Positives = 137/320 (42%), Gaps = 28/320 (8%)
Query: 1 KLWERVIERRLRKEARVTDN----QFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFI 56
K+ ER+I R+ VT QFGF T E ++ + K+ F+
Sbjct: 526 KMLERLILNRILTSEEVTRAIPKFQFGFRLQHGTPEQLHRVVNFALEALEKKEYAGSCFL 585
Query: 57 DLEKTYDRVPRE-ILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFP-ITIGV 114
D+++ +DRV +L+KA K + + IK +EG SV T DG I GV
Sbjct: 586 DIQQAFDRVWHPGLLYKA--KSLLSPQLFQLIKSFWEGRKFSV-TADGCRSSVKFIEAGV 642
Query: 115 HQGSTLSPYLFTL-VLDVLTEH-IQELAPRCMLFA----DVVLVGESR--EEVNGRLETW 166
QGS L P L+++ D+ ++ + LA +L A D+ ++ +S E L+ +
Sbjct: 643 PQGSVLGPTLYSIFTADMPNQNAVTGLAEGEVLIATYADDIAVLTKSTCIVEATDALQEY 702
Query: 167 RQALEAYGFRLSRSTTEYMECNFSGRRS-RSTLEVKVGDHIIPQVTRFKYLGSFV*NDGE 225
A + + + + S N + + R V + ++ +KYLG +
Sbjct: 703 LDAFQEWAVKWNVSINAGKCANVTFTNAIRDCPGVTINGSLLSHTHEYKYLGVILDRSLT 762
Query: 226 IEADV-----SHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWAVK 280
+ S R + + W A+ K+ L K K Y+ V P L Y + + +
Sbjct: 763 FRRHITSLQHSFRTRITKMNWLLAAR----NKLSLDNKVKIYKCIVAPGLFYAIQVYGIA 818
Query: 281 SQ-HENQVSVAEMRMLRWMS 299
++ H N++ V + +MLR +S
Sbjct: 819 ARTHLNKIRVLQAKMLRKIS 838
>RTJK_DROME (P21328) RNA-directed DNA polymerase from mobile element
jockey (EC 2.7.7.49) (Reverse transcriptase)
Length = 916
Score = 57.4 bits (137), Expect = 5e-08
Identities = 72/318 (22%), Positives = 137/318 (42%), Gaps = 24/318 (7%)
Query: 1 KLWERVIERRLRKEARVTDN----QFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFI 56
K+ ER+I RL VT QFGF T E ++ + +K+ F+
Sbjct: 526 KIMERLILNRLLTCKDVTKAIPKFQFGFRLQHGTPEQLHRVVNFALEAMENKEYAVGAFL 585
Query: 57 DLEKTYDRVPRE-ILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVH 115
D+++ +DRV +L+KA ++ + +K E + V + PI GV
Sbjct: 586 DIQQAFDRVWHPGLLYKAKRLFPPQLYLV--VKSFLEERTFHVSVDGYKSSIKPIAAGVP 643
Query: 116 QGSTLSPYLFTLVLDVLTEH--IQELAPRCMLFA----DVVLVGESR------EEVNGRL 163
QGS L P L+++ + H + E+ +L A D ++ +S+ + L
Sbjct: 644 QGSVLGPTLYSVFASDMPTHTPVTEVDEEDVLIATYADDTAVLTKSKSILAATSGLQEYL 703
Query: 164 ETWRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*ND 223
+ ++Q E + R++ + R+ S V + +I +KYLG +
Sbjct: 704 DAFQQWAENWNVRINAEKCANVT---FANRTGSCPGVSLNGRLIRHHQAYKYLGITLDRK 760
Query: 224 GEIEADVSHRIQAEWLKWRRASGVLCDK-KVPLKLKGKFYRTAVRPALLYGTECWAVKSQ 282
+++ QA K R S ++ + K+ L K Y++ + P L YG + + + ++
Sbjct: 761 LTFSRHITNIQQAFRTKVARMSWLIAPRNKLSLGCKVNIYKSILAPCLFYGLQVYGIAAK 820
Query: 283 -HENQVSVAEMRMLRWMS 299
H N++ + + + LR +S
Sbjct: 821 SHLNKIRILQAKTLRRIS 838
>Y2R2_DROME (P16425) Hypothetical 115 kDa protein in type I
retrotransposable element R1DM (ORF 2)
Length = 1021
Score = 54.3 bits (129), Expect = 4e-07
Identities = 73/271 (26%), Positives = 111/271 (40%), Gaps = 29/271 (10%)
Query: 21 QFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHL-VFIDLEKTYDRVPREILWKALEKKGV 79
QFGF GR +A R V L F+D + +D V L G
Sbjct: 544 QFGFRQGRCVEDA---WRHVKSSVGASAAQYVLGTFVDFKGAFDNVEWSAALSRLADLGC 600
Query: 80 RIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLFTLVLDVLTEHIQEL 139
R + + + G +R+ GT E P+T G QGS P+++ +++DVL +Q L
Sbjct: 601 R--EMGLWQSFFSGRRAVIRSSSGTVE-VPVTRGCPQGSISGPFIWDILMDVL---LQRL 654
Query: 140 APRCML--FADVVLV---GESR---EEVNGRL----ETWRQALEAYGFRLSRSTTEYMEC 187
P C L +AD +L+ G SR EE +L ETW + G LS S T M
Sbjct: 655 QPYCQLSAYADDLLLLVEGNSRAVLEEKGAQLMSIVETWGAEV---GDCLSTSKTVIMLL 711
Query: 188 NFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGEIEADV-SHRIQAEWLKWRRASG 246
+ RR+ + V+ +P V +YLG V + + S R + + A
Sbjct: 712 KGALRRAPT---VRFAGRNLPYVRSCRYLGITVSEGMKFLTHIASLRQRMTGVVGALARV 768
Query: 247 VLCDKKVPLKLKGKFYRTAVRPALLYGTECW 277
+ D + + Y + P +L+G W
Sbjct: 769 LRADWGFSPRARRTIYDGLMAPCVLFGAPVW 799
>PO21_SCICO (Q03279) Retrovirus-related Pol polyprotein from type I
retrotransposable element R2 [Contains: Reverse
transcriptase (EC 2.7.7.49); Endonuclease] (Fragment)
Length = 869
Score = 50.4 bits (119), Expect = 6e-06
Identities = 34/128 (26%), Positives = 59/128 (45%), Gaps = 1/128 (0%)
Query: 3 WERVIERRLRKEARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDLEKTY 62
+ +++ RR + Q ++P + +L +I + +K+LH+ +DL K +
Sbjct: 243 FNKILARRFVSCYTYDERQTAYLPIDGVCINVSMLTAIIAEAKRLRKELHIAILDLVKAF 302
Query: 63 DRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSP 122
+ V L A+ + G + I DMY T ++ +G E I GV+QG LS
Sbjct: 303 NSVYHSALIDAITEAGCPPGVVDYIADMYNNVITEMQF-EGKCELASILAGVYQGDPLSG 361
Query: 123 YLFTLVLD 130
LFTL +
Sbjct: 362 PLFTLAYE 369
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein
(ORF 2)
Length = 1308
Score = 49.3 bits (116), Expect = 1e-05
Identities = 29/106 (27%), Positives = 49/106 (45%)
Query: 25 MPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDLEKTYDRVPREILWKALEKKGVRIAYI 84
+PGR + ++L+R ++ R L + +D EK +DRV + L L+ ++
Sbjct: 562 VPGRTIFDNVFLIRDLLHFARRTGLSLAFLSLDQEKAFDRVDHQYLIGTLQAYSFGPQFV 621
Query: 85 RAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLFTLVLD 130
+K MY A V+ T GV QG LS L++L ++
Sbjct: 622 GYLKTMYASAECLVKINWSLTAPLAFGRGVRQGCPLSGQLYSLAIE 667
>YMH5_CAEEL (P34472) Hypothetical protein F58A4.5 in chromosome III
Length = 1222
Score = 49.3 bits (116), Expect = 1e-05
Identities = 42/156 (26%), Positives = 73/156 (45%), Gaps = 6/156 (3%)
Query: 1 KLWERVIERRLRKEAR--VTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDL 58
++ ER+I R+R E ++ +Q GF+ R ++ + ++K L ++F D
Sbjct: 696 RIMERIICSRIRSEYSHLLSPHQHGFLNFRSCPSSLVRSISLYHSILKNEKSLDILFFDF 755
Query: 59 EKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTED-FPITIGVHQG 117
K +D+V IL K L G+ K+ + SV+ + + +PI+ GV QG
Sbjct: 756 AKAFDKVSHPILLKKLALFGLDKLTCSWFKEFLHLRTFSVKINKFVSSNAYPISSGVPQG 815
Query: 118 STLSPYLFTLVL-DVLTEHIQELAPRCMLFADVVLV 152
S P LF L + D+L + + C FAD + +
Sbjct: 816 SVSGPLLFILFINDLLIDLEPNIHVSC--FADDIKI 849
>YSM6_CAEEL (Q10126) Hypothetical protein F52C9.6 in chromosome III
Length = 279
Score = 41.2 bits (95), Expect = 0.004
Identities = 42/199 (21%), Positives = 77/199 (38%), Gaps = 12/199 (6%)
Query: 148 DVVLVGESREEVNGRLETWRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHII 207
D+VL+ + L+ Q G ++ T+ + N S+ +
Sbjct: 16 DIVLIANHPNTASKMLQELVQKCSEVGLEINTGKTKVLR-NRLADPSKVYFGSPSSTTQL 74
Query: 208 PQVTRFKYLGSFV*NDGEIEADVSHRIQAEWLKW---RRASGVLCDKKVPLKLKGKFYRT 264
V + YLG + + ++ R +A W + + + + DKK+ + L + +
Sbjct: 75 DDVDEYIYLGRQINAHNNLMPEIHRRRRAAWAAFNGSKNTTDSITDKKIRVNL----FDS 130
Query: 265 AVRPALLYGTECWAVKSQHENQVSVAEMRMLRWMSRKTRHDRIKNDTIRERVGVAPIVEK 324
V PAL YG+E W +V + + R + T + + D RE + V V
Sbjct: 131 IVLPALTYGSEAWTFNKALSERVRITHASLERRLVGITLTQQRERDLHREDIRVMSQVRD 190
Query: 325 ----LVENRLR*FGHVERR 339
+ + +L GHV RR
Sbjct: 191 PLNFVKKRKLGWAGHVARR 209
>YO84_CAEEL (P34620) Hypothetical protein ZK1236.4 in chromosome III
Length = 364
Score = 40.0 bits (92), Expect = 0.009
Identities = 29/111 (26%), Positives = 51/111 (45%), Gaps = 6/111 (5%)
Query: 38 RRVIERYRTDKKDLHLVFIDLEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTS 97
+ + E +TD K ++D K +D+V +IL L + IR + S
Sbjct: 10 KNIDEGAQTDVK-----YLDFSKAFDKVSHDILLDKLTSIKINKHLIRWLDVFLTNRSFK 64
Query: 98 VRTQDGTTEDFPITIGVHQGSTLSPYLFTLVLDVLTEHIQELAPRCMLFAD 148
V+ + +E GV QGS +SP LF + ++ ++ ++ + C FAD
Sbjct: 65 VKVGNTLSEPKKTVCGVPQGSVISPVLFGIFVNEISANL-PVGVYCKQFAD 114
>AI2M_YEAST (P03876) Putative COX1/OXI3 intron 2 protein
Length = 789
Score = 34.7 bits (78), Expect = 0.36
Identities = 27/140 (19%), Positives = 56/140 (39%), Gaps = 17/140 (12%)
Query: 4 ERVIERRLRKEARVTDNQ------FGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFID 57
E++++ +R + N GF P + AI + ++ + +D
Sbjct: 300 EKIVQESMRMMLEIIYNNSFSYYSHGFRPNLSCLTAIIQCKNYMQYCNW------FIKVD 353
Query: 58 LEKTYDRVPREILWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQG 117
L K +D +P +L L ++ ++ + + T T+G+ QG
Sbjct: 354 LNKCFDTIPHNMLINVLNERIKDKGFMDLLYKLLRAGYVDKNNNYHNT-----TLGIPQG 408
Query: 118 STLSPYLFTLVLDVLTEHIQ 137
S +SP L + LD L ++++
Sbjct: 409 SVVSPILCNIFLDKLDKYLE 428
>RTP2_TRYBG (P15594) Retrotransposable element SLACS 132 kDa protein
(ORF2)
Length = 1182
Score = 34.3 bits (77), Expect = 0.47
Identities = 23/78 (29%), Positives = 33/78 (41%), Gaps = 2/78 (2%)
Query: 111 TIGVHQGSTLSPYLFTLVLDVLTEHIQELAPRCML--FADVVLVGESREEVNGRLETWRQ 168
T GV QG L P LF++ +Q+ P + D V V EE+ +
Sbjct: 702 TRGVRQGMVLGPLLFSIGTLATLRRLQQTFPEAQFTAYLDDVTVAAPPEELKNVCAATAE 761
Query: 169 ALEAYGFRLSRSTTEYME 186
A+EA G + TE +E
Sbjct: 762 AMEALGIVNNADKTEVLE 779
>POL_BIV27 (P19561) Pol polyprotein [Contains: Protease (EC
3.4.23.-) (P119) (Retropepsin); Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT) (P72); Integrase (IN)]
Length = 1056
Score = 33.5 bits (75), Expect = 0.81
Identities = 21/78 (26%), Positives = 41/78 (51%), Gaps = 3/78 (3%)
Query: 101 QDGTTEDFPITIGVHQGSTLSPYLFTLVLDVLTEHIQELAPRCMLFA--DVVLVGESREE 158
++G E F + + QG SP ++ + E+I++ P ML+ D +L+G +R++
Sbjct: 293 REGPIERFQWNV-LPQGWVCSPAIYQTTTQKIIENIKKSHPDVMLYQYMDDLLIGSNRDD 351
Query: 159 VNGRLETWRQALEAYGFR 176
++ R L +YGF+
Sbjct: 352 HKQIVQEIRDKLGSYGFK 369
>POL_BIV06 (P19560) Pol polyprotein [Contains: Protease (EC
3.4.23.-) (P119) (Retropepsin); Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT) (P72); Integrase (IN)]
Length = 1056
Score = 33.5 bits (75), Expect = 0.81
Identities = 21/78 (26%), Positives = 41/78 (51%), Gaps = 3/78 (3%)
Query: 101 QDGTTEDFPITIGVHQGSTLSPYLFTLVLDVLTEHIQELAPRCMLFA--DVVLVGESREE 158
++G E F + + QG SP ++ + E+I++ P ML+ D +L+G +R++
Sbjct: 293 REGPIERFQWNV-LPQGWVCSPAIYQTTTQKIIENIKKSHPDVMLYQYMDDLLIGSNRDD 351
Query: 159 VNGRLETWRQALEAYGFR 176
++ R L +YGF+
Sbjct: 352 HKQIVQEIRDKLGSYGFK 369
>RUV2_XENLA (Q9DE27) RuvB-like 2 (EC 3.6.1.-) (Reptin)
Length = 462
Score = 33.1 bits (74), Expect = 1.1
Identities = 30/128 (23%), Positives = 58/128 (44%), Gaps = 9/128 (7%)
Query: 119 TLSPYLFTLVLDVLTEHIQELAPRCMLFADVVLVGESREEVN--GRLETWRQALEAYGFR 176
TL + D+ T+ I+ L + DV+ + ++ ++ GR T + +A G
Sbjct: 161 TLKTTEMETIYDLGTKMIESLTKEKVQAGDVITIDKATGKITKLGRAFTRARDYDAMG-- 218
Query: 177 LSRSTTEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFK--YLGSFV*NDGEIEADVSHRI 234
S T++++C + R + V H I + +L F + GEI+++V +I
Sbjct: 219 ---SQTKFVQCPDGELQKRKEVVHTVSLHEIDVINSRTQGFLALFSGDTGEIKSEVREQI 275
Query: 235 QAEWLKWR 242
A+ +WR
Sbjct: 276 NAKVAEWR 283
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.326 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,665,385
Number of Sequences: 164201
Number of extensions: 1545420
Number of successful extensions: 4585
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 9
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 4565
Number of HSP's gapped (non-prelim): 35
length of query: 346
length of database: 59,974,054
effective HSP length: 111
effective length of query: 235
effective length of database: 41,747,743
effective search space: 9810719605
effective search space used: 9810719605
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC144503.9


