

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144484.11 - phase: 0
(796 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
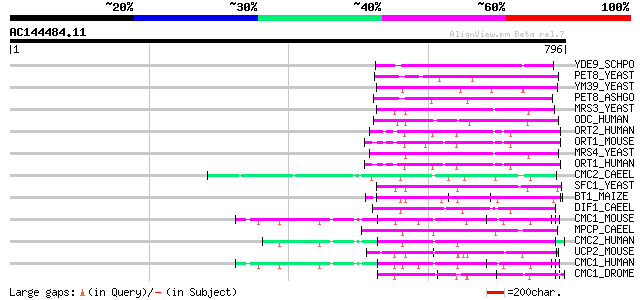
Score E
Sequences producing significant alignments: (bits) Value
YDE9_SCHPO (Q10442) Putative mitochondrial carrier C12B10.09 114 1e-24
PET8_YEAST (P38921) Putative mitochondrial carrier protein PET8 114 1e-24
YM39_YEAST (Q03829) Putative mitochondrial carrier YMR166C 110 1e-23
PET8_ASHGO (O60029) Putative mitochondrial carrier protein PET8 108 4e-23
MRS3_YEAST (P10566) Mitochondrial RNA splicing protein MRS3 96 3e-19
ODC_HUMAN (Q9BQT8) Mitochondrial 2-oxodicarboxylate carrier 94 1e-18
ORT2_HUMAN (Q9BXI2) Mitochondrial ornithine transporter 2 (Solut... 91 2e-17
ORT1_MOUSE (Q9WVD5) Mitochondrial ornithine transporter 1 (Solut... 90 3e-17
MRS4_YEAST (P23500) Mitochondrial RNA splicing protein MRS4 89 6e-17
ORT1_HUMAN (Q9Y619) Mitochondrial ornithine transporter 1 (Solut... 88 8e-17
CMC2_CAEEL (Q20799) Probable calcium-binding mitochondrial carri... 88 8e-17
SFC1_YEAST (P33303) Succinate/fumarate mitochondrial transporter... 86 3e-16
BT1_MAIZE (P29518) Brittle-1 protein, chloroplast precursor 86 3e-16
DIF1_CAEEL (Q27257) Protein dif-1 85 9e-16
CMC1_MOUSE (Q8BH59) Calcium-binding mitochondrial carrier protei... 85 9e-16
MPCP_CAEEL (P40614) Phosphate carrier protein, mitochondrial pre... 83 3e-15
CMC2_HUMAN (Q9UJS0) Calcium-binding mitochondrial carrier protei... 83 3e-15
UCP2_MOUSE (P70406) Mitochondrial uncoupling protein 2 (UCP 2) (... 82 4e-15
CMC1_HUMAN (O75746) Calcium-binding mitochondrial carrier protei... 82 4e-15
CMC1_DROME (Q9VA73) Calcium-binding mitochondrial carrier Aralar1 82 6e-15
>YDE9_SCHPO (Q10442) Putative mitochondrial carrier C12B10.09
Length = 345
Score = 114 bits (284), Expect = 1e-24
Identities = 84/259 (32%), Positives = 128/259 (48%), Gaps = 12/259 (4%)
Query: 525 ALAGGLSCALSCAL-LHPVDSIKTRVQASSMSFPEIIAKLPEIGTRGLYRGSIPAILGQF 583
AL G+ L+ L L P+D++KTR+QA + G G+YRG ++G
Sbjct: 85 ALGAGICAGLAVDLSLFPIDTLKTRLQAKG-------GFVKNGGFHGVYRGLGSILVGSA 137
Query: 584 SSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQAGLF 643
L +E K L L + Q+ ++ VR+P EV+KQR QA
Sbjct: 138 PGASLFFTTYENMKSRLSQSGLGLSDPQIHMCSASLGEIAACIVRVPTEVIKQRAQASGG 197
Query: 644 NNVGEALVGTWQQDG--LKGFFRGTGATLCREVPFYVAGMGLYAESK-KGVQKLLGRELE 700
++ T + + F+ G G T+ RE+PF + ++ K K K +
Sbjct: 198 TLSSRNILQTILKSNNVWRDFYAGYGITIAREIPFTLIQFPIWEHLKLKWRIKHSRNKNL 257
Query: 701 AWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAV 760
A E G+++GG+AA +TTPFDV+KTR+MT+Q R +S SI+ HEG L L+KG V
Sbjct: 258 AHEAAISGSIAGGIAAALTTPFDVVKTRIMTSQQR-LSYVFTIKSIVAHEGFLALYKGIV 316
Query: 761 PRFFWIAPLGAMNFAGYEL 779
PR W++ GA+ Y++
Sbjct: 317 PRVLWLSGGGAIFLGCYDV 335
Score = 32.7 bits (73), Expect = 3.9
Identities = 16/60 (26%), Positives = 30/60 (49%), Gaps = 3/60 (5%)
Query: 524 SALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGTRG---LYRGSIPAIL 580
+A++G ++ ++ AL P D +KTR+ S + + G LY+G +P +L
Sbjct: 261 AAISGSIAGGIAAALTTPFDVVKTRIMTSQQRLSYVFTIKSIVAHEGFLALYKGIVPRVL 320
>PET8_YEAST (P38921) Putative mitochondrial carrier protein PET8
Length = 284
Score = 114 bits (284), Expect = 1e-24
Identities = 84/280 (30%), Positives = 128/280 (45%), Gaps = 27/280 (9%)
Query: 524 SALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGTRGLYRGSIPAILGQF 583
S L+G + + + P+D+IKTR+QA F G +G+YRG A++
Sbjct: 7 SLLSGAAAGTSTDLVFFPIDTIKTRLQAKGGFFANG-------GYKGIYRGLGSAVVASA 59
Query: 584 SSHGLRTGIFEASKLVLVNVAPNLPELQVQS-----------IASFCSTFLGTAVRIPCE 632
L F + + V P + +L Q ++S VR+P E
Sbjct: 60 PGASL---FFISYDYMKVKSRPYISKLYSQGSEQLIDTTTHMLSSSIGEICACLVRVPAE 116
Query: 633 VLKQRLQAGLFNNVGEALVGTWQQDGLKGF----FRGTGATLCREVPFYVAGMGLYAESK 688
V+KQR Q N+ + L + D +G +RG T+ RE+PF LY K
Sbjct: 117 VVKQRTQVHSTNSSWQTLQSILRNDNKEGLRKNLYRGWSTTIMREIPFTCIQFPLYEYLK 176
Query: 689 KGVQKLLGR-ELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSIL 747
K K G+ ++E W+ G+++GG+AA TTP D +KTR+M + + S+ V I
Sbjct: 177 KTWAKANGQSQVEPWKGAICGSIAGGIAAATTTPLDFLKTRLMLNK-TTASLGSVIIRIY 235
Query: 748 RHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKN 787
R EGP F G PR WI+ GA+ YE ++K+
Sbjct: 236 REEGPAVFFSGVGPRTMWISAGGAIFLGMYETVHSLLSKS 275
>YM39_YEAST (Q03829) Putative mitochondrial carrier YMR166C
Length = 368
Score = 110 bits (275), Expect = 1e-23
Identities = 88/306 (28%), Positives = 145/306 (46%), Gaps = 47/306 (15%)
Query: 526 LAGGLSCALSCALLHPVDSIKTRVQASS--MSFPEIIAK-----LPEIGTRGLYRGSIPA 578
++GG+ + + +H +D++KTR Q + + +I+ L E RGLY G + A
Sbjct: 58 VSGGIGGKIGDSAMHSLDTVKTRQQGAPNVKKYRNMISAYRTIWLEEGVRRGLYGGYMAA 117
Query: 579 ILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRL 638
+LG F S + G +E +K ++ + + A F F+ + V +P EVLK RL
Sbjct: 118 MLGSFPSAAIFFGTYEYTKRTMIEDW-QINDTITHLSAGFLGDFISSFVYVPSEVLKTRL 176
Query: 639 QA-GLFNN-----------VGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAE 686
Q G FNN + A+ +++G + F G ATL R++PF Y +
Sbjct: 177 QLQGRFNNPFFQSGYNYSNLRNAIKTVIKEEGFRSLFFGYKATLARDLPFSALQFAFYEK 236
Query: 687 SKK---GVQKLLGR--ELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQ-------- 733
++ +++ GR EL I GA +GGLA ++TTP DV+KTR+ T Q
Sbjct: 237 FRQLAFKIEQKDGRDGELSIPNEILTGACAGGLAGIITTPMDVVKTRVQTQQPPSQSNKS 296
Query: 734 ----------GR----SVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYEL 779
GR S S+S+ ++ + EG LG F G PRF W + ++ Y++
Sbjct: 297 YSVTHPHVTNGRPAALSNSISLSLRTVYQSEGVLGFFSGVGPRFVWTSVQSSIMLLLYQM 356
Query: 780 ARKAMN 785
+ ++
Sbjct: 357 TLRGLS 362
>PET8_ASHGO (O60029) Putative mitochondrial carrier protein PET8
Length = 271
Score = 108 bits (271), Expect = 4e-23
Identities = 84/268 (31%), Positives = 127/268 (47%), Gaps = 19/268 (7%)
Query: 522 LRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGTRGLYRGSIPAILG 581
L S ++G + + + P+D++KTR+QA F G RG+YRG A++
Sbjct: 6 LASLVSGAAAGTSTDVVFFPIDTLKTRLQAKGGFFHNG-------GYRGIYRGLGSAVVA 58
Query: 582 QFSSHGLRTGIFEASKLVLVNV------APNLPELQVQSIASFCSTFLGTAVRIPCEVLK 635
L +++ K L V + L E+ ++S VR+P EV+K
Sbjct: 59 SAPGASLFFVTYDSMKQQLRPVMGRWTASEQLAEVLTHMLSSSLGEMSACLVRVPAEVIK 118
Query: 636 QRLQAGLFNNVGEALVGTWQ----QDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGV 691
QR Q N+ + L + + ++G +RG T+ RE+PF LY KK
Sbjct: 119 QRTQTHHTNSSLQTLRLILRDPTGEGVVRGLYRGWWTTIMREIPFTCIQFPLYEYLKKKW 178
Query: 692 QKLLGRE-LEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHE 750
E + AW+ G+L+GG+AA TTP DV+KTRMM + R V M +A ++ R E
Sbjct: 179 AAYAEIERVSAWQGAVCGSLAGGIAAAATTPLDVLKTRMMLHE-RRVPMLHLARTLFREE 237
Query: 751 GPLGLFKGAVPRFFWIAPLGAMNFAGYE 778
G F+G PR WI+ GA+ YE
Sbjct: 238 GARVFFRGIGPRTMWISAGGAIFLGVYE 265
Score = 32.3 bits (72), Expect = 5.1
Identities = 23/84 (27%), Positives = 40/84 (47%), Gaps = 3/84 (3%)
Query: 515 EIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEI-IAK--LPEIGTRGL 571
EI S + A+ G L+ ++ A P+D +KTR+ P + +A+ E G R
Sbjct: 183 EIERVSAWQGAVCGSLAGGIAAAATTPLDVLKTRMMLHERRVPMLHLARTLFREEGARVF 242
Query: 572 YRGSIPAILGQFSSHGLRTGIFEA 595
+RG P + + + G++EA
Sbjct: 243 FRGIGPRTMWISAGGAIFLGVYEA 266
>MRS3_YEAST (P10566) Mitochondrial RNA splicing protein MRS3
Length = 314
Score = 96.3 bits (238), Expect = 3e-19
Identities = 71/273 (26%), Positives = 128/273 (46%), Gaps = 19/273 (6%)
Query: 526 LAGGLSCALSCALLHPVDSIKTRVQ---ASSMSFPEIIAKLPEI----GTRGLYRGSIPA 578
+AG + + +++ P+D++KTR+Q A S+S +++++ I GT L++G
Sbjct: 38 IAGAFAGIMEHSVMFPIDALKTRIQSANAKSLSAKNMLSQISHISTSEGTLALWKGVQSV 97
Query: 579 ILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQS-IASFCSTFLGTAVRIPCEVLKQR 637
ILG +H + G +E K L++ + ++ I+ C+T A+ P + +KQR
Sbjct: 98 ILGAGPAHAVYFGTYEFCKKNLIDSSDTQTHHPFKTAISGACATTASDALMNPFDTIKQR 157
Query: 638 LQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGR 697
+Q +V + +Q +GL F+ TL +PF +Y S K +
Sbjct: 158 IQLNTSASVWQTTKQIYQSEGLAAFYYSYPTTLVMNIPFAAFNFVIYESSTKFLNP--SN 215
Query: 698 ELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIV---------AFSILR 748
E G++SG A +TTP D +KT + ++VS+ I+ A +I +
Sbjct: 216 EYNPLIHCLCGSISGSTCAAITTPLDCIKTVLQIRGSQTVSLEIMRKADTFSKAASAIYQ 275
Query: 749 HEGPLGLFKGAVPRFFWIAPLGAMNFAGYELAR 781
G G ++G PR P A+++ YE A+
Sbjct: 276 VYGWKGFWRGWKPRIVANMPATAISWTAYECAK 308
Score = 42.0 bits (97), Expect = 0.006
Identities = 27/94 (28%), Positives = 43/94 (45%), Gaps = 3/94 (3%)
Query: 702 WETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVS---MSIVAFSILRHEGPLGLFKG 758
+ + GA +G + V P D +KTR+ +A +S+S M I EG L L+KG
Sbjct: 34 YHQLIAGAFAGIMEHSVMFPIDALKTRIQSANAKSLSAKNMLSQISHISTSEGTLALWKG 93
Query: 759 AVPRFFWIAPLGAMNFAGYELARKAMNKNDEAKT 792
P A+ F YE +K + + + +T
Sbjct: 94 VQSVILGAGPAHAVYFGTYEFCKKNLIDSSDTQT 127
>ODC_HUMAN (Q9BQT8) Mitochondrial 2-oxodicarboxylate carrier
Length = 299
Score = 94.0 bits (232), Expect = 1e-18
Identities = 85/289 (29%), Positives = 128/289 (43%), Gaps = 29/289 (10%)
Query: 523 RSALAGGLSCALSCALLHPVDSIKTRVQASSM-----SFPEIIAKLPEI----GTRGLYR 573
R +AGG + + L+HP+D +KTR Q S+ ++ I G G Y+
Sbjct: 15 RQIVAGGSAGLVEICLMHPLDVVKTRFQIQRCATDPNSYKSLVDSFRMIFQMEGLFGFYK 74
Query: 574 GSIPAILGQFSSHGLRTGIFEASKLVL--VNVAPNLPELQVQSIASFCSTFLGTAVRIPC 631
G +P IL + ++ FE K +L V+++P L +IA S V P
Sbjct: 75 GILPPILAETPKRAVKFFTFEQYKKLLGYVSLSPAL----TFAIAGLGSGLTEAIVVNPF 130
Query: 632 EVLKQRLQAGLFNNVGE--ALVGTWQQD------GLKGFFRGTGATLCREVPFYVAGMGL 683
EV+K LQA N E + VG +Q GL+G +G ATL R F + G
Sbjct: 131 EVVKVGLQANR-NTFAEQPSTVGYARQIIKKEGWGLQGLNKGLTATLGRHGVFNMVYFGF 189
Query: 684 YAESKKGVQKLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSI-- 741
Y K + LE W +G LSG +A+V+ PFDV K+R+ Q +
Sbjct: 190 YYNVKNMIPVNKDPILEFWRKFGIGLLSGTIASVINIPFDVAKSRIQGPQPVPGEIKYRT 249
Query: 742 ---VAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKN 787
++ + EG L L+KG +P+ + P GA+ YE + +N
Sbjct: 250 CFKTMATVYQEEGILALYKGLLPKIMRLGPGGAVMLLVYEYTYSWLQEN 298
Score = 33.1 bits (74), Expect = 3.0
Identities = 25/90 (27%), Positives = 38/90 (41%), Gaps = 5/90 (5%)
Query: 700 EAWETIAVGALSGGLAAVVTTPFDVMKTRMM----TAQGRSVSMSIVAF-SILRHEGPLG 754
EA I G +G + + P DV+KTR S + +F I + EG G
Sbjct: 12 EASRQIVAGGSAGLVEICLMHPLDVVKTRFQIQRCATDPNSYKSLVDSFRMIFQMEGLFG 71
Query: 755 LFKGAVPRFFWIAPLGAMNFAGYELARKAM 784
+KG +P P A+ F +E +K +
Sbjct: 72 FYKGILPPILAETPKRAVKFFTFEQYKKLL 101
>ORT2_HUMAN (Q9BXI2) Mitochondrial ornithine transporter 2 (Solute
carrier family 25, member 2)
Length = 301
Score = 90.5 bits (223), Expect = 2e-17
Identities = 83/305 (27%), Positives = 135/305 (44%), Gaps = 41/305 (13%)
Query: 516 IPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPE--------IG 567
I A L + AGG +C L+ P D+IK ++Q +FP++ L + +G
Sbjct: 7 IQAAIDLTAGAAGGTACVLTG---QPFDTIKVKMQ----TFPDLYKGLTDCFLKTYAQVG 59
Query: 568 TRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVA-----PNLPELQVQSIASFCSTF 622
RG Y+G+ PA++ + + + + + + VA L +LQ + SF S F
Sbjct: 60 LRGFYKGTGPALMAYVAENSVLFMCYGFCQQFVRKVAGMDKQAKLSDLQTAAAGSFASAF 119
Query: 623 LGTAVRIPCEVLKQRLQ-----------AGLFNNVGEALVGTWQQDGLKGFFRGTGATLC 671
A+ P E++K RLQ A N + + G ++DG GF+ G +TL
Sbjct: 120 AALAL-CPTELVKCRLQTMYEMEMSGKIAKSHNTIWSVVKGILKKDGPLGFYHGLSSTLL 178
Query: 672 REVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGLAAV----VTTPFDVMKT 727
+E P Y G Y S+ GR + + + LSGG+A + + P D +K+
Sbjct: 179 QEGPGYFFFFGGYELSRSFFAS--GRSKDELGPVHL-MLSGGVAGICLWLIVFPVDCIKS 235
Query: 728 RM--MTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMN 785
R+ ++ G+ S++R+EG + L+ G P F YE +RK M
Sbjct: 236 RIQVLSMYGKQAGFIGTLLSVVRNEGIVALYSGLKATMIRAIPANGALFVAYEYSRKMMM 295
Query: 786 KNDEA 790
K EA
Sbjct: 296 KQLEA 300
>ORT1_MOUSE (Q9WVD5) Mitochondrial ornithine transporter 1 (Solute
carrier family 25, member 15)
Length = 301
Score = 89.7 bits (221), Expect = 3e-17
Identities = 83/312 (26%), Positives = 138/312 (43%), Gaps = 47/312 (15%)
Query: 509 AVPPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPE--- 565
A+ ++++ AG+ AGG +C L+ P D++K ++Q +FP++ L +
Sbjct: 6 AIQAAIDLTAGA------AGGTACVLTG---QPFDTMKVKMQ----TFPDLYRGLTDCCL 52
Query: 566 -----IGTRGLYRGSIPAILGQFSSHGLRTGIFE-----ASKLVLVNVAPNLPELQVQSI 615
+G RG Y+G+ PA++ + + + + K+V ++ L +LQ +
Sbjct: 53 KTYSQVGFRGFYKGTSPALIANIAENSVLFMCYGFCQQVVRKVVGLDQQAKLSDLQNAAA 112
Query: 616 ASFCSTFLGTAVRIPCEVLKQRLQ-----------AGLFNNVGEALVGTWQQDGLKGFFR 664
SF S F V P E++K RLQ A N V + +++DG GF+
Sbjct: 113 GSFASAF-AALVLCPTELVKCRLQTMYEMETSGKIAASQNTVWSVVKEIFRKDGPLGFYH 171
Query: 665 GTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGLAAV----VTT 720
G +TL REVP Y G Y S+ GR + + + LSGG +
Sbjct: 172 GLSSTLLREVPGYFFFFGGYELSRSFFAS--GRSKDELGPVPL-MLSGGFGGICLWLAVY 228
Query: 721 PFDVMKTR--MMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYE 778
P D +K+R +++ G+ + SI+++EG L+ G P P F YE
Sbjct: 229 PVDCIKSRIQVLSMTGKQTGLVRTFLSIVKNEGITALYSGLKPTMIRAFPANGALFLAYE 288
Query: 779 LARKAMNKNDEA 790
+RK M EA
Sbjct: 289 YSRKLMMNQLEA 300
>MRS4_YEAST (P23500) Mitochondrial RNA splicing protein MRS4
Length = 304
Score = 88.6 bits (218), Expect = 6e-17
Identities = 72/290 (24%), Positives = 126/290 (42%), Gaps = 20/290 (6%)
Query: 516 IPAGSVLRSAL-AGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEI-------G 567
+P+ + L S L AG + + +L+ P+D++KTRVQA+ ++ + +I G
Sbjct: 17 LPSHAPLHSQLLAGAFAGIMEHSLMFPIDALKTRVQAAGLNKAASTGMISQISKISTMEG 76
Query: 568 TRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQS-IASFCSTFLGTA 626
+ L++G ILG +H + G +E K L++ +++ ++ +T A
Sbjct: 77 SMALWKGVQSVILGAGPAHAVYFGTYEFCKARLISPEDMQTHQPMKTALSGTIATIAADA 136
Query: 627 VRIPCEVLKQRLQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAE 686
+ P + +KQRLQ V +Q +G F+ TL +PF +Y
Sbjct: 137 LMNPFDTVKQRLQLDTNLRVWNVTKQIYQNEGFAAFYYSYPTTLAMNIPFAAFNFMIYES 196
Query: 687 SKKGVQKLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIV---- 742
+ K G +SG A +TTP D +KT + +VS+ I+
Sbjct: 197 ASKFFNP--QNSYNPLIHCLCGGISGATCAALTTPLDCIKTVLQVRGSETVSIEIMKDAN 254
Query: 743 -----AFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKN 787
+ +IL G G ++G PR P A+++ YE A+ + KN
Sbjct: 255 TFGRASRAILEVHGWKGFWRGLKPRIVANIPATAISWTAYECAKHFLMKN 304
Score = 34.3 bits (77), Expect = 1.3
Identities = 23/91 (25%), Positives = 38/91 (41%), Gaps = 3/91 (3%)
Query: 705 IAVGALSGGLAAVVTTPFDVMKTRMMTA---QGRSVSMSIVAFSILRHEGPLGLFKGAVP 761
+ GA +G + + P D +KTR+ A + S M I EG + L+KG
Sbjct: 27 LLAGAFAGIMEHSLMFPIDALKTRVQAAGLNKAASTGMISQISKISTMEGSMALWKGVQS 86
Query: 762 RFFWIAPLGAMNFAGYELARKAMNKNDEAKT 792
P A+ F YE + + ++ +T
Sbjct: 87 VILGAGPAHAVYFGTYEFCKARLISPEDMQT 117
>ORT1_HUMAN (Q9Y619) Mitochondrial ornithine transporter 1 (Solute
carrier family 25, member 15) (SP1855)
Length = 301
Score = 88.2 bits (217), Expect = 8e-17
Identities = 82/312 (26%), Positives = 137/312 (43%), Gaps = 47/312 (15%)
Query: 509 AVPPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPE--- 565
A+ ++++ AG+ AGG +C L+ P D++K ++Q +FP++ L +
Sbjct: 6 AIQAAIDLTAGA------AGGTACVLTG---QPFDTMKVKMQ----TFPDLYRGLTDCCL 52
Query: 566 -----IGTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVA-----PNLPELQVQSI 615
+G RG Y+G+ PA++ + + + + + V+ VA L +LQ +
Sbjct: 53 KTYSQVGFRGFYKGTSPALIANIAENSVLFMCYGFCQQVVRKVAGLDKQAKLSDLQNAAA 112
Query: 616 ASFCSTFLGTAVRIPCEVLKQRLQ-----------AGLFNNVGEALVGTWQQDGLKGFFR 664
SF S F V P E++K RLQ A N V + ++DG GF+
Sbjct: 113 GSFASAF-AALVLCPTELVKCRLQTMYEMETSGKIAKSQNTVWSVIKSILRKDGPLGFYH 171
Query: 665 GTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGLAAV----VTT 720
G +TL REVP Y G Y S+ GR + + + LSGG+ +
Sbjct: 172 GLSSTLLREVPGYFFFFGGYELSRSFFAS--GRSKDELGPVPL-MLSGGVGGICLWLAVY 228
Query: 721 PFDVMKTR--MMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYE 778
P D +K+R +++ G+ +++++EG L+ G P P F YE
Sbjct: 229 PVDCIKSRIQVLSMSGKQAGFIRTFINVVKNEGITALYSGLKPTMIRAFPANGALFLAYE 288
Query: 779 LARKAMNKNDEA 790
+RK M EA
Sbjct: 289 YSRKLMMNQLEA 300
>CMC2_CAEEL (Q20799) Probable calcium-binding mitochondrial carrier
F55A11.4
Length = 588
Score = 88.2 bits (217), Expect = 8e-17
Identities = 107/541 (19%), Positives = 216/541 (39%), Gaps = 55/541 (10%)
Query: 284 DETKEESVGISAQKVASNIFSIPLTNVERLKTTLSTVSLTELIEMLPQLGKTTKDHPDKK 343
+E E + G + QK + L+++ K ++ + + + ++ LGK TK+
Sbjct: 5 NEQTESTSGAAEQKEDDEEQYVQLSSLGEYKDEVTPLLSPKHVPLV--LGKVTKEAAIAT 62
Query: 344 KLFSVQDFFRYTESEGRRFFEELDRDGDGQVTLEDLEIAMRRRK--LPRRYAKEFMSRTR 401
E + R ++ LD D DG + + DL +A++ +P A MS+
Sbjct: 63 HSALHGGMSEEKERQIRDIYDRLDIDNDGTIDIRDLTLALKHETPHIPANLAPVIMSKMS 122
Query: 402 SHLFSRSFGWKQFLSFMEQKEPTILRAYTSLCLTKSGTLKKSEILESLKNSGLPANEDNA 461
R + F S++ + E + + + G + E+ K+ G+P ++ A
Sbjct: 123 PDDEGR-VDFYSFSSYVLENEQKLAEMFADMDRNHDGLVDVVEMKNYCKDIGVPLDDHKA 181
Query: 462 AAMMRFLNADTEESISYGHFRNFMLLLPSDRLQEDPRSIWFEAATVVAVPPSVEIP---- 517
++ ++ S+ F+ FM+L PS L +D W ++ + +IP
Sbjct: 182 QHIVNKMDQTGSASVDLKEFQEFMMLYPSSDL-KDIVDFW-RHNLIIDIGEDSQIPEDFS 239
Query: 518 -----AGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEI----IAKL--PEI 566
G R +AGG + A+S P D IK +Q +S + KL E
Sbjct: 240 QQEMQEGIWWRHLVAGGAAGAVSRTCTAPFDRIKVYLQVNSSKTNRLGVMSCLKLLHAEG 299
Query: 567 GTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTA 626
G + +RG+ ++ ++ ++ K ++ N ++ + C+ A
Sbjct: 300 GIKSFWRGNGINVIKIAPESAIKFMCYDQLKRLIQKKKGN---EEISTFERLCAGSAAGA 356
Query: 627 VR----IPCEVLKQRLQAGLFNNVGEALV----GTWQQDGLKGFFRGTGATLCREVPFYV 678
+ P EV+K RL + ++ + ++G++ F++G L +P+
Sbjct: 357 ISQSTIYPMEVMKTRLALRKTGQLDRGIIHFAHKMYTKEGIRCFYKGYLPNLIGIIPYAG 416
Query: 679 AGMGLYAESKKGVQKLL---GRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGR 735
+ +Y K+ + E +A G S + + PF +++TR+
Sbjct: 417 IDLAIYETLKRTYVRYYETNSSEPGVLALLACGTCSSTCGQLSSYPFALVRTRLQ----- 471
Query: 736 SVSMSIVAFS------------ILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKA 783
++SI +S IL++EG G ++G P F + P ++++ YE R
Sbjct: 472 --ALSITRYSPQPDTMFGQFKYILQNEGVTGFYRGITPNFLKVIPAVSISYVVYEKVRTG 529
Query: 784 M 784
+
Sbjct: 530 L 530
>SFC1_YEAST (P33303) Succinate/fumarate mitochondrial transporter
(Regulator of acetyl-CoA synthetase activity)
Length = 322
Score = 86.3 bits (212), Expect = 3e-16
Identities = 77/305 (25%), Positives = 127/305 (41%), Gaps = 41/305 (13%)
Query: 526 LAGGLSCALSCALLHPVDSIKTRVQA-------SSMSFPEIIAKLPEI----GTRGLYRG 574
+AGG + HP+D+IK R+Q + P I I G LY+G
Sbjct: 15 MAGGTAGLFEALCCHPLDTIKVRMQIYRRVAGIEHVKPPGFIKTGRTIYQKEGFLALYKG 74
Query: 575 SIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRI-PCEV 633
++G +R +E + +LVN + +A + + + P EV
Sbjct: 75 LGAVVIGIIPKMAIRFSSYEFYRTLLVNKESGIVSTGNTFVAGVGAGITEAVLVVNPMEV 134
Query: 634 LKQRLQAG-----------LFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMG 682
+K RLQA +NN A +++G+ +RG T R+ A
Sbjct: 135 VKIRLQAQHLTPSEPNAGPKYNNAIHAAYTIVKEEGVSALYRGVSLTAARQATNQGANFT 194
Query: 683 LYAESKKGVQKLLGRE-LEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMS- 740
+Y++ K+ +Q + L +WET +G +SG + P D +KTR+ + +S+S+
Sbjct: 195 VYSKLKEFLQNYHQMDVLPSWETSCIGLISGAIGPFSNAPLDTIKTRLQ--KDKSISLEK 252
Query: 741 --------IVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAM------NK 786
+ +L+ EG L+KG PR +AP A+ F YE R+ + K
Sbjct: 253 QSGMKKIITIGAQLLKEEGFRALYKGITPRVMRVAPGQAVTFTVYEYVREHLENLGIFKK 312
Query: 787 NDEAK 791
ND K
Sbjct: 313 NDTPK 317
>BT1_MAIZE (P29518) Brittle-1 protein, chloroplast precursor
Length = 436
Score = 86.3 bits (212), Expect = 3e-16
Identities = 66/281 (23%), Positives = 130/281 (45%), Gaps = 17/281 (6%)
Query: 526 LAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEI----GTRGLYRGSIPAILG 581
++G ++ A+S + P+++I+T + S+ + I G GL+RG+ +L
Sbjct: 139 VSGAIAGAVSRTFVAPLETIRTHLMVGSIGVDSMAGVFQWIMQNEGWTGLFRGNAVNVLR 198
Query: 582 QFSSHGLRTGIFEASKLVLVNVAPNLPELQVQS--IASFCSTFLGTAVRIPCEVLKQR-- 637
S + ++ +K L P++ + + +A + F T P E++K R
Sbjct: 199 VAPSKAIEHFTYDTAKKFLTPKGDEPPKIPIPTPLVAGALAGFASTLCTYPMELIKTRVT 258
Query: 638 LQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGR 697
++ +++NV A V + +G +RG +L VP+ Y K+ ++ GR
Sbjct: 259 IEKDVYDNVAHAFVKILRDEGPSELYRGLTPSLIGVVPYAACNFYAYETLKRLYRRATGR 318
Query: 698 ----ELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQ--GRSVSMSIV--AFSILRH 749
++ T+ +G+ +G +A+ T P +V + +M GR V +++ + IL+
Sbjct: 319 RPGADVGPVATLLIGSAAGAIASSATFPLEVARKQMQVGAVGGRQVYQNVLHAIYCILKK 378
Query: 750 EGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAM-NKNDE 789
EG GL++G P + P + F YE +K + +K DE
Sbjct: 379 EGAGGLYRGLGPSCIKLMPAAGIAFMCYEACKKILVDKEDE 419
Score = 61.2 bits (147), Expect = 1e-08
Identities = 47/172 (27%), Positives = 72/172 (41%), Gaps = 8/172 (4%)
Query: 630 PCEVLKQRLQAGLFNNVGEALVGTW--QQDGLKGFFRGTGATLCREVPFYVAGMGLYAES 687
P E ++ L G A V W Q +G G FRG + R P Y +
Sbjct: 154 PLETIRTHLMVGSIGVDSMAGVFQWIMQNEGWTGLFRGNAVNVLRVAPSKAIEHFTYDTA 213
Query: 688 KKGVQKLLGR--ELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFS 745
KK + ++ + GAL+G + + T P +++KTR+ + +++
Sbjct: 214 KKFLTPKGDEPPKIPIPTPLVAGALAGFASTLCTYPMELIKTRVTIEKDVYDNVAHAFVK 273
Query: 746 ILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYE----LARKAMNKNDEAKTG 793
ILR EGP L++G P + P A NF YE L R+A + A G
Sbjct: 274 ILRDEGPSELYRGLTPSLIGVVPYAACNFYAYETLKRLYRRATGRRPGADVG 325
Score = 51.6 bits (122), Expect = 8e-06
Identities = 45/193 (23%), Positives = 79/193 (40%), Gaps = 19/193 (9%)
Query: 511 PPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEI----IAKLPEI 566
PP + IP V AG L+ S +P++ IKTRV + + + L +
Sbjct: 224 PPKIPIPTPLV-----AGALAGFASTLCTYPMELIKTRVTIEKDVYDNVAHAFVKILRDE 278
Query: 567 GTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASF----CSTF 622
G LYRG P+++G +E K + P V +A+ +
Sbjct: 279 GPSELYRGLTPSLIGVVPYAACNFYAYETLKRLYRRATGRRPGADVGPVATLLIGSAAGA 338
Query: 623 LGTAVRIPCEVLKQRLQAG------LFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPF 676
+ ++ P EV ++++Q G ++ NV A+ +++G G +RG G + + +P
Sbjct: 339 IASSATFPLEVARKQMQVGAVGGRQVYQNVLHAIYCILKKEGAGGLYRGLGPSCIKLMPA 398
Query: 677 YVAGMGLYAESKK 689
Y KK
Sbjct: 399 AGIAFMCYEACKK 411
>DIF1_CAEEL (Q27257) Protein dif-1
Length = 312
Score = 84.7 bits (208), Expect = 9e-16
Identities = 83/292 (28%), Positives = 127/292 (43%), Gaps = 35/292 (11%)
Query: 521 VLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPE-----------IIAKLPEIGTR 569
VL + +AGG+ + + + HP D++K R+Q M P + + + G
Sbjct: 4 VLLNFIAGGVGGSCTVIVGHPFDTVKVRIQTMPMPKPGEKPQFTGALDCVKRTVSKEGFF 63
Query: 570 GLYRGSIPAILGQFSSHGLRTGIFEASK-LVLVNVAPNLPELQVQSIASFCSTFLGTAVR 628
LY+G ++G + G K L + + + +Q + + F T V
Sbjct: 64 ALYKGMAAPLVGVSPLFAVFFGGCAVGKWLQQTDPSQEMTFIQNANAGALAGVFT-TIVM 122
Query: 629 IPCEVLKQRLQAGLFNNVG---------EALVGTWQQDGLKGFFRGTGATLCREVPFYVA 679
+P E +K LQ + G + + ++Q G+ +RGTGATL R++P A
Sbjct: 123 VPGERIKCLLQVQQAGSAGSGVHYDGPLDVVKKLYKQGGISSIYRGTGATLLRDIPASAA 182
Query: 680 GMGLYAESKKGVQKLLGRELEAWETIAVGA--LSGGLAAV----VTTPFDVMKTRMMTAQ 733
+ +Y KK K G A T++ GA ++GGLA + V P DV+K+R+ TA
Sbjct: 183 YLSVYEYLKK---KFSGE--GAQRTLSPGATLMAGGLAGIANWGVCIPADVLKSRLQTAP 237
Query: 734 GRSVSMSI--VAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKA 783
I V +LR EGP LFKG P P A F G EL A
Sbjct: 238 EGKYPDGIRGVLREVLREEGPRALFKGFWPVMLRAFPANAACFFGLELTLAA 289
>CMC1_MOUSE (Q8BH59) Calcium-binding mitochondrial carrier protein
Aralar1 (Mitochondrial aspartate glutamate carrier 1)
(Solute carrier family 25, member 12)
Length = 677
Score = 84.7 bits (208), Expect = 9e-16
Identities = 75/273 (27%), Positives = 119/273 (43%), Gaps = 20/273 (7%)
Query: 528 GGLSCALSCALLHPVDSIKTRVQAS------------SMSFPEIIAKLPEIGTRGLYRGS 575
G ++ A+ ++P+D +KTR+Q SF L G GLYRG
Sbjct: 333 GSVAGAVGATAVYPIDLVKTRMQNQRGTGSVVGELMYKNSFDCFKKVLRYEGFFGLYRGL 392
Query: 576 IPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLK 635
IP ++G ++ + + + ++P L + +A C+ P E++K
Sbjct: 393 IPQLIGVAPEKAIKLTVNDFVRDKFTKRDGSIP-LPAEILAGGCAGGSQVIFTNPLEIVK 451
Query: 636 QRLQAGLFNNVGEAL--VGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQK 693
RLQ G + + Q GL G ++G A R++PF +YA K +
Sbjct: 452 IRLQVAGEITTGPRVSALNVLQDLGLFGLYKGAKACFLRDIPFSAIYFPVYAHCKLLLAD 511
Query: 694 LLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTA--QGRSVSMSIVAF--SILRH 749
GR + + GAL+G AA + TP DV+KTR+ A G++ +V ILR
Sbjct: 512 ENGR-VGGINLLTAGALAGVPAASLVTPADVIKTRLQVAARAGQTTYSGVVDCFRKILRE 570
Query: 750 EGPLGLFKGAVPRFFWIAPLGAMNFAGYELARK 782
EGP +KG R F +P + YEL ++
Sbjct: 571 EGPSAFWKGTAARVFRSSPQFGVTLVTYELLQR 603
Score = 64.7 bits (156), Expect = 9e-10
Identities = 102/482 (21%), Positives = 196/482 (40%), Gaps = 62/482 (12%)
Query: 324 ELIEMLPQLGKTTKDHPDKKKLFSVQDFFRYT------ESEGRRFFEELDRDGDGQVTLE 377
+++++L + TKD L S Q+F + +S F+ D+ G+G+VT E
Sbjct: 55 KIVQLLAGVADQTKDG-----LISYQEFLAFESVLCAPDSMFIVAFQLFDKSGNGEVTFE 109
Query: 378 DLEIAMRR----RKLPRRYAKEFMSRTRSHLFSRSFGWKQFLSFMEQKEPTILR-AYTSL 432
+++ + +P + EF+ H + + +F F+++ + R A+
Sbjct: 110 NVKEIFGQTIIHHHIPFNWDCEFIRLHFGHNRKKHLNYVEFTQFLQELQLEHARQAFALK 169
Query: 433 CLTKSGTLKK---SEILESLKNSGL-PANEDNAAAMMRFLNADTEESISYGHFRNFMLLL 488
+KSG + S+++ ++++ L P E+N ++ T +S+ +F F LL
Sbjct: 170 DKSKSGMISGLDFSDVMVTIRSHMLTPFVEEN---LVSAAGGGTSHQVSFSYFNAFNSLL 226
Query: 489 PSDRLQEDPRSIWFEAATVVAVPPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTR 548
+ L R I+ +T+ +E+ +SA+ G L +L+ + +
Sbjct: 227 NNMELV---RKIY---STLAGTRKDIEVTKEEFAQSAIRYGQVTPLEIDILYQLADLYNA 280
Query: 549 VQASSMSFPEIIAKLPEIGTRGLYRGSIPAILGQFS---SHGLRTGIFEASKLVLVNVAP 605
+++ E IA L E G++P L + S GL I+ + +A
Sbjct: 281 SGRLTLADIERIAPLAE--------GALPYNLAELQRQQSPGLGRPIW-------LQIAE 325
Query: 606 NLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQ--AGLFNNVGEALVGT--------WQ 655
+ + S+A +G P +++K R+Q G + VGE + +
Sbjct: 326 SAYRFTLGSVAGA----VGATAVYPIDLVKTRMQNQRGTGSVVGELMYKNSFDCFKKVLR 381
Query: 656 QDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGLA 715
+G G +RG L P + + + K G + I G +GG
Sbjct: 382 YEGFFGLYRGLIPQLIGVAPEKAIKLTVNDFVRDKFTKRDG-SIPLPAEILAGGCAGGSQ 440
Query: 716 AVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFA 775
+ T P +++K R+ A + + A ++L+ G GL+KGA F P A+ F
Sbjct: 441 VIFTNPLEIVKIRLQVAGEITTGPRVSALNVLQDLGLFGLYKGAKACFLRDIPFSAIYFP 500
Query: 776 GY 777
Y
Sbjct: 501 VY 502
Score = 48.1 bits (113), Expect = 9e-05
Identities = 32/119 (26%), Positives = 55/119 (45%), Gaps = 17/119 (14%)
Query: 685 AESKKGVQKLLGRELEAWETIA-------VGALSGGLAAVVTTPFDVMKTRMMTAQGRSV 737
AE ++ LGR + W IA +G+++G + A P D++KTRM +G
Sbjct: 305 AELQRQQSPGLGRPI--WLQIAESAYRFTLGSVAGAVGATAVYPIDLVKTRMQNQRGTGS 362
Query: 738 SMSIVAFS--------ILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKND 788
+ + + +LR+EG GL++G +P+ +AP A+ + R K D
Sbjct: 363 VVGELMYKNSFDCFKKVLRYEGFFGLYRGLIPQLIGVAPEKAIKLTVNDFVRDKFTKRD 421
>MPCP_CAEEL (P40614) Phosphate carrier protein, mitochondrial
precursor (PTP)
Length = 340
Score = 82.8 bits (203), Expect = 3e-15
Identities = 73/301 (24%), Positives = 133/301 (43%), Gaps = 27/301 (8%)
Query: 505 ATVVAVPPSVEIPAGSVLR-SALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIA-- 561
A+ V+ P VE +G AL G LSC ++ + P+D +K R+Q + + I
Sbjct: 26 ASAVSAPGQVEFGSGKYYAYCALGGVLSCGITHTAIVPLDLVKCRIQVNPEKYTGIATGF 85
Query: 562 --KLPEIGTRGLYRGSIPAILGQFSSHGL-RTGIFEASKLVLVNVAPN----LPELQVQS 614
+ E G R L +G P +LG +S+ GL + G +E K V ++ L +
Sbjct: 86 RTTIAEEGARALVKGWAPTLLG-YSAQGLGKFGFYEIFKNVYADMLGEENAYLYRTSLYL 144
Query: 615 IASFCSTFLGTAVRIPCEVLKQRLQA--GLFNNVGEALVGTWQQDGLKGFFRGTGATLCR 672
AS + F + P E K R+Q G + ++ +GL GF++G R
Sbjct: 145 AASASAEFFADILLAPMEATKVRIQTSPGAPPTLRGCAPMIYKAEGLTGFYKGLPPLWMR 204
Query: 673 EVPFYVAGMGLYAESKKGVQKLL--------GRELEAWETIAVGALSGGLAAVVTTPFDV 724
++P+ + + ++ + + + + + + T G ++G A+V+ P D
Sbjct: 205 QIPYTMMKFACFEKTVEALYQYVVPKPRAECSKAEQLVVTFVAGYIAGVFCAIVSHPADT 264
Query: 725 MKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAM 784
+ +++ + A IL+ G G++KG VPR I L A+ + Y+ + A+
Sbjct: 265 VVSKL------NQDSQATAGGILKKLGFAGVWKGLVPRIIMIGTLTALQWFIYDSVKVAL 318
Query: 785 N 785
N
Sbjct: 319 N 319
>CMC2_HUMAN (Q9UJS0) Calcium-binding mitochondrial carrier protein
Aralar2 (Mitochondrial aspartate glutamate carrier 2)
(Solute carrier family 25, member 13) (Citrin)
Length = 675
Score = 82.8 bits (203), Expect = 3e-15
Identities = 75/273 (27%), Positives = 126/273 (45%), Gaps = 20/273 (7%)
Query: 528 GGLSCALSCALLHPVDSIKTRVQ--ASSMSFP-EIIAK---------LPEIGTRGLYRGS 575
G ++ A+ ++P+D +KTR+Q S+ SF E++ K L G GLYRG
Sbjct: 335 GSVAGAVGATAVYPIDLVKTRMQNQRSTGSFVGELMYKNSFDCFKKVLRYEGFFGLYRGL 394
Query: 576 IPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLK 635
+P +LG ++ + + + ++ ++P L + +A C+ P E++K
Sbjct: 395 LPQLLGVAPEKAIKLTVNDFVRDKFMHKDGSVP-LAAEILAGGCAGGSQVIFTNPLEIVK 453
Query: 636 QRLQAGLFNNVGEAL--VGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQK 693
RLQ G + + + G G ++G A R++PF YA K
Sbjct: 454 IRLQVAGEITTGPRVSALSVVRDLGFFGIYKGAKACFLRDIPFSAIYFPCYAHVKASFAN 513
Query: 694 LLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQ--GRSVSMSIVAF--SILRH 749
G ++ + GA++G AA + TP DV+KTR+ A G++ ++ ILR
Sbjct: 514 EDG-QVSPGSLLLAGAIAGMPAASLVTPADVIKTRLQVAARAGQTTYSGVIDCFRKILRE 572
Query: 750 EGPLGLFKGAVPRFFWIAPLGAMNFAGYELARK 782
EGP L+KGA R F +P + YEL ++
Sbjct: 573 EGPKALWKGAGARVFRSSPQFGVTLLTYELLQR 605
Score = 53.1 bits (126), Expect = 3e-06
Identities = 90/453 (19%), Positives = 180/453 (38%), Gaps = 45/453 (9%)
Query: 363 FEELDRDGDGQVTLEDLEIAMRR----RKLPRRYAKEFMSRTRSHLFSRSFGWKQFLSFM 418
F+ D+ G G+VT ED++ + + +P + EF+ R + +F F+
Sbjct: 96 FQLFDKAGKGEVTFEDVKQVFGQTTIHQHIPFNWDSEFVQLHFGKERKRHLTYAEFTQFL 155
Query: 419 -EQKEPTILRAYTSLCLTKSGTLKK---SEILESLKNSGL-PANEDNAAAMMRFLNADTE 473
E + +A+ ++G + +I+ +++ L P E+ ++ T
Sbjct: 156 LEIQLEHAKQAFVQRDNARTGRVTAIDFRDIMVTIRPHVLTPFVEE---CLVAAAGGTTS 212
Query: 474 ESISYGHFRNFMLLLPSDRLQEDPRSIWFEAATVVAVPPSVEIPAGSVLRSALAGGLSCA 533
+S+ +F F LL + L R I+ +T+ VE+ + +A G
Sbjct: 213 HQVSFSYFNGFNSLLNNMELI---RKIY---STLAGTRKDVEVTKEEFVLAAQKFGQVTP 266
Query: 534 LSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGTRGLYRGSIPAILGQFSSHGLRTGIF 593
+ +L + + +++ E IA L E G++P L + +
Sbjct: 267 MEVDILFQLADLYEPRGRMTLADIERIAPLEE--------GTLPFNLAEAQR---QKASG 315
Query: 594 EASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQAG----------LF 643
++++ VL+ VA + + S+A +G P +++K R+Q ++
Sbjct: 316 DSARPVLLQVAESAYRFGLGSVAGA----VGATAVYPIDLVKTRMQNQRSTGSFVGELMY 371
Query: 644 NNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWE 703
N + + +G G +RG L P + + + G A E
Sbjct: 372 KNSFDCFKKVLRYEGFFGLYRGLLPQLLGVAPEKAIKLTVNDFVRDKFMHKDGSVPLAAE 431
Query: 704 TIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRF 763
+A G +GG + T P +++K R+ A + + A S++R G G++KGA F
Sbjct: 432 ILA-GGCAGGSQVIFTNPLEIVKIRLQVAGEITTGPRVSALSVVRDLGFFGIYKGAKACF 490
Query: 764 FWIAPLGAMNFAGYELARKAM-NKNDEAKTGNL 795
P A+ F Y + + N++ + G+L
Sbjct: 491 LRDIPFSAIYFPCYAHVKASFANEDGQVSPGSL 523
>UCP2_MOUSE (P70406) Mitochondrial uncoupling protein 2 (UCP 2)
(UCPH)
Length = 309
Score = 82.4 bits (202), Expect = 4e-15
Identities = 74/297 (24%), Positives = 136/297 (44%), Gaps = 29/297 (9%)
Query: 513 SVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRVQ-----------ASSMSFPEIIA 561
+ ++P + ++ AG +C ++ + P+D+ K R+Q A+S + ++
Sbjct: 6 ATDVPPTATVKFLGAGTAAC-IADLITFPLDTAKVRLQIQGESQGLVRTAASAQYRGVLG 64
Query: 562 KLPEI----GTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIAS 617
+ + G R LY G + + Q S +R G++++ K + + + + +A
Sbjct: 65 TILTMVRTEGPRSLYNGLVAGLQRQMSFASVRIGLYDSVKQFYTKGSEHAG-IGSRLLAG 123
Query: 618 FCSTFLGTAVRIPCEVLKQRLQAGL-------FNNVGEALVGTWQQDGLKGFFRGTGATL 670
+ L AV P +V+K R QA + + EA +++G++G ++GT +
Sbjct: 124 STTGALAVAVAQPTDVVKVRFQAQARAGGGRRYQSTVEAYKTIAREEGIRGLWKGTSPNV 183
Query: 671 CREVPFYVAGMGLYAESKKGVQK--LLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTR 728
R A + Y K + K L+ +L T A GA G V+ +P DV+KTR
Sbjct: 184 ARNAIVNCAELVTYDLIKDTLLKANLMTDDLPCHFTSAFGA--GFCTTVIASPVDVVKTR 241
Query: 729 MM-TAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAM 784
M +A G+ S A ++LR EGP +KG +P F + + F YE ++A+
Sbjct: 242 YMNSALGQYHSAGHCALTMLRKEGPRAFYKGFMPSFLRLGSWNVVMFVTYEQLKRAL 298
Score = 48.9 bits (115), Expect = 5e-05
Identities = 51/198 (25%), Positives = 81/198 (40%), Gaps = 21/198 (10%)
Query: 608 PELQVQSIASFCSTFLGTAVRIPCEVLKQRLQA-----GLFNNVGEA----LVGTW---- 654
P V+ + + + + + P + K RLQ GL A ++GT
Sbjct: 11 PTATVKFLGAGTAACIADLITFPLDTAKVRLQIQGESQGLVRTAASAQYRGVLGTILTMV 70
Query: 655 QQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEA-WETIAVGALSGG 713
+ +G + + G A L R++ F +GLY K+ K G E + G+ +G
Sbjct: 71 RTEGPRSLYNGLVAGLQRQMSFASVRIGLYDSVKQFYTK--GSEHAGIGSRLLAGSTTGA 128
Query: 714 LAAVVTTPFDVMKTRMMTAQ----GRSVSMSIVAF-SILRHEGPLGLFKGAVPRFFWIAP 768
LA V P DV+K R GR ++ A+ +I R EG GL+KG P A
Sbjct: 129 LAVAVAQPTDVVKVRFQAQARAGGGRRYQSTVEAYKTIAREEGIRGLWKGTSPNVARNAI 188
Query: 769 LGAMNFAGYELARKAMNK 786
+ Y+L + + K
Sbjct: 189 VNCAELVTYDLIKDTLLK 206
>CMC1_HUMAN (O75746) Calcium-binding mitochondrial carrier protein
Aralar1 (Mitochondrial aspartate glutamate carrier 1)
(Solute carrier family 25, member 12)
Length = 678
Score = 82.4 bits (202), Expect = 4e-15
Identities = 72/273 (26%), Positives = 123/273 (44%), Gaps = 20/273 (7%)
Query: 528 GGLSCALSCALLHPVDSIKTRVQ---ASSMSFPEIIAK---------LPEIGTRGLYRGS 575
G ++ A+ ++P+D +KTR+Q S E++ K L G GLYRG
Sbjct: 333 GSVAGAVGATAVYPIDLVKTRMQNQRGSGSVVGELMYKNSFDCFKKVLRYEGFFGLYRGL 392
Query: 576 IPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLK 635
IP ++G ++ + + + ++P L + +A C+ P E++K
Sbjct: 393 IPQLIGVAPEKAIKLTVNDFVRDKFTRRDGSVP-LPAEVLAGGCAGGSQVIFTNPLEIVK 451
Query: 636 QRLQAGLFNNVGEAL--VGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQK 693
RLQ G + + + G+ G ++G A R++PF +YA K +
Sbjct: 452 IRLQVAGEITTGPRVSALNVLRDLGIFGLYKGAKACFLRDIPFSAIYFPVYAHCKLLLAD 511
Query: 694 LLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTA--QGRSVSMSIVAF--SILRH 749
G + +A GA++G AA + TP DV+KTR+ A G++ ++ ILR
Sbjct: 512 ENGH-VGGLNLLAAGAMAGVPAASLVTPADVIKTRLQVAARAGQTTYSGVIDCFRKILRE 570
Query: 750 EGPLGLFKGAVPRFFWIAPLGAMNFAGYELARK 782
EGP +KG R F +P + A YE+ ++
Sbjct: 571 EGPSAFWKGTAARVFRSSPQFGVTLAHYEVLQR 603
Score = 64.3 bits (155), Expect = 1e-09
Identities = 101/482 (20%), Positives = 194/482 (39%), Gaps = 62/482 (12%)
Query: 324 ELIEMLPQLGKTTKDHPDKKKLFSVQDFFRYT------ESEGRRFFEELDRDGDGQVTLE 377
+++++L + TKD L S Q+F + +S F+ D+ G+G+VT E
Sbjct: 55 KIVQLLAGVADQTKDG-----LISYQEFLAFESVLCAPDSMFIVAFQLFDKSGNGEVTFE 109
Query: 378 DLEIAMRR----RKLPRRYAKEFMSRTRSHLFSRSFGWKQFLSFMEQKEPTILR-AYTSL 432
+++ + +P + EF+ H + + +F F+++ + R A+
Sbjct: 110 NVKEIFGQTIIHHHIPFNWDCEFIRLHFGHNRKKHLNYTEFTQFLQELQLEHARQAFALK 169
Query: 433 CLTKSGTLKK---SEILESLKNSGL-PANEDNAAAMMRFLNADTEESISYGHFRNFMLLL 488
+KSG + S+I+ ++++ L P E+N ++ +S+ +F F LL
Sbjct: 170 DKSKSGMISGLDFSDIMVTIRSHMLTPFVEEN---LVSAAGGSISHQVSFSYFNAFNSLL 226
Query: 489 PSDRLQEDPRSIWFEAATVVAVPPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTR 548
+ L R I+ +T+ VE+ +SA+ G L +L+ + +
Sbjct: 227 NNMELV---RKIY---STLAGTRKDVEVTKEEFAQSAIRYGQVTPLEIDILYQLADLYNA 280
Query: 549 VQASSMSFPEIIAKLPEIGTRGLYRGSIPAILGQFS---SHGLRTGIFEASKLVLVNVAP 605
+++ E IA L E G++P L + S GL I+ + +A
Sbjct: 281 SGRLTLADIERIAPLAE--------GALPYNLAELQRQQSPGLGRPIW-------LQIAE 325
Query: 606 NLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQAG----------LFNNVGEALVGTWQ 655
+ + S+A +G P +++K R+Q ++ N + +
Sbjct: 326 SAYRFTLGSVAGA----VGATAVYPIDLVKTRMQNQRGSGSVVGELMYKNSFDCFKKVLR 381
Query: 656 QDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGLA 715
+G G +RG L P + + + + G E +A G +GG
Sbjct: 382 YEGFFGLYRGLIPQLIGVAPEKAIKLTVNDFVRDKFTRRDGSVPLPAEVLA-GGCAGGSQ 440
Query: 716 AVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFA 775
+ T P +++K R+ A + + A ++LR G GL+KGA F P A+ F
Sbjct: 441 VIFTNPLEIVKIRLQVAGEITTGPRVSALNVLRDLGIFGLYKGAKACFLRDIPFSAIYFP 500
Query: 776 GY 777
Y
Sbjct: 501 VY 502
Score = 47.0 bits (110), Expect = 2e-04
Identities = 31/119 (26%), Positives = 55/119 (46%), Gaps = 17/119 (14%)
Query: 685 AESKKGVQKLLGRELEAWETIA-------VGALSGGLAAVVTTPFDVMKTRMMTAQGRSV 737
AE ++ LGR + W IA +G+++G + A P D++KTRM +G
Sbjct: 305 AELQRQQSPGLGRPI--WLQIAESAYRFTLGSVAGAVGATAVYPIDLVKTRMQNQRGSGS 362
Query: 738 SMSIVAFS--------ILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKND 788
+ + + +LR+EG GL++G +P+ +AP A+ + R + D
Sbjct: 363 VVGELMYKNSFDCFKKVLRYEGFFGLYRGLIPQLIGVAPEKAIKLTVNDFVRDKFTRRD 421
>CMC1_DROME (Q9VA73) Calcium-binding mitochondrial carrier Aralar1
Length = 695
Score = 82.0 bits (201), Expect = 6e-15
Identities = 73/277 (26%), Positives = 122/277 (43%), Gaps = 29/277 (10%)
Query: 528 GGLSCALSCALLHPVDSIKTRVQ-----------ASSMSFPEIIAKLPEIGTRGLYRGSI 576
G + A+ +++P+D +KTR+Q A S+ + G GLYRG +
Sbjct: 349 GSFAGAVGATVVYPIDLVKTRMQNQRAGSYIGEVAYRNSWDCFKKVVRHEGFMGLYRGLL 408
Query: 577 PAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQ 636
P ++G ++ + + + L + N+P + +A C+ P E++K
Sbjct: 409 PQLMGVAPEKAIKLTVNDLVRDKLTDKKGNIPTW-AEVLAGGCAGASQVVFTNPLEIVKI 467
Query: 637 RLQAGLFNNVGEALVGT----W---QQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKK 689
RLQ GE G+ W ++ GL G ++G A L R+VPF YA +K
Sbjct: 468 RLQVA-----GEIASGSKIRAWSVVRELGLFGLYKGARACLLRDVPFSAIYFPTYAHTKA 522
Query: 690 GVQKLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRM--MTAQGRSVSMSI--VAFS 745
+ G +A GA++G AA + TP DV+KTR+ + G++ +
Sbjct: 523 MMADKDGYN-HPLTLLAAGAIAGVPAASLVTPADVIKTRLQVVARSGQTTYTGVWDATKK 581
Query: 746 ILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARK 782
I+ EGP +KG R F +P + YEL ++
Sbjct: 582 IMAEEGPRAFWKGTAARVFRSSPQFGVTLVTYELLQR 618
Score = 50.4 bits (119), Expect = 2e-05
Identities = 46/186 (24%), Positives = 77/186 (40%), Gaps = 15/186 (8%)
Query: 614 SIASFCSTFLGTAVRIPCEVLKQRLQ---AGLFNNVGE-ALVGTW-------QQDGLKGF 662
++ SF +G V P +++K R+Q AG + +GE A +W + +G G
Sbjct: 347 TLGSFAGA-VGATVVYPIDLVKTRMQNQRAGSY--IGEVAYRNSWDCFKKVVRHEGFMGL 403
Query: 663 FRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGLAAVVTTPF 722
+RG L P + + + + G + W + G +G V T P
Sbjct: 404 YRGLLPQLMGVAPEKAIKLTVNDLVRDKLTDKKGN-IPTWAEVLAGGCAGASQVVFTNPL 462
Query: 723 DVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARK 782
+++K R+ A + I A+S++R G GL+KGA P A+ F Y +
Sbjct: 463 EIVKIRLQVAGEIASGSKIRAWSVVRELGLFGLYKGARACLLRDVPFSAIYFPTYAHTKA 522
Query: 783 AMNKND 788
M D
Sbjct: 523 MMADKD 528
Score = 48.9 bits (115), Expect = 5e-05
Identities = 31/105 (29%), Positives = 53/105 (49%), Gaps = 13/105 (12%)
Query: 699 LEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAF--------SILRHE 750
LE+ +G+ +G + A V P D++KTRM + S + VA+ ++RHE
Sbjct: 340 LESSYRFTLGSFAGAVGATVVYPIDLVKTRMQNQRAGSY-IGEVAYRNSWDCFKKVVRHE 398
Query: 751 GPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDEAKTGNL 795
G +GL++G +P+ +AP A+ +L R + K GN+
Sbjct: 399 GFMGLYRGLLPQLMGVAPEKAIKLTVNDLVRDKLTD----KKGNI 439
Score = 38.9 bits (89), Expect = 0.054
Identities = 36/160 (22%), Positives = 69/160 (42%), Gaps = 11/160 (6%)
Query: 526 LAGGLSCALSCALLHPVDSIKTRVQA----SSMSFPEIIAKLPEIGTRGLYRGSIPAILG 581
LAGG + A +P++ +K R+Q +S S + + E+G GLY+G+ +L
Sbjct: 446 LAGGCAGASQVVFTNPLEIVKIRLQVAGEIASGSKIRAWSVVRELGLFGLYKGARACLLR 505
Query: 582 QFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQA- 640
+ + +K ++ + L + + + + V P +V+K RLQ
Sbjct: 506 DVPFSAIYFPTYAHTKAMMADKDGYNHPLTLLAAGAIAGVPAASLV-TPADVIKTRLQVV 564
Query: 641 -----GLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVP 675
+ V +A ++G + F++GT A + R P
Sbjct: 565 ARSGQTTYTGVWDATKKIMAEEGPRAFWKGTAARVFRSSP 604
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.134 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 89,954,868
Number of Sequences: 164201
Number of extensions: 3823761
Number of successful extensions: 11427
Number of sequences better than 10.0: 266
Number of HSP's better than 10.0 without gapping: 74
Number of HSP's successfully gapped in prelim test: 192
Number of HSP's that attempted gapping in prelim test: 10164
Number of HSP's gapped (non-prelim): 822
length of query: 796
length of database: 59,974,054
effective HSP length: 118
effective length of query: 678
effective length of database: 40,598,336
effective search space: 27525671808
effective search space used: 27525671808
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 70 (31.6 bits)
Medicago: description of AC144484.11


