

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144503.4 - phase: 0
(366 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
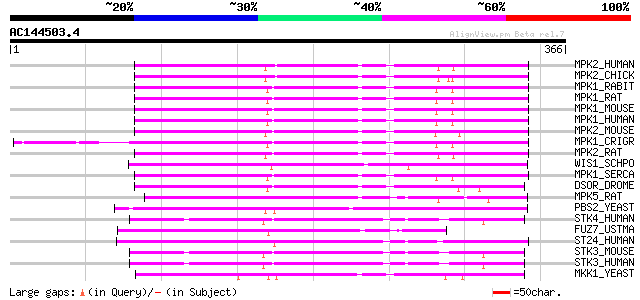
Score E
Sequences producing significant alignments: (bits) Value
MPK2_HUMAN (P36507) Dual specificity mitogen-activated protein k... 169 1e-41
MPK2_CHICK (Q90891) Dual specificity mitogen-activated protein k... 169 1e-41
MPK1_RABIT (P29678) Dual specificity mitogen-activated protein k... 167 5e-41
MPK1_RAT (Q01986) Dual specificity mitogen-activated protein kin... 166 9e-41
MPK1_MOUSE (P31938) Dual specificity mitogen-activated protein k... 166 9e-41
MPK1_HUMAN (Q02750) Dual specificity mitogen-activated protein k... 166 9e-41
MPK2_MOUSE (Q63932) Dual specificity mitogen-activated protein k... 166 1e-40
MPK1_CRIGR (Q63980) Dual specificity mitogen-activated protein k... 166 1e-40
MPK2_RAT (P36506) Dual specificity mitogen-activated protein kin... 165 2e-40
WIS1_SCHPO (P33886) Protein kinase wis1 (EC 2.7.1.-) (Protein ki... 164 4e-40
MPK1_SERCA (Q91447) Dual specificity mitogen-activated protein k... 162 2e-39
DSOR_DROME (Q24324) Dual specificity mitogen-activated protein k... 160 4e-39
MPK5_RAT (Q62862) Dual specificity mitogen-activated protein kin... 155 2e-37
PBS2_YEAST (P08018) Polymyxin B resistance protein kinase (EC 2.... 153 8e-37
STK4_HUMAN (Q13043) Serine/threonine-protein kinase 4 (EC 2.7.1.... 151 2e-36
FUZ7_USTMA (Q99078) Dual specificity protein kinase FUZ7 (EC 2.7... 151 2e-36
ST24_HUMAN (Q9Y6E0) Serine/threonine-protein kinase 24 (EC 2.7.1... 151 3e-36
STK3_MOUSE (Q9JI10) Serine/threonine-protein kinase 3 (EC 2.7.1.... 150 5e-36
STK3_HUMAN (Q13188) Serine/threonine-protein kinase 3 (EC 2.7.1.... 150 6e-36
MKK1_YEAST (P32490) MAP kinase kinase MKK1/SSP32 (EC 2.7.1.-) 149 1e-35
>MPK2_HUMAN (P36507) Dual specificity mitogen-activated protein
kinase kinase 2 (EC 2.7.1.-) (MAP kinase kinase 2)
(MAPKK 2) (ERK activator kinase 2) (MAPK/ERK kinase 2)
(MEK2)
Length = 400
Score = 169 bits (427), Expect = 1e-41
Identities = 107/310 (34%), Positives = 169/310 (54%), Gaps = 58/310 (18%)
Query: 83 ELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNV 142
+ ER++ +G+G+GG V KV HR +G A K+I+ + ++R QI RE+Q+L + + +
Sbjct: 71 DFERISELGAGNGGVVTKVQHRPSGLIMARKLIHLEIKPAIRNQIIRELQVLHECNSPYI 130
Query: 143 VKCHEMYDHNAEIQVLLEYMDGGSL-----EGKHIPQENQLADVARQILRGLAYLHRRH- 196
V + + + EI + +E+MDGGSL E K IP+E L V+ +LRGLAYL +H
Sbjct: 131 VGFYGAFYSDGEISICMEHMDGGSLDQVLKEAKRIPEE-ILGKVSIAVLRGLAYLREKHQ 189
Query: 197 IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQ 256
I+HRD+KPSN+L+NSR ++K+ DFGV L +M NS VGT +YM+PER+ G
Sbjct: 190 IMHRDVKPSNILVNSRGEIKLCDFGVSGQLIDSM--ANSFVGTRSYMAPERL-----QGT 242
Query: 257 YDAYAGDIWSLGVSILEFYMGRFPF----------------AVGRQGDWAS--------- 291
+ + DIWS+G+S++E +GR+P G +G+ S
Sbjct: 243 HYSVQSDIWSMGLSLVELAVGRYPIPPPDAKELEAIFGRPVVDGEEGEPHSISPRPRPPG 302
Query: 292 ------------------LMCAICMSQPPEAPT-TASPEFRDFVSRCLQRDPSRRWTASR 332
L+ I PP+ P +P+F++FV++CL ++P+ R
Sbjct: 303 RPVSGHGMDSRPAMAIFELLDYIVNEPPPKLPNGVFTPDFQEFVNKCLIKNPAERADLKM 362
Query: 333 LLSHPFLVRN 342
L +H F+ R+
Sbjct: 363 LTNHTFIKRS 372
>MPK2_CHICK (Q90891) Dual specificity mitogen-activated protein
kinase kinase 2 (EC 2.7.1.-) (MAP kinase kinase 2)
(MAPKK 2) (ERK activator kinase 2) (MAPK/ERK kinase 2)
(MEK2)
Length = 398
Score = 169 bits (427), Expect = 1e-41
Identities = 107/310 (34%), Positives = 170/310 (54%), Gaps = 58/310 (18%)
Query: 83 ELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNV 142
+ ER++ +G+G+GG V KV H+ +G A K+I+ + ++R QI RE+Q+L + + +
Sbjct: 69 DFERISELGAGNGGVVTKVQHKPSGLIMARKLIHLEIKPAIRNQIIRELQVLHECNSPYI 128
Query: 143 VKCHEMYDHNAEIQVLLEYMDGGSL-----EGKHIPQENQLADVARQILRGLAYLHRRH- 196
V + + + EI + +E+MDGGSL E K IP+E L V+ +LRGLAYL +H
Sbjct: 129 VGFYGAFYSDGEISICMEHMDGGSLDQVLKEAKRIPEE-ILGKVSIAVLRGLAYLREKHQ 187
Query: 197 IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQ 256
I+HRD+KPSN+L+NSR ++K+ DFGV L +M NS VGT +YMSPER+ G
Sbjct: 188 IMHRDVKPSNILVNSRGEIKLCDFGVSGQLIDSM--ANSFVGTRSYMSPERL-----QGT 240
Query: 257 YDAYAGDIWSLGVSILEFYMGRFPF----------------AVGRQGD------WA---- 290
+ + DIWS+G+S++E +GR+P G +G+ WA
Sbjct: 241 HYSVQSDIWSMGLSLVELSIGRYPIPPPDSKELEAIFGRPVVDGAEGESHSVSPWARPPG 300
Query: 291 -----------------SLMCAICMSQPPEAPT-TASPEFRDFVSRCLQRDPSRRWTASR 332
L+ I PP+ P + +F++FV++CL ++P+ R
Sbjct: 301 RPISGHGMDSRPAMAIFELLDYIVNEPPPKLPNGVFTQDFQEFVNKCLIKNPAERADLKM 360
Query: 333 LLSHPFLVRN 342
L++H F+ R+
Sbjct: 361 LMNHTFIKRS 370
>MPK1_RABIT (P29678) Dual specificity mitogen-activated protein
kinase kinase 1 (EC 2.7.1.-) (MAP kinase kinase 1)
(MAPKK 1) (ERK activator kinase 1) (MAPK/ERK kinase 1)
(MEK1)
Length = 392
Score = 167 bits (422), Expect = 5e-41
Identities = 104/306 (33%), Positives = 168/306 (53%), Gaps = 54/306 (17%)
Query: 83 ELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNV 142
+ E+++ +G+G+GG V+KV H+ +G A K+I+ + ++R QI RE+Q+L + + +
Sbjct: 66 DFEKISELGAGNGGVVFKVSHKPSGLVMARKLIHLEIKPAIRNQIIRELQVLHECNSPYI 125
Query: 143 VKCHEMYDHNAEIQVLLEYMDGGSLE-----GKHIPQENQLADVARQILRGLAYLHRRH- 196
V + + + EI + +E+MDGGSL+ IP E L V+ +++GL YL +H
Sbjct: 126 VGFYGAFYSDGEISICMEHMDGGSLDQVLKKAGRIP-EQILGKVSIAVIKGLTYLREKHK 184
Query: 197 IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQ 256
I+HRD+KPSN+L+NSR ++K+ DFGV L +M NS VGT +YMSPER+ G
Sbjct: 185 IMHRDVKPSNILVNSRGEIKLCDFGVSGQLIDSM--ANSFVGTRSYMSPERL-----QGT 237
Query: 257 YDAYAGDIWSLGVSILEFYMGRFP------------FAVGRQGDWA-------------- 290
+ + DIWS+G+S++E +GR+P F +GD A
Sbjct: 238 HYSVQSDIWSMGLSLVEMAVGRYPIPPPDAKELELMFGCQVEGDAAETPPRPRTPGRPLS 297
Query: 291 -------------SLMCAICMSQPPEAPTTA-SPEFRDFVSRCLQRDPSRRWTASRLLSH 336
L+ I PP+ P+ S EF+DFV++CL ++P+ R +L+ H
Sbjct: 298 SYGMDSRPPMAIFELLDYIVNEPPPKLPSAVFSLEFQDFVNKCLIKNPAERADLKQLMVH 357
Query: 337 PFLVRN 342
F+ R+
Sbjct: 358 AFIKRS 363
>MPK1_RAT (Q01986) Dual specificity mitogen-activated protein kinase
kinase 1 (EC 2.7.1.-) (MAP kinase kinase 1) (MAPKK 1)
(ERK activator kinase 1) (MAPK/ERK kinase 1) (MEK1)
Length = 392
Score = 166 bits (420), Expect = 9e-41
Identities = 104/306 (33%), Positives = 168/306 (53%), Gaps = 54/306 (17%)
Query: 83 ELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNV 142
+ E+++ +G+G+GG V+KV H+ +G A K+I+ + ++R QI RE+Q+L + + +
Sbjct: 66 DFEKISELGAGNGGVVFKVSHKPSGLVMARKLIHLEIKPAIRNQIIRELQVLHECNSPYI 125
Query: 143 VKCHEMYDHNAEIQVLLEYMDGGSLE-----GKHIPQENQLADVARQILRGLAYLHRRH- 196
V + + + EI + +E+MDGGSL+ IP E L V+ +++GL YL +H
Sbjct: 126 VGFYGAFYSDGEISICMEHMDGGSLDQVLKKAGRIP-EQILGKVSIAVIKGLTYLREKHK 184
Query: 197 IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQ 256
I+HRD+KPSN+L+NSR ++K+ DFGV L +M NS VGT +YMSPER+ G
Sbjct: 185 IMHRDVKPSNILVNSRGEIKLCDFGVSGQLIDSM--ANSFVGTRSYMSPERL-----QGT 237
Query: 257 YDAYAGDIWSLGVSILEFYMGRFP------------FAVGRQGDWA-------------- 290
+ + DIWS+G+S++E +GR+P F +GD A
Sbjct: 238 HYSVQSDIWSMGLSLVEMAVGRYPIPPPDAKELELLFGCQVEGDAAETPPRPRTPGRPLS 297
Query: 291 -------------SLMCAICMSQPPEAPT-TASPEFRDFVSRCLQRDPSRRWTASRLLSH 336
L+ I PP+ P+ S EF+DFV++CL ++P+ R +L+ H
Sbjct: 298 SYGMDSRPPMAIFELLDYIVNEPPPKLPSGVFSLEFQDFVNKCLIKNPAERADLKQLMVH 357
Query: 337 PFLVRN 342
F+ R+
Sbjct: 358 AFIKRS 363
>MPK1_MOUSE (P31938) Dual specificity mitogen-activated protein
kinase kinase 1 (EC 2.7.1.-) (MAP kinase kinase 1)
(MAPKK 1) (ERK activator kinase 1) (MAPK/ERK kinase 1)
(MEK1)
Length = 392
Score = 166 bits (420), Expect = 9e-41
Identities = 104/306 (33%), Positives = 168/306 (53%), Gaps = 54/306 (17%)
Query: 83 ELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNV 142
+ E+++ +G+G+GG V+KV H+ +G A K+I+ + ++R QI RE+Q+L + + +
Sbjct: 66 DFEKISELGAGNGGVVFKVSHKPSGLVMARKLIHLEIKPAIRNQIIRELQVLHECNSPYI 125
Query: 143 VKCHEMYDHNAEIQVLLEYMDGGSLE-----GKHIPQENQLADVARQILRGLAYLHRRH- 196
V + + + EI + +E+MDGGSL+ IP E L V+ +++GL YL +H
Sbjct: 126 VGFYGAFYSDGEISICMEHMDGGSLDQVLKKAGRIP-EQILGKVSIAVIKGLTYLREKHK 184
Query: 197 IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQ 256
I+HRD+KPSN+L+NSR ++K+ DFGV L +M NS VGT +YMSPER+ G
Sbjct: 185 IMHRDVKPSNILVNSRGEIKLCDFGVSGQLIDSM--ANSFVGTRSYMSPERL-----QGT 237
Query: 257 YDAYAGDIWSLGVSILEFYMGRFP------------FAVGRQGDWA-------------- 290
+ + DIWS+G+S++E +GR+P F +GD A
Sbjct: 238 HYSVQSDIWSMGLSLVEMAVGRYPIPPPDAKELELLFGCHVEGDAAETPPRPRTPGRPLS 297
Query: 291 -------------SLMCAICMSQPPEAPT-TASPEFRDFVSRCLQRDPSRRWTASRLLSH 336
L+ I PP+ P+ S EF+DFV++CL ++P+ R +L+ H
Sbjct: 298 SYGMDSRPPMAIFELLDYIVNEPPPKLPSGVFSLEFQDFVNKCLIKNPAERADLKQLMVH 357
Query: 337 PFLVRN 342
F+ R+
Sbjct: 358 AFIKRS 363
>MPK1_HUMAN (Q02750) Dual specificity mitogen-activated protein
kinase kinase 1 (EC 2.7.1.-) (MAP kinase kinase 1)
(MAPKK 1) (ERK activator kinase 1) (MAPK/ERK kinase 1)
(MEK1)
Length = 392
Score = 166 bits (420), Expect = 9e-41
Identities = 104/306 (33%), Positives = 168/306 (53%), Gaps = 54/306 (17%)
Query: 83 ELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNV 142
+ E+++ +G+G+GG V+KV H+ +G A K+I+ + ++R QI RE+Q+L + + +
Sbjct: 66 DFEKISELGAGNGGVVFKVSHKPSGLVMARKLIHLEIKPAIRNQIIRELQVLHECNSPYI 125
Query: 143 VKCHEMYDHNAEIQVLLEYMDGGSLE-----GKHIPQENQLADVARQILRGLAYLHRRH- 196
V + + + EI + +E+MDGGSL+ IP E L V+ +++GL YL +H
Sbjct: 126 VGFYGAFYSDGEISICMEHMDGGSLDQVLKKAGRIP-EQILGKVSIAVIKGLTYLREKHK 184
Query: 197 IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQ 256
I+HRD+KPSN+L+NSR ++K+ DFGV L +M NS VGT +YMSPER+ G
Sbjct: 185 IMHRDVKPSNILVNSRGEIKLCDFGVSGQLIDSM--ANSFVGTRSYMSPERL-----QGT 237
Query: 257 YDAYAGDIWSLGVSILEFYMGRFP------------FAVGRQGDWA-------------- 290
+ + DIWS+G+S++E +GR+P F +GD A
Sbjct: 238 HYSVQSDIWSMGLSLVEMAVGRYPIPPPDAKELELMFGCQVEGDAAETPPRPRTPGRPLS 297
Query: 291 -------------SLMCAICMSQPPEAPT-TASPEFRDFVSRCLQRDPSRRWTASRLLSH 336
L+ I PP+ P+ S EF+DFV++CL ++P+ R +L+ H
Sbjct: 298 SYGMDSRPPMAIFELLDYIVNEPPPKLPSGVFSLEFQDFVNKCLIKNPAERADLKQLMVH 357
Query: 337 PFLVRN 342
F+ R+
Sbjct: 358 AFIKRS 363
>MPK2_MOUSE (Q63932) Dual specificity mitogen-activated protein
kinase kinase 2 (EC 2.7.1.-) (MAP kinase kinase 2)
(MAPKK 2) (ERK activator kinase 2) (MAPK/ERK kinase 2)
(MEK2)
Length = 401
Score = 166 bits (419), Expect = 1e-40
Identities = 108/311 (34%), Positives = 167/311 (52%), Gaps = 59/311 (18%)
Query: 83 ELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNV 142
+ ER++ +G+G+GG V K HR +G A K+I+ + +VR QI RE+Q+L + + +
Sbjct: 71 DFERISELGAGNGGVVTKARHRPSGFIMARKLIHLEIKPAVRNQIIRELQVLHECNSPYI 130
Query: 143 VKCHEMYDHNAEIQVLLEYMDGGSL-----EGKHIPQENQLADVARQILRGLAYLHRRH- 196
V + + + EI + +E+MDGGSL E K IP E+ L V+ +LRGLAYL +H
Sbjct: 131 VGFYGAFYSDGEISICMEHMDGGSLDQVLKEAKRIP-EDILGKVSIAVLRGLAYLREKHQ 189
Query: 197 IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQ 256
I+HRD+KPSN+L+NSR ++K+ DFGV L +M NS VGT +YMSPER+ G
Sbjct: 190 IMHRDVKPSNILVNSRGEIKLCDFGVSGQLIDSM--ANSFVGTRSYMSPERL-----QGT 242
Query: 257 YDAYAGDIWSLGVSILEFYMGRF------------------------------------- 279
+ + DIWS+G+S++E +GR+
Sbjct: 243 HYSVQSDIWSMGLSLVELAIGRYPIPPPDAKELEASFGRPVVDGADGEPHSVSPRPRPPG 302
Query: 280 -PFAVGRQGDWASLMCA------ICMSQPPEAPT-TASPEFRDFVSRCLQRDPSRRWTAS 331
P +VG D M I PP+ P+ S +F++FV++CL ++P+ R
Sbjct: 303 RPISVGHGMDSRPAMAIFELLDYIVNEPPPKLPSGVFSSDFQEFVNKCLIKNPAERADLK 362
Query: 332 RLLSHPFLVRN 342
L++H F+ R+
Sbjct: 363 LLMNHAFIKRS 373
>MPK1_CRIGR (Q63980) Dual specificity mitogen-activated protein
kinase kinase 1 (EC 2.7.1.-) (MAP kinase kinase 1)
(MAPKK 1) (ERK activator kinase 1) (MAPK/ERK kinase 1)
(MEK1)
Length = 393
Score = 166 bits (419), Expect = 1e-40
Identities = 122/386 (31%), Positives = 192/386 (49%), Gaps = 75/386 (19%)
Query: 3 PIQLPPPTSTGSATASGSPNNNNSRPQRRRRDHLTLPLPQRDTNLAVPLPLPPSGGSGGG 62
PIQL P T GSA S N +++ + L L QR+ L L G
Sbjct: 8 PIQLNP-TPDGSAVNGTSSAETNLEALQKKLEELELEEQQRN-RLEAFLTQKQKVGE--- 62
Query: 63 GNGSGSGSGGASQQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEES 122
+ + E+++ +G+G+GG V+KV H+ +G A K+I+ + +
Sbjct: 63 ----------------LKDDDFEKISELGAGNGGVVFKVSHKPSGLVMARKLIHLEIKPA 106
Query: 123 VRRQIHREIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLE-----GKHIPQENQ 177
+R QI RE+Q+L + + +V + ++ + EI + +E+MDGGSL+ IP E
Sbjct: 107 IRNQIIRELQVLHECNSPYIVGFYGVFYSDGEISICMEHMDGGSLDQVLKKAGRIP-EQI 165
Query: 178 LADVARQILRGLAYLHRRH-IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSS 236
L V+ +++GL YL +H I+HRD+KPSN+L+NSR ++K+ DFGV L +M NS
Sbjct: 166 LGKVSIAVIKGLTYLREKHKIMHRDVKPSNILVNSRGEIKLCDFGVSGQLIDSM--ANSF 223
Query: 237 VGTIAYMSPERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFP------------FAVG 284
VGT +YMSPER+ G + + DIWS+G+S++E +GR+P F
Sbjct: 224 VGTRSYMSPERL-----QGTHYSVQSDIWSMGLSLVEMAVGRYPIPPPDAKELELLFGCQ 278
Query: 285 RQGDWA---------------------------SLMCAICMSQPPEAPT-TASPEFRDFV 316
+GD A L+ I P + P+ S EF+DFV
Sbjct: 279 VEGDAAETPPRPRTPGRPLSSYGMDSRPPMAIFELLDYIVNEPPAKLPSGVFSLEFQDFV 338
Query: 317 SRCLQRDPSRRWTASRLLSHPFLVRN 342
++CL ++P+ R +LL H F+ R+
Sbjct: 339 NKCLIKNPAERADLKQLLVHAFIKRS 364
>MPK2_RAT (P36506) Dual specificity mitogen-activated protein kinase
kinase 2 (EC 2.7.1.-) (MAP kinase kinase 2) (MAPKK 2)
(ERK activator kinase 2) (MAPK/ERK kinase 2) (MEK2)
Length = 400
Score = 165 bits (417), Expect = 2e-40
Identities = 108/310 (34%), Positives = 167/310 (53%), Gaps = 58/310 (18%)
Query: 83 ELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNV 142
+ ER++ +G+G+GG V K HR +G A K+I+ + +VR QI RE+Q+L + + +
Sbjct: 71 DFERISELGAGNGGVVTKARHRPSGLIMARKLIHLEIKPAVRNQIIRELQVLHECNSPYI 130
Query: 143 VKCHEMYDHNAEIQVLLEYMDGGSL-----EGKHIPQENQLADVARQILRGLAYLHRRH- 196
V + + + EI + +E+MDGGSL E K IP E+ L V+ +LRGLAYL +H
Sbjct: 131 VGFYGAFYSDGEISICMEHMDGGSLDQVLKEAKRIP-EDILGKVSIAVLRGLAYLREKHQ 189
Query: 197 IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQ 256
I+HRD+KPSN+L+NSR ++K+ DFGV L +M NS VGT +YMSPER+ G
Sbjct: 190 IMHRDVKPSNILVNSRGEIKLCDFGVSGQLIDSM--ANSFVGTRSYMSPERL-----QGT 242
Query: 257 YDAYAGDIWSLGVSILEFYMGRFPF----------------AVGRQGDWAS--------- 291
+ + DIWS+G+S++E +GR+P G G+ S
Sbjct: 243 HYSVQSDIWSMGLSLVELAIGRYPIPPPDAKELEASFGRPVVDGADGEPHSVSPRPRPPG 302
Query: 292 ------------------LMCAICMSQPPEAPT-TASPEFRDFVSRCLQRDPSRRWTASR 332
L+ I PP+ P+ S +F++FV++CL ++P+ R
Sbjct: 303 RPISGHGMDSRPAMAIFELLDYIVNEPPPKLPSGVFSSDFQEFVNKCLIKNPAERADLKL 362
Query: 333 LLSHPFLVRN 342
L +H F+ R+
Sbjct: 363 LTNHAFIKRS 372
>WIS1_SCHPO (P33886) Protein kinase wis1 (EC 2.7.1.-) (Protein
kinase sty2)
Length = 605
Score = 164 bits (414), Expect = 4e-40
Identities = 98/269 (36%), Positives = 146/269 (53%), Gaps = 8/269 (2%)
Query: 79 IPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVD 138
I SE+ +L +G G+ G VYK +H+ G ALK I EE+ QI E+ IL
Sbjct: 315 INMSEIIKLEELGKGNYGVVYKALHQPTGVTMALKEIRLSLEEATFNQIIMELDILHKAV 374
Query: 139 DVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGKH---IPQENQLADVARQILRGLAYLHRR 195
+V + + + + +EYMD GS++ + I E LA A +++GL L
Sbjct: 375 SPYIVDFYGAFFVEGSVFICMEYMDAGSMDKLYAGGIKDEGVLARTAYAVVQGLKTLKEE 434
Query: 196 H-IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDIND 254
H I+HRD+KP+N+L+NS QVK+ DFGV L ++ N +G +YM+PERI
Sbjct: 435 HNIIHRDVKPTNVLVNSNGQVKLCDFGVSGNLVASISKTN--IGCQSYMAPERIRVGGPT 492
Query: 255 GQYDAYA--GDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPEF 312
Y D+WSLG++ILE +G +P+ + + AIC PP P + SPE
Sbjct: 493 NGVLTYTVQADVWSLGLTILEMALGAYPYPPESYTSIFAQLSAICDGDPPSLPDSFSPEA 552
Query: 313 RDFVSRCLQRDPSRRWTASRLLSHPFLVR 341
RDFV++CL ++PS R L +HP+L++
Sbjct: 553 RDFVNKCLNKNPSLRPDYHELANHPWLLK 581
>MPK1_SERCA (Q91447) Dual specificity mitogen-activated protein
kinase kinase 1 (EC 2.7.1.-) (MAP kinase kinase 1)
(MAPKK 1) (ERK activator kinase 1) (MAPK/ERK kinase 1)
(MEK1) (Fragment)
Length = 388
Score = 162 bits (409), Expect = 2e-39
Identities = 102/308 (33%), Positives = 165/308 (53%), Gaps = 56/308 (18%)
Query: 83 ELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNV 142
+ E+++ +G+G+GG V+KV H+ +G A K+I+ + ++R QI RE+Q+L + + +
Sbjct: 60 DFEKISELGAGNGGVVFKVSHKPSGLIMARKLIHLEIKPAIRNQIIRELQVLHECNSPYI 119
Query: 143 VKCHEMYDHNAEIQVLLEYMDGGSLE-----GKHIPQENQLADVARQILRGLAYLHRRH- 196
V + + + EI + +E+MDGGSL+ IP E L V+ +++GL YL +H
Sbjct: 120 VGFYGAFYSDGEISICMEHMDGGSLDQVLKKAGRIP-EQILGKVSIAVIKGLTYLREKHK 178
Query: 197 IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQ 256
I+HRD+KPSN+L+NSR ++K+ DFGV L +M NS VGT +YMSPER+ G
Sbjct: 179 IMHRDVKPSNILVNSRGEIKLCDFGVSGQLIDSM--ANSFVGTRSYMSPERL-----QGT 231
Query: 257 YDAYAGDIWSLGVSILEFYMGRFP------------FAVGRQGDWA-------------- 290
+ + DIWS+G+S++E +GR+P F +GD
Sbjct: 232 HYSVQSDIWSMGLSLVEMAIGRYPIPPPDSKELELMFGCPVEGDSPVTETSPRQRAPGRP 291
Query: 291 ---------------SLMCAICMSQPPEAPT-TASPEFRDFVSRCLQRDPSRRWTASRLL 334
L+ I PP+ P EF+DFV++CL ++P+ R +L+
Sbjct: 292 MSSYGSDSRPPMAIFELLDYIVNEPPPKLPNGVFGSEFQDFVNKCLIKNPAERADLKQLM 351
Query: 335 SHPFLVRN 342
H F+ R+
Sbjct: 352 IHAFIKRS 359
>DSOR_DROME (Q24324) Dual specificity mitogen-activated protein
kinase kinase dSOR1 (EC 2.7.1.-) (Downstream of RAF)
(MAPKK)
Length = 393
Score = 160 bits (406), Expect = 4e-39
Identities = 102/287 (35%), Positives = 154/287 (53%), Gaps = 38/287 (13%)
Query: 83 ELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNV 142
+LE+L +GSG+GG V KV H A K+I+ + ++++QI RE+++L + + ++
Sbjct: 83 DLEKLGELGSGNGGVVMKVRHTHTHLIMARKLIHLEVKPAIKKQILRELKVLHECNFPHI 142
Query: 143 VKCHEMYDHNAEIQVLLEYMDGGSLE-----GKHIPQENQLADVARQILRGLAYLHRRH- 196
V + + + EI + +EYMDGGSL+ IP E+ L + +L+GL+YL H
Sbjct: 143 VGFYGAFYSDGEISICMEYMDGGSLDLILKRAGRIP-ESILGRITLAVLKGLSYLRDNHA 201
Query: 197 IVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQ 256
I+HRD+KPSN+L+NS ++KI DFGV L +M NS VGT +YMSPER+ G
Sbjct: 202 IIHRDVKPSNILVNSSGEIKICDFGVSGQLIDSM--ANSFVGTRSYMSPERL-----QGT 254
Query: 257 YDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMC--AICMSQPPEAPTTA------ 308
+ + DIWSLG+S++E +G +P S+ A QP + P
Sbjct: 255 HYSVQSDIWSLGLSLVEMAIGMYPIPPPNTATLESIFADNAEESGQPTDEPRAMAIFELL 314
Query: 309 ----------------SPEFRDFVSRCLQRDPSRRWTASRLLSHPFL 339
S EF+DFV CL++ P R LLSHP++
Sbjct: 315 DYIVNEPPPKLEHKIFSTEFKDFVDICLKKQPDERADLKTLLSHPWI 361
>MPK5_RAT (Q62862) Dual specificity mitogen-activated protein kinase
kinase 5 (EC 2.7.1.-) (MAP kinase kinase 5) (MAPKK 5)
(MAPK/ERK kinase 5)
Length = 448
Score = 155 bits (392), Expect = 2e-37
Identities = 92/258 (35%), Positives = 146/258 (55%), Gaps = 14/258 (5%)
Query: 90 IGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDDVNVVKCHEMY 149
+G G+GGTVYK H +G+ A+KVI +++QI E++IL D ++ + +
Sbjct: 172 LGHGNGGTVYKAYHVPSGKILAVKVILLDITLELQKQIMSELEILYKCDSSYIIGFYGAF 231
Query: 150 DHNAEIQVLLEYMDGGSLEGKHIPQENQLADVARQILRGLAYLHRRHIVHRDIKPSNLLI 209
I + E+MDGGSL+ E+ L +A +++GL YL I+HRD+KPSN+L+
Sbjct: 232 FVENRISICTEFMDGGSLDVYRKIPEHVLGRIAVAVVKGLTYLWSLKILHRDVKPSNMLV 291
Query: 210 NSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDINDGQYDAYAGDIWSLGV 269
N+ QVK+ DFGV L ++ + VGT AYM+PERI+ + QY ++ D+WSLG+
Sbjct: 292 NTSGQVKLCDFGVSTQLVNSI--AKTYVGTNAYMAPERISGE----QYGIHS-DVWSLGI 344
Query: 270 SILEFYMGRFPF--AVGRQGDWASLMCAICMSQPPEAPTTASPEFRD----FVSRCLQRD 323
S +E +GRFP+ QG L C+ ++P EF + F+++C+++
Sbjct: 345 SFMELALGRFPYPQIQKNQGSLMPLQLLQCIVD-EDSPVLPLGEFSEPFVHFITQCMRKQ 403
Query: 324 PSRRWTASRLLSHPFLVR 341
P R L+ HPF+V+
Sbjct: 404 PKERPAPEELMGHPFIVQ 421
>PBS2_YEAST (P08018) Polymyxin B resistance protein kinase (EC
2.7.1.-)
Length = 668
Score = 153 bits (386), Expect = 8e-37
Identities = 101/282 (35%), Positives = 154/282 (53%), Gaps = 14/282 (4%)
Query: 70 SGGASQQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHR 129
S G+S ++ + ELE L+ +G G+ G V KV+H+ A K + +E+ RQI
Sbjct: 348 SNGSSSRITL--DELEFLDELGHGNYGNVSKVLHKPTNVIMATKEVRLELDEAKFRQILM 405
Query: 130 EIQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSL-----EGKHIP--QENQLADVA 182
E+++L + +V + + + + +EYMDGGSL E I E QLA +A
Sbjct: 406 ELEVLHKCNSPYIVDFYGAFFIEGAVYMCMEYMDGGSLDKIYDESSEIGGIDEPQLAFIA 465
Query: 183 RQILRGLAYLHRRH-IVHRDIKPSNLLINSRK-QVKIADFGVGRILNQTMDPCNSSVGTI 240
++ GL L +H I+HRD+KP+N+L ++ + VK+ DFGV N +++G
Sbjct: 466 NAVIHGLKELKEQHNIIHRDVKPTNILCSANQGTVKLCDFGVSG--NLVASLAKTNIGCQ 523
Query: 241 AYMSPERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQ 300
+YM+PERI + D DIWSLG+SILE +GR+P+ + S + AI
Sbjct: 524 SYMAPERIKSLNPDRATYTVQSDIWSLGLSILEMALGRYPYPPETYDNIFSQLSAIVDGP 583
Query: 301 PPEAPTTA-SPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVR 341
PP P+ S + +DFVS CLQ+ P RR T + L HP+LV+
Sbjct: 584 PPRLPSDKFSSDAQDFVSLCLQKIPERRPTYAALTEHPWLVK 625
>STK4_HUMAN (Q13043) Serine/threonine-protein kinase 4 (EC 2.7.1.37)
(STE20-like kinase MST1) (MST-1) (Mammalian STE20-like
protein kinase 1) (Serine/threonine-protein kinase
Krs-2)
Length = 487
Score = 151 bits (382), Expect = 2e-36
Identities = 98/271 (36%), Positives = 148/271 (54%), Gaps = 26/271 (9%)
Query: 80 PFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDD 139
P + L ++G GS G+VYK +H+ G+ A+K + ES ++I +EI I++ D
Sbjct: 26 PEEVFDVLEKLGEGSYGSVYKAIHKETGQIVAIKQV---PVESDLQEIIKEISIMQQCDS 82
Query: 140 VNVVKCHEMYDHNAEIQVLLEYMDGGS------LEGKHIPQENQLADVARQILRGLAYLH 193
+VVK + Y N ++ +++EY GS L K + E+++A + + L+GL YLH
Sbjct: 83 PHVVKYYGSYFKNTDLWIVMEYCGAGSVSDIIRLRNKTLT-EDEIATILQSTLKGLEYLH 141
Query: 194 RRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDIN 253
+HRDIK N+L+N+ K+ADFGV L TM N+ +GT +M+PE I
Sbjct: 142 FMRKIHRDIKAGNILLNTEGHAKLADFGVAGQLTDTMAKRNTVIGTPFWMAPE----VIQ 197
Query: 254 DGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPE-- 311
+ Y+ A DIWSLG++ +E G+ P+A M AI M PT PE
Sbjct: 198 EIGYNCVA-DIWSLGITAIEMAEGKPPYAD------IHPMRAIFMIPTNPPPTFRKPELW 250
Query: 312 ---FRDFVSRCLQRDPSRRWTASRLLSHPFL 339
F DFV +CL + P +R TA++LL HPF+
Sbjct: 251 SDNFTDFVKQCLVKSPEQRATATQLLQHPFV 281
>FUZ7_USTMA (Q99078) Dual specificity protein kinase FUZ7 (EC
2.7.1.-)
Length = 435
Score = 151 bits (382), Expect = 2e-36
Identities = 91/223 (40%), Positives = 135/223 (59%), Gaps = 14/223 (6%)
Query: 72 GASQQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREI 131
G +L + +L+ L+ +G+G+GGTV KV+H +G A KV++ + SVR+QI RE+
Sbjct: 97 GVEYKLDLKNEDLKTLSELGAGNGGTVTKVLHEKSGTVMAKKVVFIDAKPSVRKQILREL 156
Query: 132 QILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEG---KHIPQENQL-ADVARQILR 187
QIL + + +V + Y + I + +E+M SL+G K+ P ++ +A +
Sbjct: 157 QILHECNSPYIVSFYGAYLNEPHICMCMEFMQKDSLDGIYKKYGPISPEICGKIAVAVSH 216
Query: 188 GLAYLHRRH-IVHRDIKPSNLLINSRKQVKIADFGV-GRILNQTMDPCNSSVGTIAYMSP 245
GL YL+ H I+HRD+KPSN+L+N Q+KI DFGV G ++N D + VGT YMSP
Sbjct: 217 GLTYLYDVHRIIHRDVKPSNILVNGAGQIKICDFGVSGELINSIAD---TFVGTSTYMSP 273
Query: 246 ERINTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGD 288
ERI D QY + D+WSLGVSI++ +GRFPFA + D
Sbjct: 274 ERIQGD----QY-SVKSDVWSLGVSIIDVALGRFPFAENEEDD 311
>ST24_HUMAN (Q9Y6E0) Serine/threonine-protein kinase 24 (EC
2.7.1.37) (STE20-like kinase MST3) (MST-3) (Mammalian
STE20-like protein kinase 3)
Length = 443
Score = 151 bits (381), Expect = 3e-36
Identities = 95/275 (34%), Positives = 143/275 (51%), Gaps = 11/275 (4%)
Query: 71 GGASQQLVIPFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHRE 130
GG++ P +L +IG GS G V+K + + A+K+I E I +E
Sbjct: 23 GGSTNLKADPEELFTKLEKIGKGSFGEVFKGIDNRTQKVVAIKIIDLEEAEDEIEDIQQE 82
Query: 131 IQILRDVDDVNVVKCHEMYDHNAEIQVLLEYMDGGSLEGKHIP---QENQLADVARQILR 187
I +L D V K + Y + ++ +++EY+ GGS P E Q+A + R+IL+
Sbjct: 83 ITVLSQCDSPYVTKYYGSYLKDTKLWIIMEYLGGGSALDLLEPGPLDETQIATILREILK 142
Query: 188 GLAYLHRRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPER 247
GL YLH +HRDIK +N+L++ +VK+ADFGV L T N+ VGT +M+PE
Sbjct: 143 GLDYLHSEKKIHRDIKAANVLLSEHGEVKLADFGVAGQLTDTQIKRNTFVGTPFWMAPE- 201
Query: 248 INTDINDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTT 307
I YD+ A DIWSLG++ +E G P + + ++ I + PP
Sbjct: 202 ---VIKQSAYDSKA-DIWSLGITAIELARGEPPHS---ELHPMKVLFLIPKNNPPTLEGN 254
Query: 308 ASPEFRDFVSRCLQRDPSRRWTASRLLSHPFLVRN 342
S ++FV CL ++PS R TA LL H F++RN
Sbjct: 255 YSKPLKEFVEACLNKEPSFRPTAKELLKHKFILRN 289
>STK3_MOUSE (Q9JI10) Serine/threonine-protein kinase 3 (EC 2.7.1.37)
(STE20-like kinase MST2) (MST-2) (Mammalian STE20-like
protein kinase 2)
Length = 497
Score = 150 bits (379), Expect = 5e-36
Identities = 97/271 (35%), Positives = 148/271 (53%), Gaps = 26/271 (9%)
Query: 80 PFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDD 139
P + L ++G GS G+V+K +H+ +G+ A+K + ES ++I +EI I++ D
Sbjct: 23 PEEVFDVLEKLGEGSYGSVFKAIHKESGQVVAIKQV---PVESDLQEIIKEISIMQQCDS 79
Query: 140 VNVVKCHEMYDHNAEIQVLLEYMDGGS------LEGKHIPQENQLADVARQILRGLAYLH 193
VVK + Y N ++ +++EY GS L K + E+++A + + L+GL YLH
Sbjct: 80 PYVVKYYGSYFKNTDLWIVMEYCGAGSVSDIIRLRNKTLT-EDEIATILKSTLKGLEYLH 138
Query: 194 RRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDIN 253
+HRDIK N+L+N+ K+ADFGV L TM N+ +GT +M+PE I
Sbjct: 139 FMRKIHRDIKAGNILLNTEGHAKLADFGVAGQLTDTMAKRNTVIGTPFWMAPE----VIQ 194
Query: 254 DGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPE-- 311
+ Y+ A DIWSLG++ +E G+ P+A M AI M PT PE
Sbjct: 195 EIGYNCVA-DIWSLGITSIEMAEGKPPYAD------IHPMRAIFMIPTNPPPTFRKPELW 247
Query: 312 ---FRDFVSRCLQRDPSRRWTASRLLSHPFL 339
F DFV +CL + P +R TA++LL HPF+
Sbjct: 248 SDDFTDFVKKCLVKSPEQRATATQLLQHPFI 278
>STK3_HUMAN (Q13188) Serine/threonine-protein kinase 3 (EC 2.7.1.37)
(STE20-like kinase MST2) (MST-2) (Mammalian STE20-like
protein kinase 2) (Serine/threonine-protein kinase
Krs-1)
Length = 491
Score = 150 bits (378), Expect = 6e-36
Identities = 97/271 (35%), Positives = 149/271 (54%), Gaps = 26/271 (9%)
Query: 80 PFSELERLNRIGSGSGGTVYKVVHRINGRAYALKVIYGHHEESVRRQIHREIQILRDVDD 139
P + L ++G GS G+V+K +H+ +G+ A+K + ES ++I +EI I++ D
Sbjct: 23 PEEVFDVLEKLGEGSYGSVFKAIHKESGQVVAIKQV---PVESDLQEIIKEISIMQQCDS 79
Query: 140 VNVVKCHEMYDHNAEIQVLLEYMDGGS------LEGKHIPQENQLADVARQILRGLAYLH 193
VVK + Y N ++ +++EY GS L K + E+++A + + L+GL YLH
Sbjct: 80 PYVVKYYGSYFKNTDLWIVMEYCGAGSVSDIIRLRNKTLI-EDEIATILKSTLKGLEYLH 138
Query: 194 RRHIVHRDIKPSNLLINSRKQVKIADFGVGRILNQTMDPCNSSVGTIAYMSPERINTDIN 253
+HRDIK N+L+N+ K+ADFGV L TM N+ +GT +M+PE I
Sbjct: 139 FMRKIHRDIKAGNILLNTEGHAKLADFGVAGQLTDTMAKRNTVIGTPFWMAPE----VIQ 194
Query: 254 DGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQGDWASLMCAICMSQPPEAPTTASPE-- 311
+ Y+ A DIWSLG++ +E G+ P+A M AI M PT PE
Sbjct: 195 EIGYNCVA-DIWSLGITSIEMAEGKPPYAD------IHPMRAIFMIPTNPPPTFRKPELW 247
Query: 312 ---FRDFVSRCLQRDPSRRWTASRLLSHPFL 339
F DFV +CL ++P +R TA++LL HPF+
Sbjct: 248 SDDFTDFVKKCLVKNPEQRATATQLLQHPFI 278
>MKK1_YEAST (P32490) MAP kinase kinase MKK1/SSP32 (EC 2.7.1.-)
Length = 508
Score = 149 bits (376), Expect = 1e-35
Identities = 94/276 (34%), Positives = 151/276 (54%), Gaps = 28/276 (10%)
Query: 84 LERLNRIGSGSGGTVYKVVHRINGRAYALKVIYG-HHEESVRRQIHREIQILRDVDDVNV 142
+E L +G G+GG+V K + + +ALKVI + + ++QI RE+Q R +
Sbjct: 221 IETLGILGEGAGGSVSKCKLKNGSKIFALKVINTLNTDPEYQKQIFRELQFNRSFQSEYI 280
Query: 143 VKCHEMY--DHNAEIQVLLEYMDGGSLEG--KHIPQ------ENQLADVARQILRGLAYL 192
V+ + M+ D N+ I + +EYM G SL+ K++ + E L +A +LRGL+YL
Sbjct: 281 VRYYGMFTDDENSSIYIAMEYMGGRSLDAIYKNLLERGGRISEKVLGKIAEAVLRGLSYL 340
Query: 193 HRRHIVHRDIKPSNLLINSRKQVKIADFGV-GRILNQTMDPCNSSVGTIAYMSPERINTD 251
H + ++HRDIKP N+L+N QVK+ DFGV G +N + GT YM+PERI
Sbjct: 341 HEKKVIHRDIKPQNILLNENGQVKLCDFGVSGEAVNSL---ATTFTGTSFYMAPERI--- 394
Query: 252 INDGQYDAYAGDIWSLGVSILEFYMGRFPFAVGRQ----GDWASLMCAIC----MSQPPE 303
GQ + D+WSLG++ILE G+FP + + + LM + + PE
Sbjct: 395 --QGQPYSVTSDVWSLGLTILEVANGKFPCSSEKMAANIAPFELLMWILTFTPELKDEPE 452
Query: 304 APTTASPEFRDFVSRCLQRDPSRRWTASRLLSHPFL 339
+ SP F+ F+ CL++D R + ++++HP++
Sbjct: 453 SNIIWSPSFKSFIDYCLKKDSRERPSPRQMINHPWI 488
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.136 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,764,288
Number of Sequences: 164201
Number of extensions: 2339221
Number of successful extensions: 30419
Number of sequences better than 10.0: 1954
Number of HSP's better than 10.0 without gapping: 1715
Number of HSP's successfully gapped in prelim test: 246
Number of HSP's that attempted gapping in prelim test: 19749
Number of HSP's gapped (non-prelim): 5306
length of query: 366
length of database: 59,974,054
effective HSP length: 112
effective length of query: 254
effective length of database: 41,583,542
effective search space: 10562219668
effective search space used: 10562219668
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC144503.4


