

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144481.6 - phase: 0
(754 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
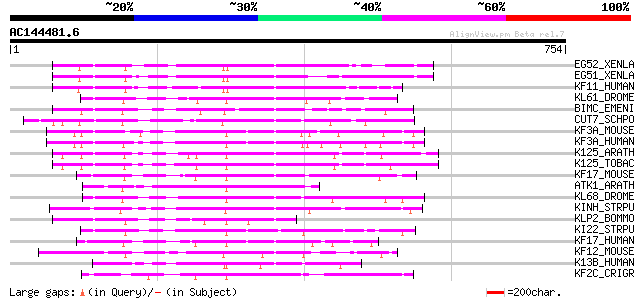
EG52_XENLA (Q91783) Kinesin-related motor protein Eg5 2 150 1e-35
EG51_XENLA (P28025) Kinesin-related motor protein Eg5 1 149 2e-35
KF11_HUMAN (P52732) Kinesin-like protein KIF11 (Kinesin-related ... 146 2e-34
KL61_DROME (P46863) Bipolar kinesin KRP-130 (Kinesin-like protei... 142 3e-33
BIMC_EMENI (P17120) Kinesin-like protein bimC 142 4e-33
CUT7_SCHPO (P24339) Kinesin-like protein cut7 140 2e-32
KF3A_MOUSE (P28741) Kinesin-like protein KIF3A (Microtubule plus... 138 6e-32
KF3A_HUMAN (Q9Y496) Kinesin-like protein KIF3A (Microtubule plus... 137 1e-31
K125_ARATH (P82266) Probable 125 kDa kinesin-related protein 137 1e-31
K125_TOBAC (O23826) 125 kDa kinesin-related protein 136 2e-31
KF17_MOUSE (Q99PW8) Kinesin-like protein KIF17 (MmKIF17) 134 7e-31
ATK1_ARATH (Q07970) Kinesin 1 (Kinesin-like protein A) 134 9e-31
KL68_DROME (P46867) Kinesin-like protein KLP68D 132 3e-30
KINH_STRPU (P35978) Kinesin heavy chain 127 8e-29
KLP2_BOMMO (P46874) Kinesin-like protein KLP2 (Fragment) 127 1e-28
KI22_STRPU (P46872) Kinesin-II 85 kDa subunit (KRP-85/95 85 kDa ... 127 1e-28
KF17_HUMAN (Q9P2E2) Kinesin-like protein KIF17 (KIF3-related mot... 125 3e-28
KF12_MOUSE (Q9D2Z8) Kinesin-like protein KIF12 125 5e-28
K13B_HUMAN (Q9NQT8) Kinesin-like protein KIF13B (Kinesin-like pr... 124 7e-28
KF2C_CRIGR (P70096) Kinesin-like protein KIF2C (Mitotic centrome... 124 9e-28
>EG52_XENLA (Q91783) Kinesin-related motor protein Eg5 2
Length = 1067
Score = 150 bits (379), Expect = 1e-35
Identities = 153/552 (27%), Positives = 248/552 (44%), Gaps = 86/552 (15%)
Query: 59 IEVIARIRDYP--DRKDKPLSVLQASSNSRSIRVKAD-----FGYRDFTLDGVSVSEEEE 111
I+V+ R R + +RK SVL+ S + + V+ G + +T D V ++
Sbjct: 19 IQVVVRCRPFNQLERKASSHSVLECESQRKEVCVRTGEVNDKLGKKTYTFDMVFGPAAKQ 78
Query: 112 LDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ-----------AGIVY 160
+D+ Y+ V ++ V +G CTI YG TG+GK+ TM G AGI+
Sbjct: 79 IDV-YRSVVCPILDEVIMGYNCTIFAYGQTGTGKTFTMEGERSSDEEFTWEQDPLAGIIP 137
Query: 161 RALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGG 220
R L I K +G V+V++LEIYNEE++DLLS + G
Sbjct: 138 RTLHQIF--------------EKLSEIGTEFSVKVSLLEIYNEELFDLLSPSPDVG---- 179
Query: 221 FGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSR 280
+ G K IS + ++ + +++ +R STL N SSR
Sbjct: 180 -----ERLQMFDDPRNKRGVIIKGLEEISVHNKDEVYQILERGAAKRKTASTLMNAYSSR 234
Query: 281 SHCMVILDV----PTVGG-------RLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNI 329
SH + + + T+ G +L LVD+AGSENI ++G A+ + INQ +
Sbjct: 235 SHSVFSVTIHMKETTIDGEELVKIGKLNLVDLAGSENIGRSGAVDKRAR-EAGNINQSLL 293
Query: 330 ALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEY 389
L RV+ ++ H+P+R+SKLT +LQDS ++K +I SP + +T+STL+Y
Sbjct: 294 TLGRVITALVERAPHIPYRESKLTRILQDSL-GGRTKTSIIATVSPASINLEETMSTLDY 352
Query: 390 GAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMEN--KLREKERNEAHKKL 447
++AK I+ P K + I + + + +N L + + K+
Sbjct: 353 ASRAKNIMNKPEVNQKLTKKALIKEYTEEIERLKRELATAREKNGVYLSNENYEQLQGKV 412
Query: 448 MKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFVELERKR 507
+ +EE I K+ A EEEI +R L + +K+LEEC
Sbjct: 413 LSQEEMITEYSEKI---AAMEEEI-----KRIGELFADNKKELEEC-------------- 450
Query: 508 MEERILQ-QQEEVEILRKRLEEIELQLCSSSKQERKDENESKEMEPNGFMRKLLSVYKST 566
ILQ +++E+E + L+E + QL + E K++ +G KLLS + T
Sbjct: 451 --TTILQCKEKELEATQNNLQESKEQLAQEAFVVSAMETTEKKL--HGTANKLLSTVRET 506
Query: 567 --DDLGMVKSMD 576
D G+ + +D
Sbjct: 507 TRDVSGLHEKLD 518
>EG51_XENLA (P28025) Kinesin-related motor protein Eg5 1
Length = 1060
Score = 149 bits (377), Expect = 2e-35
Identities = 159/556 (28%), Positives = 251/556 (44%), Gaps = 94/556 (16%)
Query: 59 IEVIARIRDYP--DRKDKPLSVLQASSNSRSIRVKAD-----FGYRDFTLDGVSVSEEEE 111
I+V+ R R + +RK SVL+ S + + V+ G + +T D V ++
Sbjct: 12 IQVVVRCRPFNQLERKASSHSVLECDSQRKEVYVRTGEVNDKLGKKTYTFDMVFGPAAKQ 71
Query: 112 LDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ-----------AGIVY 160
+++ Y+ V ++ V +G CTI YG TG+GK+ TM G AGI+
Sbjct: 72 IEV-YRSVVCPILDEVIMGYNCTIFAYGQTGTGKTFTMEGERSSDEEFTWEQDPLAGIIP 130
Query: 161 RALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGG 220
R L I + SE G V+V++LEIYNEE++DLLS + G
Sbjct: 131 RTLHQIF---EKLSEN-----------GTEFSVKVSLLEIYNEELFDLLSPSPDVG---- 172
Query: 221 FGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSR 280
+ G K IS + ++ +++ RR STL N SSR
Sbjct: 173 -----ERLQMFDDPRNKRGVIIKGLEEISVHNKDEVYHILERGAARRKTASTLMNAYSSR 227
Query: 281 SHCMVILDV----PTVGG-------RLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNI 329
SH + + + TV G +L LVD+AGSENI ++G A+ + INQ +
Sbjct: 228 SHSVFSVTIHMKETTVDGEELVKIGKLNLVDLAGSENIGRSGAVDKRAR-EAGNINQSLL 286
Query: 330 ALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEY 389
L RV+ ++ H+P+R+SKLT +LQDS ++K +I SP + +T+STL+Y
Sbjct: 287 TLGRVITALVERTPHIPYRESKLTRILQDSL-GGRTKTSIIATVSPASINLEETVSTLDY 345
Query: 390 GAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAHKKLMK 449
+AK I+ P K + K KE E ++L
Sbjct: 346 ANRAKSIMNKPEVNQK-------------------------LTKKALIKEYTEEIERL-- 378
Query: 450 KEEEIAALRTKVETAPASE--EEINLKVNERTRHLRQELEK--KLEECQRMTNEFVELER 505
+ E+AA R K +SE E++ KV + + + EK +EE + +E +
Sbjct: 379 -KRELAAAREKNGVYLSSENYEQLQGKVLSQEEMITEYTEKITAMEEELKSISELFADNK 437
Query: 506 KRMEE--RILQ-QQEEVEILRKRLEEIELQLCSSSKQERKDENESKEMEPNGFMRKLLSV 562
K +EE ILQ +++E+E + L+E + QL S E K++ +G KLLS
Sbjct: 438 KELEECTTILQCKEKELEETQNHLQESKEQLAQESFVVSAFETTEKKL--HGTANKLLST 495
Query: 563 YKST--DDLGMVKSMD 576
+ T D G+ + +D
Sbjct: 496 VRETTRDVSGLHEKLD 511
>KF11_HUMAN (P52732) Kinesin-like protein KIF11 (Kinesin-related
motor protein Eg5) (Kinesin-like spindle protein HKSP)
(Thyroid receptor interacting protein 5) (TRIP5)
(Kinesin-like protein 1)
Length = 1057
Score = 146 bits (368), Expect = 2e-34
Identities = 148/506 (29%), Positives = 239/506 (46%), Gaps = 63/506 (12%)
Query: 59 IEVIARIRDY--PDRKDKPLSVLQASSNSRSIRVK----ADFGYRD-FTLDGVSVSEEEE 111
I+V+ R R + +RK S+++ + + V+ AD R +T D V + ++
Sbjct: 19 IQVVVRCRPFNLAERKASAHSIVECDPVRKEVSVRTGGLADKSSRKTYTFDMVFGASTKQ 78
Query: 112 LDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ-----------AGIVY 160
+D+ Y+ V ++ V +G CTI YG TG+GK+ TM G AGI+
Sbjct: 79 IDV-YRSVVCPILDEVIMGYNCTIFAYGQTGTGKTFTMEGERSPNEEYTWEEDPLAGIIP 137
Query: 161 RALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGG 220
R L I + TD+ G V+V++LEIYNEE++DLL+ +
Sbjct: 138 RTLHQIF-EKLTDN-------------GTEFSVKVSLLEIYNEELFDLLNPSSD------ 177
Query: 221 FGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSR 280
+ + K V+ K + T + +E +I ++K +R +TL N SSR
Sbjct: 178 VSERLQMFDDPRNKRGVIIKGLEEITVHNKDEVYQI---LEKGAAKRTTAATLMNAYSSR 234
Query: 281 SHCMVILDV----PTVGG-------RLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNI 329
SH + + + T+ G +L LVD+AGSENI ++G A+ + INQ +
Sbjct: 235 SHSVFSVTIHMKETTIDGEELVKIGKLNLVDLAGSENIGRSGAVDKRAR-EAGNINQSLL 293
Query: 330 ALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEY 389
L RV+ ++ HVP+R+SKLT +LQDS +++ +I SP + +T+STLEY
Sbjct: 294 TLGRVITALVERTPHVPYRESKLTRILQDSL-GGRTRTSIIATISPASLNLEETLSTLEY 352
Query: 390 GAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERN--EAHKKL 447
+AK I+ P K + I + + + +N + E N KL
Sbjct: 353 AHRAKNILNKPEVNQKLTKKALIKEYTEEIERLKRDLAAAREKNGVYISEENFRVMSGKL 412
Query: 448 MKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFVELERKR 507
+EE+I L +E A EEE+N +V E + EL++ + Q T E +E +K
Sbjct: 413 TVQEEQIVEL---IEKIGAVEEELN-RVTELFMDNKNELDQCKSDLQNKTQE-LETTQKH 467
Query: 508 MEERILQQQEEVEILRKRLEEIELQL 533
++E LQ +E E + LE E +L
Sbjct: 468 LQETKLQLVKE-EYITSALESTEEKL 492
>KL61_DROME (P46863) Bipolar kinesin KRP-130 (Kinesin-like protein
Klp61F)
Length = 1066
Score = 142 bits (358), Expect = 3e-33
Identities = 134/473 (28%), Positives = 218/473 (45%), Gaps = 85/473 (17%)
Query: 97 RDFTLDGVSVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCS--- 153
+ FT D E ++ D+ Y V I V G CT+ YG TG+GK+HTM G
Sbjct: 62 KKFTFDRSFGPESKQCDV-YSVVVSPLIEEVLNGYNCTVFAYGQTGTGKTHTMVGNETAE 120
Query: 154 --------KQAGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEI 205
GI+ RAL + + + V ++++ LE+YNEE+
Sbjct: 121 LKSSWEDDSDIGIIPRALSHLFDELRM--------------MEVEYTMRISYLELYNEEL 166
Query: 206 YDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISGNEA------GKISKE 259
DLLST+ + +K+++ K K + I G E + K
Sbjct: 167 CDLLSTD----------------DTTKIRIFDDSTK-KGSVIIQGLEEIPVHSKDDVYKL 209
Query: 260 IQKVEKRRIVKSTLCNDRSSRSHCM--VILDVPTVG---------GRLMLVDMAGSENIE 308
++K ++RR +TL N +SSRSH + +++ + G G+L LVD+AGSEN+
Sbjct: 210 LEKGKERRKTATTLMNAQSSRSHTVFSIVVHIRENGIEGEDMLKIGKLNLVDLAGSENVS 269
Query: 309 QAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKIL 368
+AG +T INQ + L RV+ ++ + HVP+R+SKLT LLQ+S ++K
Sbjct: 270 KAGNEKGIRVRETVNINQSLLTLGRVITALVDRAPHVPYRESKLTRLLQESL-GGRTKTS 328
Query: 369 MILCASPDPKEIHKTISTLEYGAKAKCIVRGPH--------TPVKEEDSSSTVILGSRIA 420
+I SP K+I +T+STLEY +AK I P T +KE + +A
Sbjct: 329 IIATISPGHKDIEETLSTLEYAHRAKNIQNKPEVNQKLTKKTVLKEYTEEIDKLKRDLMA 388
Query: 421 AMDEFIMKLQMEN------KLREKERNEAHKKLMKKEEEIAALRTKVETAPASEEEINLK 474
A D+ + L E KL + R K L+ K AL+ +++ E+++
Sbjct: 389 ARDKNGIYLAEETYGEITLKLESQNRELNEKMLLLK-----ALKDELQNKEKIFSEVSMS 443
Query: 475 VNERTRHLRQELEKKLEECQRMTNEFVELERKRMEERILQQQEEVEILRKRLE 527
+ E+T+ L KK EE T + L +K + + + +E+ E++ ++
Sbjct: 444 LVEKTQEL-----KKTEENLLNTKGTLLLTKKVLTKTKRRYKEKKELVASHMK 491
>BIMC_EMENI (P17120) Kinesin-like protein bimC
Length = 1184
Score = 142 bits (357), Expect = 4e-33
Identities = 147/533 (27%), Positives = 240/533 (44%), Gaps = 96/533 (18%)
Query: 59 IEVIARIRDYPDRKDKPLS-VLQASSNSRSIRVKADFG-----YRDFTLDGVSVSEEEEL 112
I V+ R R +R+ K S V+ + + V+ G + +T D V + +++
Sbjct: 82 IHVVVRCRGRNEREVKENSGVVLQTEGVKGKTVELSMGPNAVSNKTYTFDKVFSAAADQI 141
Query: 113 DLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFG--------CSKQAGIVYRALR 164
+ Y+ V + + G CTI YG TG+GK++TM G S AGI+ R L
Sbjct: 142 TV-YEDVVLPIVTEMLAGYNCTIFAYGQTGTGKTYTMSGDMTDTLGILSDNAGIIPRVLY 200
Query: 165 DILGD-GDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGF 223
+ DT+S V+ + +E+YNEE+ DLLS
Sbjct: 201 SLFAKLADTEST-----------------VKCSFIELYNEELRDLLS------------- 230
Query: 224 GWSKSNASKVKLEVMGKKAKNATYISGNEAGKIS------KEIQKVEKRRIVKSTLCNDR 277
++ N + KK +T + G E I K +Q+ +R V +T CND
Sbjct: 231 --AEENPKLKIYDNEQKKGHMSTLVQGMEETYIDSATAGIKLLQQGSHKRQVAATKCNDL 288
Query: 278 SSRSHCMVILDVP-----------TVGGRLMLVDMAGSENIEQAGQTGFEAKMQTAK--I 324
SSRSH + + V G+L LVD+AGSENI G++G E K T I
Sbjct: 289 SSRSHTVFTITVNIKRTTESGEEYVCPGKLNLVDLAGSENI---GRSGAENKRATEAGLI 345
Query: 325 NQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTI 384
N+ + L RV+ ++ + H+P+R+SKLT LLQDS ++K +I SP + +TI
Sbjct: 346 NKSLLTLGRVINALVDKSQHIPYRESKLTRLLQDSL-GGRTKTCIIATMSPARSNLEETI 404
Query: 385 STLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAH 444
STL+Y +AK I P +ST+ + + I KL+ E + + RN +
Sbjct: 405 STLDYAFRAKNIRNKPQI-------NSTMPKMTLLREFTAEIEKLKAE-LIATRHRNGVY 456
Query: 445 KKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNEFVELE 504
+ EE ++ + E+ EE K+ + L K++E +T++F +L+
Sbjct: 457 MSVESYEE----MKMENESRRIISEEQRAKIES----MESSLRHKVQELLTLTSKFNDLK 508
Query: 505 RKRME--------ERILQQQEEV-EILRKRLEEIELQLCSSSKQERKDENESK 548
+ + +LQQ + V + R +LEE E+ C+ + E + ++ K
Sbjct: 509 KDNDDTLAALCSTNDVLQQTDIVLQNTRAQLEEEEMLRCAHEETEHQLQDVGK 561
>CUT7_SCHPO (P24339) Kinesin-like protein cut7
Length = 1085
Score = 140 bits (352), Expect = 2e-32
Identities = 158/588 (26%), Positives = 257/588 (42%), Gaps = 94/588 (15%)
Query: 19 TPRSKHRLNFNGVKPAPTPPHPHPSPHPNFNNKDSPPEHP--------IEVIARIRDYPD 70
TP S R N K P + +P+ ++ +SP +H I V+ R+R D
Sbjct: 27 TPNSHFRSASNPRKRREPPTID--TGYPDRSDTNSPTDHALHDENETNINVVVRVRGRTD 84
Query: 71 ---RKDKPLSVLQASSNSRSIRVKAD----FGYRDFTLDGVSVSEEEELDLFYKKFVESR 123
R + L+V + + + +++D + + D V E ++L LF + V
Sbjct: 85 QEVRDNSSLAVSTSGAMGAELAIQSDPSSMLVTKTYAFDKVFGPEADQLMLF-ENSVAPM 143
Query: 124 INGVKLGDKCTIMMYGPTGSGKSHTMFG--------CSKQAGIVYRALRDILGDGDTDSE 175
+ V G CTI YG TG+GK++TM G S+ AG++ RAL + D ++
Sbjct: 144 LEQVLNGYNCTIFAYGQTGTGKTYTMSGDLSDSDGILSEGAGLIPRALYQLFSSLDNSNQ 203
Query: 176 GSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKL 235
V+ + E+YNEEI DLL + F + N +
Sbjct: 204 --------------EYAVKCSYYELYNEEIRDLLVSEELRKPARVFEDTSRRGNVVITGI 249
Query: 236 EVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVIL-------- 287
E +YI AG + +++ RR V +T CND SSRSH + +
Sbjct: 250 E--------ESYIKN--AGDGLRLLREGSHRRQVAATKCNDLSSRSHSIFTITLHRKVSS 299
Query: 288 ---------------DVPTVGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALK 332
D +L +VD+AGSENI ++G A+ +T INQ + L
Sbjct: 300 GMTDETNSLTINNNSDDLLRASKLHMVDLAGSENIGRSGAENKRAR-ETGMINQSLLTLG 358
Query: 333 RVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAK 392
RV+ ++ H+P+R+SKLT LLQDS K+K MI+ S + +TISTLEY A+
Sbjct: 359 RVINALVEKAHHIPYRESKLTRLLQDSL-GGKTKTSMIVTVSSTNTNLEETISTLEYAAR 417
Query: 393 AKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENK-----LREKERNEAHKKL 447
AK I + P + V++ + ++ L K L E E ++
Sbjct: 418 AKSI---RNKPQNNQLVFRKVLIKDLVLDIERLKNDLNATRKKNGVYLAESTYKELMDRV 474
Query: 448 MKKEEEIAALRTKVETAPASEEEINLKVN-ERTRHL---RQELEKKLEECQ-RMTNEFVE 502
K+ K+E ++N+K + E+ +++ QE +K++E Q ++ N E
Sbjct: 475 QNKDLLCQEQARKLEVL-----DLNVKSSREQLQYVSKSNQEHKKEVEALQLQLVNSSTE 529
Query: 503 LERKRMEERILQQQEEVEI-LRKRLEEIELQLCSSSKQERKDENESKE 549
LE + E L+ + +EI RK+ E E ++ + + + ESKE
Sbjct: 530 LESVKSENEKLKNELVLEIEKRKKYETNEAKITTVATDLSQYYRESKE 577
>KF3A_MOUSE (P28741) Kinesin-like protein KIF3A (Microtubule plus
end-directed kinesin motor 3A)
Length = 701
Score = 138 bits (347), Expect = 6e-32
Identities = 169/584 (28%), Positives = 257/584 (43%), Gaps = 105/584 (17%)
Query: 50 NKDSPPEH--PIEVIARIRDYPDRKDKPLSVLQASSNSR---SIRV-KADFGY---RDFT 100
NK PE ++V+ R R +R +K + QA S +I V K D + FT
Sbjct: 4 NKSEKPESCDNVKVVVRCRPLNER-EKSMCYRQAVSVDEMRGTITVHKTDSSNEPPKTFT 62
Query: 101 LDGVSVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAGIVY 160
D V E ++LD+ Y I+ V G TI YG TG+GK+ TM G G
Sbjct: 63 FDTVFGPESKQLDV-YNLTARPIIDSVLEGYNGTIFAYGQTGTGKTFTMEGVRAVPG--- 118
Query: 161 RALRDILGDGDTDSEG----SDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGG 216
LR ++ + G ++GD+ R V+V+ LEIYNEE+ DLL
Sbjct: 119 --LRGVIPNSFAHIFGHIAKAEGDT--------RFLVRVSYLEIYNEEVRDLL------- 161
Query: 217 GGGGFGFGWSKSNASKVKLEV-MGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCN 275
G ++ +VK +G K+ + N A + + + K R V +T N
Sbjct: 162 -------GKDQTQRLEVKERPDVGVYIKDLSAYVVNNADDMDRIMTLGHKNRSVGATNMN 214
Query: 276 DRSSRSHCMVILDVPTVG-----------GRLMLVDMAGSENIEQAGQTGFEAKMQTAKI 324
+ SSRSH + + + G+L LVD+AGSE + G TG K T KI
Sbjct: 215 EHSSRSHAIFTITIECSEKGVDGNMHVRMGKLHLVDLAGSERQAKTGATGQRLKEAT-KI 273
Query: 325 NQGNIALKRVVESIANGDS-HVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKT 383
N L V+ ++ +G S HVP+R+SKLT LLQDS SK +M P +T
Sbjct: 274 NLSLSTLGNVISALVDGKSTHVPYRNSKLTRLLQDSL-GGNSKTMMCANIGPADYNYDET 332
Query: 384 ISTLEYGAKAK--------------CIVRGPHTPVKE--------EDSSSTVILGSRIAA 421
ISTL Y +AK ++R ++E E+ S + I GS
Sbjct: 333 ISTLRYANRAKNIKNKARINEDPKDALLRQFQKEIEELKKKLEEGEEVSGSDISGSE--- 389
Query: 422 MDEFIMKLQMENKLREKERNEAHK------KLMKKEEEIAALRTKVETAPASEEEINLKV 475
D+ +L + + R+K R++A K K+++ + +I R +ET EEE K
Sbjct: 390 EDDEEGELGEDGEKRKKRRDQAGKKKVSPDKMVEMQAKIDEERKALETKLDMEEEERNKA 449
Query: 476 NERTRHLRQELEKKLEECQRMTNEFVELERK------------RMEERILQQQE-EVEIL 522
++L K +E Q + + LE+K +E++L++ E+E
Sbjct: 450 RAELERREKDLLKAQQEHQSLLEKLSALEKKVIVGGVDLLAKAEEQEKLLEESNMELEER 509
Query: 523 RKRLEEIELQLCSSSKQERKDENE---SKEMEPNGFMRKLLSVY 563
R+R E++ +L +QER D E S + E G +KL V+
Sbjct: 510 RRRAEQLRKEL-EEKEQERLDIEEKYTSLQEEAQGKTKKLKKVW 552
>KF3A_HUMAN (Q9Y496) Kinesin-like protein KIF3A (Microtubule plus
end-directed kinesin motor 3A)
Length = 702
Score = 137 bits (344), Expect = 1e-31
Identities = 168/581 (28%), Positives = 258/581 (43%), Gaps = 98/581 (16%)
Query: 50 NKDSPPEH--PIEVIARIRDYPDRKDKPLSVLQASSNSR---SIRV-KADFGY---RDFT 100
NK PE ++V+ R R +R +K + QA S +I V K D + FT
Sbjct: 4 NKSEKPESCDNVKVVVRCRPLNER-EKSMCYKQAVSVDEMRGTITVHKTDSSNEPPKTFT 62
Query: 101 LDGVSVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCS---KQAG 157
D V E ++LD+ Y I+ V G TI YG TG+GK+ TM G + G
Sbjct: 63 FDTVFGPESKQLDV-YNLTARPIIDSVLEGYNGTIFAYGQTGTGKTFTMEGVRAIPELRG 121
Query: 158 IVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGG 217
I+ + I G ++GD+ R V+V+ LEIYNEE+ DLL
Sbjct: 122 IIPNSFAHIFGH----IAKAEGDT--------RFLVRVSYLEIYNEEVRDLL-------- 161
Query: 218 GGGFGFGWSKSNASKVKLEV-MGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCND 276
G ++ +VK +G K+ + N A + + + K R V +T N+
Sbjct: 162 ------GKDQTQRLEVKERPDVGVYIKDLSAYVVNNADDMDRIMTLGHKNRSVGATNMNE 215
Query: 277 RSSRSHCMVILDVPTVG-----------GRLMLVDMAGSENIEQAGQTGFEAKMQTAKIN 325
SSRSH + + + G+L LVD+AGSE + G TG K T KIN
Sbjct: 216 HSSRSHAIFTITIECSEKGIDGNMHVRMGKLHLVDLAGSERQAKTGATGQRLKEAT-KIN 274
Query: 326 QGNIALKRVVESIANGDS-HVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTI 384
L V+ ++ +G S HVP+R+SKLT LLQDS SK +M P +TI
Sbjct: 275 LSLSTLGNVISALVDGKSTHVPYRNSKLTRLLQDSL-GGNSKTMMCANIGPADYNYDETI 333
Query: 385 STLEYGAKAKCI---VRGPHTP--------------VKEEDSSSTVILGSRIAAM---DE 424
STL Y +AK I R P +K++ I GS I+ D+
Sbjct: 334 STLRYANRAKNIKNKARINEDPKDALLRQFQKEIEELKKKLEEGEEISGSDISGSEEDDD 393
Query: 425 FIMKLQMENKLREKERNEAHKKLM----------KKEEEIAALRTKVETAPASEEEINLK 474
++ + + R+K R++ KK + K +EE AL TK++ + +
Sbjct: 394 EEGEVGEDGEKRKKRRDQTGKKKVSPDKMIEMQAKIDEERKALETKLDMEEEERNKARAE 453
Query: 475 VNERTRHL---RQELEKKLEECQRMTNEFVE-----LERKRMEERILQQQE-EVEILRKR 525
+ +R + L +QE + LE+ + + + L + +E++L++ E+E RKR
Sbjct: 454 LEKREKDLLKAQQEHQSLLEKLSALEKKVIVGGVDLLAKAEEQEKLLEESNMELEERRKR 513
Query: 526 LEEIELQLCSSSKQERKDENE---SKEMEPNGFMRKLLSVY 563
E++ +L +QER D E S + E G +KL V+
Sbjct: 514 AEQLRREL-EEKEQERLDIEEKYTSLQEEAQGKTKKLKKVW 553
>K125_ARATH (P82266) Probable 125 kDa kinesin-related protein
Length = 1056
Score = 137 bits (344), Expect = 1e-31
Identities = 155/578 (26%), Positives = 260/578 (44%), Gaps = 96/578 (16%)
Query: 59 IEVIARIRDYPD---RKDKPLSVLQASSNSRSIRVKADFGY----RDFTLDGVSVSEEEE 111
++V+ R R + D R + P VL + R + V + R FT D V ++
Sbjct: 13 VQVLLRCRPFSDDELRSNAP-QVLTCNDLQREVAVSQNIAGKHIDRVFTFDKVFGPSAQQ 71
Query: 112 LDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFG-CSK-----------QAGIV 159
DL Y + V +N V G CTI YG TG+GK++TM G C + +AG++
Sbjct: 72 KDL-YDQAVVPIVNEVLEGFNCTIFAYGQTGTGKTYTMEGECRRSKSAPCGGLPAEAGVI 130
Query: 160 YRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGG 219
RA++ I DT EG + S V+VT LE+YNEEI DLL+
Sbjct: 131 PRAVKQIF---DT-LEGQQAEYS----------VKVTFLELYNEEITDLLAPE------- 169
Query: 220 GFGFGWSKSNASKVKLEVMGKK-------AKNATYISGNE------AGKISKEIQKVEKR 266
+ S+V E KK K + G E A +I +++ +
Sbjct: 170 ---------DLSRVAAEEKQKKPLPLMEDGKGGVLVRGLEEEIVTSANEIFTLLERGSSK 220
Query: 267 RIVKSTLCNDRSSRSHCMVILDVPTVG-----------GRLMLVDMAGSENIEQAGQTGF 315
R T N +SSRSH + + + G+L LVD+AGSENI ++G
Sbjct: 221 RRTAETFLNKQSSRSHSLFSITIHIKEATPEGEELIKCGKLNLVDLAGSENISRSGARDG 280
Query: 316 EAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASP 375
A+ + +IN+ + L RV+ ++ HVP+RDSKLT LL+DS ++K +I SP
Sbjct: 281 RAR-EAGEINKSLLTLGRVISALVEHLGHVPYRDSKLTRLLRDSL-GGRTKTCIIATVSP 338
Query: 376 DPKEIHKTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKL 435
+ +T+STL+Y +AK I P K S+ L I + + + +N +
Sbjct: 339 AVHCLEETLSTLDYAHRAKNIRNKPEVNQKMMKSTLIKDLYGEIERLKAEVYASREKNGV 398
Query: 436 -REKER---NEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLE 491
KER E+ +K+M E+I + ++E EE+ K + R +L KL+
Sbjct: 399 YMPKERYYQEESERKVM--AEQIEQMGGQIENYQKQLEELQDKYVGQVREC-SDLTTKLD 455
Query: 492 ECQRMTNEFVEL-----ERKRMEERILQQQEEVEILRKRLEEIELQLCS--SSKQERKDE 544
++ ++ ++ E + + +++++ + +K+ E + +Q S E+ +
Sbjct: 456 ITEKNLSQTCKVLASTNEELKKSQYAMKEKDFIISEQKKSENVLVQQACILQSNLEKATK 515
Query: 545 NESKEMEPNGFMRKLLSVYKSTDDLGMVKSMDLDMDDQ 582
+ S + G KL S D+ +V + +++ +Q
Sbjct: 516 DNSSLHQKIGREDKL-----SADNRKVVDNYQVELSEQ 548
>K125_TOBAC (O23826) 125 kDa kinesin-related protein
Length = 1006
Score = 136 bits (343), Expect = 2e-31
Identities = 150/567 (26%), Positives = 255/567 (44%), Gaps = 75/567 (13%)
Query: 59 IEVIARIRDYPD---RKDKPLSVLQASSNSRSIRVKADFGY----RDFTLDGVSVSEEEE 111
++V+ R R + + R + P V+ + R + V + R FT D V ++
Sbjct: 10 VQVLLRCRPFSNDELRNNAP-QVVTCNDYQREVAVSQNIAGKHIDRIFTFDKVFGPSAQQ 68
Query: 112 LDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSK------------QAGIV 159
DL Y + + +N V G CTI YG TG+GK++TM G K +AG++
Sbjct: 69 RDL-YDQAIVPIVNEVLEGFNCTIFAYGQTGTGKTYTMEGECKRSKSGPNGELPQEAGVI 127
Query: 160 YRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGG 219
RA++ + DT E + + S V+VT LE+YNEEI DLL+
Sbjct: 128 PRAVKQVF---DT-LESQNAEYS----------VKVTFLELYNEEITDLLAPED------ 167
Query: 220 GFGFGWSKSNASKVKLEVMGKKAKNATYISGNE------AGKISKEIQKVEKRRIVKSTL 273
+ + K +L +M + K + G E A +I +++ +R TL
Sbjct: 168 ---LKVALEDRQKKQLPLM-EDGKGGVLVRGLEEEIVTSANEIFTLLERGSAKRRTAETL 223
Query: 274 CNDRSSRSHCMVILDVPTVG-----------GRLMLVDMAGSENIEQAGQTGFEAKMQTA 322
N +SSRSH + + + G+L LVD+AGSENI ++G A+ +
Sbjct: 224 LNKQSSRSHSLFSITIHIKEATPEGEELIKCGKLNLVDLAGSENISRSGAREGRAR-EAG 282
Query: 323 KINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHK 382
+IN+ + L RV+ ++ H+P+RDSKLT LL+DS ++K +I SP + +
Sbjct: 283 EINKSLLTLGRVINALVEHLGHIPYRDSKLTRLLRDSL-GGRTKTCIIATVSPAVHCLEE 341
Query: 383 TISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLR-EKER- 440
T+STL+Y +AK I P K S+ L I + + + +N + KER
Sbjct: 342 TLSTLDYAHRAKNIKNKPEVNQKMMKSTLIKDLYGEIERLKAEVYAAREKNGVYIPKERY 401
Query: 441 --NEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTN 498
E +K M ++I + +E EE+ + + + + +L KL+ Q+ N
Sbjct: 402 YQEENERKAM--ADQIEQMGVSIENHQKQFEELQSRHDSQVQQC-SDLTCKLDVTQKQLN 458
Query: 499 EFVELERKRMEERILQQQ---EEVEILRKRLEEIELQLCSSSKQERKDENESKEMEPNGF 555
+ +L EE++ Q Q +E + + ++ E L + R D +S + + F
Sbjct: 459 QTSKL-LAYTEEQLRQSQYTLKERDFIISEQKKAENALAHQACVLRADLEKSIQENASLF 517
Query: 556 MRKLLSVYKSTDDLGMVKSMDLDMDDQ 582
+ STD+ +V + ++ Q
Sbjct: 518 QKIAREDKLSTDNRSLVNNFQAELAKQ 544
>KF17_MOUSE (Q99PW8) Kinesin-like protein KIF17 (MmKIF17)
Length = 1038
Score = 134 bits (338), Expect = 7e-31
Identities = 128/490 (26%), Positives = 204/490 (41%), Gaps = 73/490 (14%)
Query: 92 ADFGYRDFTLDGVSVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFG 151
AD + FT DG E + Y + + GV G TI YG TGSGKS TM G
Sbjct: 45 ADEPPKQFTFDGAYYIEHFT-EQIYNEIAYPLVEGVTEGYNGTIFAYGQTGSGKSFTMQG 103
Query: 152 CSK---QAGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDL 208
Q GI+ RA + S + V+ + LEIYNE+++DL
Sbjct: 104 LPDPPCQRGIIPRAFEHVF-------------ESVQCAENTKFLVRASYLEIYNEDVHDL 150
Query: 209 LSTNGGGGGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISG------NEAGKISKEIQK 262
L + +K +LE + + + Y+ G + + + ++
Sbjct: 151 LGAD------------------TKQRLE-LKEHPEKGVYVKGLSMHTVHNVAQCERVMET 191
Query: 263 VEKRRIVKSTLCNDRSSRSHCMVILDVPTV-----------GGRLMLVDMAGSENIEQAG 311
K R V TL N SSRSH + +++ G+L LVD+AGSE + G
Sbjct: 192 GWKNRAVGYTLMNKDSSRSHSIFTINIEIYAVDERGKDHLRAGKLNLVDLAGSERQSKTG 251
Query: 312 QTGFEAKMQTAKINQGNIALKRVVESIANGD-SHVPFRDSKLTMLLQDSFEDDKSKILMI 370
TG E + KIN AL V+ ++ +G H+P+RDSKLT LLQDS +K LM+
Sbjct: 252 ATG-ERLKEATKINLSLSALGNVISALVDGRCKHIPYRDSKLTRLLQDSL-GGNTKTLMV 309
Query: 371 LCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVI--LGSRIAAMDEFIMK 428
C SP +T+STL Y +AK I P ED ++ I + + +
Sbjct: 310 ACLSPADNNYDETLSTLRYANRAKNIKNKPRI---NEDPKDALLREYQEEIKRLKAILAQ 366
Query: 429 LQMENKLREKERNEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEK 488
L + ++ EE++ + T + A ++ I + ER L+ + E
Sbjct: 367 QMGPGNLSALLSTQTPPGPVQSEEKLLSPTTVQQDTEAEKQLIREEYEERLARLKADYEA 426
Query: 489 KLEECQRMTNEFV------ELERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQERK 542
+ E R+ + +++ ++E + +++E IL+ + LC + R
Sbjct: 427 EQESRVRLQEDITAMRNSYDVKLSTLQENLRKEKETEAILKAEV------LCKTEVMSRA 480
Query: 543 DENESKEMEP 552
+ E P
Sbjct: 481 ELASGPEYSP 490
>ATK1_ARATH (Q07970) Kinesin 1 (Kinesin-like protein A)
Length = 793
Score = 134 bits (337), Expect = 9e-31
Identities = 112/337 (33%), Positives = 168/337 (49%), Gaps = 38/337 (11%)
Query: 99 FTLDGVSVSEEEELDLFYK--KFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFG---CS 153
FT D V E + ++F++ + V+S ++G K+ I YG TGSGK++TM G
Sbjct: 478 FTFDKVFNHEASQEEVFFEISQLVQSALDGYKV----CIFAYGQTGSGKTYTMMGRPEAP 533
Query: 154 KQAGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNG 213
Q G++ R+L I + S G+ G K +QV++LEIYNE I DLLSTN
Sbjct: 534 DQKGLIPRSLEQIFQA--SQSLGAQGWKYK---------MQVSMLEIYNETIRDLLSTNR 582
Query: 214 GGGGGGGFGFGWSKSNASKVKLEVMGKK-AKNATYISGNEAGKISKEIQKVEKRRIVKST 272
+ + +V G + T GKIS +Q+ + R V T
Sbjct: 583 TTSMDLVRADSGTSGKQYTITHDVNGHTHVSDLTIFDVCSVGKISSLLQQAAQSRSVGKT 642
Query: 273 LCNDRSSRSHCMVILDVPTVG--------GRLMLVDMAGSENIEQAGQTGFEAKMQTAKI 324
N++SSRSH + + + V G L L+D+AGSE + ++G TG K +T I
Sbjct: 643 QMNEQSSRSHFVFTMRISGVNESTEQQVQGVLNLIDLAGSERLSKSGATGDRLK-ETQAI 701
Query: 325 NQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTI 384
N+ AL V+ ++A + HVPFR+SKLT LLQ D SK LM + SPDP +++
Sbjct: 702 NKSLSALSDVIFALAKKEDHVPFRNSKLTYLLQPCLGGD-SKTLMFVNISPDPTSAGESL 760
Query: 385 STLEYGAKAK-CIVRGPHTPVKEEDSSSTVILGSRIA 420
+L + A+ C + P +ST +L SR++
Sbjct: 761 CSLRFAARVNACEIGIPRR------QTSTKLLDSRLS 791
Score = 42.0 bits (97), Expect = 0.006
Identities = 52/243 (21%), Positives = 105/243 (42%), Gaps = 26/243 (10%)
Query: 323 KINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHK 382
K+ + +A K+ V S+ + + ++ L D ++ + A + I +
Sbjct: 203 KVKEEKMAAKQKVTSLEDMYKRLQEYNTSLQQYNSKLQTDLETVRAALTRAEKEKSSILE 262
Query: 383 TISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNE 442
+STL RG ++++ SSS V+ I D + ++ ++ R++
Sbjct: 263 NLSTL----------RGHSKSLQDQLSSSRVLQDDAIKQKDSLLSEVTNLRNELQQVRDD 312
Query: 443 AHKKLMKKEEEIAALRTKVETAPASEEEINLKVN-----ERTRHLRQE----LEKKL--- 490
+++++ ++ +R E S +E+++ E T L++E LE++L
Sbjct: 313 RDRQVVQSQKLSEEIRKYQENVGKSSQELDILTAKSGSLEETCSLQKERLNMLEQQLAIA 372
Query: 491 EECQRMTNEFVELERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQERKDENESKEM 550
E Q+M + V L R EE Q+ + L+ RL ++E QLC +K N E+
Sbjct: 373 NERQKMADASVSLTRTEFEE----QKHLLCELQDRLADMEHQLCEGELLRKKLHNTILEL 428
Query: 551 EPN 553
+ N
Sbjct: 429 KGN 431
>KL68_DROME (P46867) Kinesin-like protein KLP68D
Length = 784
Score = 132 bits (332), Expect = 3e-30
Identities = 140/506 (27%), Positives = 215/506 (41%), Gaps = 76/506 (15%)
Query: 99 FTLDGVSVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGC---SKQ 155
FT D + + L Y + V ++ V G I YG TG+GK+ TM G +
Sbjct: 67 FTYDAAYDASATQTTL-YHEVVFPLVSSVLEGFNGCIFAYGQTGTGKTFTMEGVRGNDEL 125
Query: 156 AGIVYRALRDI-LGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGG 214
GI+ R I L T++ + V V+ LEIY EE+ DLL N
Sbjct: 126 MGIIPRTFEQIWLHINRTEN--------------FQFLVDVSYLEIYMEELRDLLKPN-- 169
Query: 215 GGGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLC 274
S +V+ G N I+ + K +Q K R V T
Sbjct: 170 -------------SKHLEVRERGSGVYVPNLHAINCKSVEDMIKVMQVGNKNRTVGFTNM 216
Query: 275 NDRSSRSHCMVILDVPTVG--------GRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQ 326
N+ SSRSH + ++ + G+L L+D+AGSE + G + K + +KIN
Sbjct: 217 NEHSSRSHAIFMIKIEMCDTETNTIKVGKLNLIDLAGSERQSKTGASAERLK-EASKINL 275
Query: 327 GNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTIST 386
+L V+ ++A HVP+RDSKLT LLQDS SK +MI P ++T++T
Sbjct: 276 ALSSLGNVISALAESSPHVPYRDSKLTRLLQDSL-GGNSKTIMIANIGPSNYNYNETLTT 334
Query: 387 LEYGAKAKCIVRGPHTPVKEEDSSSTVI--LGSRIAAMDEFIMKLQMENKLRE------- 437
L YG++AK I + P+K ED + I + I Q + ++
Sbjct: 335 LRYGSRAKSI---QNQPIKNEDPQDAKLKEYQEEIERLKRLIGPQQQQRSEKQVTAKKQR 391
Query: 438 --KERNEAHKKLMKKEEEIAALRTKVE--TAPASEEEINLKVNE-RTRHLRQELEKKLEE 492
K + E K M +++ + VE + P E + K NE +ELE++ E
Sbjct: 392 VKKPKKETVTKEMSDSLQVSTIEQPVEDDSDPEGAESESDKENEAEVAKSNEELERERVE 451
Query: 493 CQRMTNEFVELERKRM-----------EERILQQQEEVEILRKRLEEIELQ----LCSSS 537
++ + ELE + + E +I +++ VEI ++ EIE+Q L +
Sbjct: 452 NSKLAAKLAELEGQLVRGGKNLLDTYSERQIELEKKLVEIAERKKREIEIQQQLELQEET 511
Query: 538 KQERKDENESKEMEPNGFMRKLLSVY 563
E ++ N S E E RKL Y
Sbjct: 512 TLEIRERNVSLEQEVELKKRKLSKCY 537
>KINH_STRPU (P35978) Kinesin heavy chain
Length = 1031
Score = 127 bits (320), Expect = 8e-29
Identities = 131/543 (24%), Positives = 235/543 (43%), Gaps = 74/543 (13%)
Query: 54 PPEHPIEVIARIRDYPDRKDKPLSVLQASSNSRSIRVKADFGYRDFTLDGVSVSEEEELD 113
P E I+V+ R+R + + + +++ D + EE
Sbjct: 4 PAECNIKVVCRVRPMNATEQNTSHICTKFISEEQVQIGGKLNMFDRIFKPNTTQEE---- 59
Query: 114 LFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTM---FGCSKQAGIVYRALRDILGDG 170
Y K + V G TI YG T SGK+ TM G + GI+ R ++DI
Sbjct: 60 -VYNKAARQIVKDVLDGYNGTIFAYGQTSSGKTFTMEGVMGNPQYMGIIPRIVQDIF--- 115
Query: 171 DTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNA 230
+ D S F ++V+ EIY + I DLL SK+N
Sbjct: 116 ---NHIYQMDESLEF------HIKVSYFEIYMDRIRDLLDV--------------SKTNL 152
Query: 231 SKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVP 290
S + + K AT + ++ I++ + R + T N+ SSRSH + ++ V
Sbjct: 153 SVHEDKNRVPFVKGATERFASSPEEVMDVIEEGKSNRHIAVTNMNEHSSRSHSIFLIQVK 212
Query: 291 T--------VGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGD 342
+ G+L LVD+AGSE + + G G + IN+ AL V+ ++A+G
Sbjct: 213 QENMETKKKLSGKLYLVDLAGSEKVSKTGAEGTVLD-EAKNINKSLSALGNVISALADGK 271
Query: 343 -SHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPH 401
SH+P+RDSK+T +LQ+S ++ +++C SP ++ STL +G +AK I
Sbjct: 272 KSHIPYRDSKMTRILQESL-GGNARTTIVICCSPSSFNESESKSTLMFGQRAKTIKNTVT 330
Query: 402 TPVK----------EEDSSSTVILGSRIAAMDEFIMKLQMENKLREKERNEAHKKLMKKE 451
++ E++ L +++ ++ + + + + KE+ + +++K+
Sbjct: 331 VNMELTAEEWRNRYEKEKEKNGRLKAQLLILENELQRWRAGESVPVKEQGNKNDEILKE- 389
Query: 452 EEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLE-ECQRMTNEFVELERKRMEE 510
++ K T SEEE N E+ + Q EK E + Q E ++ + EE
Sbjct: 390 ----MMKPKQMTVHVSEEEKNKWEEEKVKLYEQLDEKDSEIDNQSRLTEKLKQQMLEQEE 445
Query: 511 RILQQQEEVEILRKRLEEIELQLCSSSKQERKD------------ENESKEMEPNGFMRK 558
+ Q + E+L+ ++ +E + +++K+E K+ + +SKE+E M +
Sbjct: 446 LLSSMQRDYELLQSQMGRLEAE-NAAAKEEAKEVLQALEEMAVNYDEKSKEVEDKNRMNE 504
Query: 559 LLS 561
LS
Sbjct: 505 TLS 507
Score = 32.3 bits (72), Expect = 4.7
Identities = 28/104 (26%), Positives = 49/104 (46%), Gaps = 6/104 (5%)
Query: 448 MKKEEEIAALRTKVETAPASEEEINLKVNE----RTRHLRQELEKKLEECQRMTNEFVEL 503
MK E + + R K+ A +E E ++ +E R Q+ E K++ E E
Sbjct: 587 MKTEVKTMSQRCKILEASNAENETKIRTSEDELDSCRMTIQQHEAKMKSLSENIRE-TEG 645
Query: 504 ERKRMEERILQQQEEVEILRKRLEEIELQLCSSSKQERKDENES 547
+++ +E+ + EE+ LR EEI L K+E +D+ +S
Sbjct: 646 KKRHLEDSLDMLNEEIVKLRAA-EEIRLTDQEDKKREEEDKMQS 688
>KLP2_BOMMO (P46874) Kinesin-like protein KLP2 (Fragment)
Length = 378
Score = 127 bits (319), Expect = 1e-28
Identities = 113/367 (30%), Positives = 178/367 (47%), Gaps = 69/367 (18%)
Query: 59 IEVIARIRDYPDRKD--KPLSVLQASSNSRSI-RVKADFGY-RDFTLDGVSVSEEEELDL 114
I+V R+R R+ K L V++ +N + R+ + FT D ++++
Sbjct: 14 IQVFVRLRPLNQRERDLKSLGVVEVHNNKEVVVRISQQNSITKKFTFDRAFAPYANQVEV 73
Query: 115 FYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ----------AGIVYRALR 164
Y++ V I V G CT+ YG TG+GK+HTM G + AGI+ RAL
Sbjct: 74 -YQEVVSPLIEEVLAGYNCTVFAYGQTGTGKTHTMVGENTGDETTWQKDPLAGIIPRALS 132
Query: 165 DILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFG 224
+ + + V+V+ LE+YNEE++DLL+T
Sbjct: 133 QLFDELRISN--------------TEYTVRVSYLELYNEELFDLLAT------------- 165
Query: 225 WSKSNASKVKLEVMGKKAKNATYISGNEAGKI--SKEIQKV----EKRRIVKSTLCNDRS 278
S+ N+ E + +K N ++G E + KE+ ++ ++R+ V STL N +S
Sbjct: 166 -SEDNSKLRIYEDVTRKGSNI--VNGLEEITVYNKKEVFRIMAQGQERKKVASTLMNAQS 222
Query: 279 SRSHCMVILDVPTVG------------GRLMLVDMAGSENIEQAGQTGFEAKMQTAK--- 323
SRSH + + V G+L LVD+AGSENI +AG AK + A+
Sbjct: 223 SRSHTVFTIVVHMKENSLPEGEELVKIGKLNLVDLAGSENISKAGSDN-PAKRERARECV 281
Query: 324 -INQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHK 382
INQ + L RV+ ++ HVP+R+SKLT +LQ+S ++K +I SP K++ +
Sbjct: 282 NINQSLLTLGRVITALVERHPHVPYRESKLTRILQESL-GGRTKTSIIATISPGHKDLEE 340
Query: 383 TISTLEY 389
T+STLEY
Sbjct: 341 TMSTLEY 347
>KI22_STRPU (P46872) Kinesin-II 85 kDa subunit (KRP-85/95 85 kDa
subunit)
Length = 699
Score = 127 bits (319), Expect = 1e-28
Identities = 140/487 (28%), Positives = 219/487 (44%), Gaps = 71/487 (14%)
Query: 97 RDFTLDGVSVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQA 156
+ FT D V ++ D+ Y + ++ + G TI YG TG+GK+ TM G Q
Sbjct: 56 KSFTFDTVFAPGAKQTDV-YNQTARPIVDAIIEGYNGTIFAYGQTGTGKTFTMEGVRSQP 114
Query: 157 ---GIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNG 213
GI+ + I G + E VR V+V+ LEIYNEE+ DLL
Sbjct: 115 ELRGIIPNSFAHIFGHIAKEQEN------------VRFLVRVSYLEIYNEEVKDLL---- 158
Query: 214 GGGGGGGFGFGWSKSNASKVKLEV-MGKKAKNATYISGNEAGKISKEIQKVEKRRIVKST 272
G + + +VK +G K+ + N A + + + K R V +T
Sbjct: 159 ----------GKDQQHRLEVKERPDVGVYVKDLSAFVVNNADDMDRIMTLGNKNRSVGAT 208
Query: 273 LCNDRSSRSHCM--VILDVPTVG---------GRLMLVDMAGSENIEQAGQTGFEAKMQT 321
N+ SSRSH + + L+ +G G+L +VD+AGSE + G TG K T
Sbjct: 209 NMNESSSRSHAIFTITLERSDMGLDKEQHVRVGKLHMVDLAGSERQTKTGATGQRLKEAT 268
Query: 322 AKINQGNIALKRVVESIANGDS-HVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEI 380
KIN L V+ S+ +G S H+P+R+SKLT LLQDS + ++CA+ P E
Sbjct: 269 -KINLSLSTLGNVISSLVDGKSTHIPYRNSKLTRLLQDSLGGNAK---TVMCANIGPAEY 324
Query: 381 H--KTISTLEYGAKAKCIV------RGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQME 432
+ +TISTL Y +AK I P + E L +I+ E + E
Sbjct: 325 NYDETISTLRYANRAKNIKNKAKINEDPKDALLREFQKEIEELKKQISESGEGLDD-DEE 383
Query: 433 NKLREKERNEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEE 492
+ E EA + +KK+ + K + +P + K++E + L E +K + E
Sbjct: 384 SGSEESGDEEAGEGGVKKKRK--GKNPKRKLSPEIMAAMQKKIDEEKKAL--EEKKDMVE 439
Query: 493 CQRMTNEFVELERKRMEERILQQQEEVEILRKRLEEIELQ--------LCSSSKQERKDE 544
R T V E +R E + + Q++ +IL ++L I+ + L S +QE+ E
Sbjct: 440 EDRNT---VHRELQRRESELHKAQDDQKILNEKLNAIQKKLIVGGVDLLAKSEEQEQLLE 496
Query: 545 NESKEME 551
+ EM+
Sbjct: 497 QSALEMK 503
>KF17_HUMAN (Q9P2E2) Kinesin-like protein KIF17 (KIF3-related motor
protein)
Length = 1029
Score = 125 bits (315), Expect = 3e-28
Identities = 136/460 (29%), Positives = 190/460 (40%), Gaps = 91/460 (19%)
Query: 92 ADFGYRDFTLDGVSVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFG 151
AD + FT DG + + + Y + + GV G TI YG TGSGKS TM G
Sbjct: 45 ADEPPKQFTFDG-AYHVDHVTEQIYNEIAYPLVEGVTEGYNGTIFAYGQTGSGKSFTMQG 103
Query: 152 CS---KQAGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDL 208
Q GI+ RA + S + V+ + LEIYNE++ DL
Sbjct: 104 LPDPPSQRGIIPRAFEHVF-------------ESVQCAENTKFLVRASYLEIYNEDVRDL 150
Query: 209 LSTNGGGGGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISG------NEAGKISKEIQK 262
L + +K KLE+ K Y+ G + + ++
Sbjct: 151 LGAD------------------TKQKLELKEHPEKGV-YVKGLSMHTVHSVAQCEHIMET 191
Query: 263 VEKRRIVKSTLCNDRSSRSHCMVILDVPTVG-----------GRLMLVDMAGSENIEQAG 311
K R V TL N SSRSH + + + G+L LVD+AGSE + G
Sbjct: 192 GWKNRSVGYTLMNKDSSRSHSIFTISIEMSAVDERGKDHLRAGKLNLVDLAGSERQSKTG 251
Query: 312 QTGFEAKMQTAKINQGNIALKRVVESIANGD-SHVPFRDSKLTMLLQDSFEDDKSKILMI 370
TG K T KIN AL V+ ++ +G HVP+RDSKLT LLQDS +K LM+
Sbjct: 252 ATGERLKEAT-KINLSLSALGNVISALVDGRCKHVPYRDSKLTRLLQDSL-GGNTKTLMV 309
Query: 371 LCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKEEDS--------------------- 409
C SP +T+STL Y +AK I P +D+
Sbjct: 310 ACLSPADNNYDETLSTLRYANRAKNIRNKPRINEDPKDALLREYQEEIKKLKAILTQQMS 369
Query: 410 -SSTVILGSRIAAMDEFIMKLQMENKLRE----KERNEAHKKLMKKEEEIAALRTKVETA 464
SS L SR D +Q+E KL + EA K+L+++E E R K +
Sbjct: 370 PSSLSALLSRQVPPD----PVQVEEKLLPQPVIQHDMEAEKQLIREEYEERLARLKADY- 424
Query: 465 PASEEEINLKVNERTRHLRQELEKK---LEECQRMTNEFV 501
+E+E ++ E +R + + LEE R E V
Sbjct: 425 -KAEQESRARLEEDITAMRNSYDVRLSTLEENLRKETEAV 463
>KF12_MOUSE (Q9D2Z8) Kinesin-like protein KIF12
Length = 642
Score = 125 bits (313), Expect = 5e-28
Identities = 137/526 (26%), Positives = 221/526 (41%), Gaps = 107/526 (20%)
Query: 40 PHPSPHPNFNNKDSPPEHPIEVIARIRDYP--DRKDKPLSVLQASSNSRSIRVKADFGYR 97
P P N E PI+V+ R+R + + S L S +R+++V D +R
Sbjct: 7 PDGDPARNLEQGPEGSETPIQVVLRVRPMSTVELRRGEQSALHCSG-TRTLQVSPDVAFR 65
Query: 98 -DFTLDGVSVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQ- 155
LDG E+ K+ E + G CT+ +G TGSGK++T+ G Q
Sbjct: 66 FGAVLDGARTQEDVFRACGVKRLGELALRGFS----CTVFTFGQTGSGKTYTLTGPPPQG 121
Query: 156 ---------AGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIY 206
AGI+ R +L + L ++ + LEIYNE+++
Sbjct: 122 EGVPVPPSLAGIMQRTFTWLL--------------DRVQHLDSPVTLRASYLEIYNEQVW 167
Query: 207 DLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISGNEAGKISKEIQKVEKR 266
DLLS G W+K+ G + + + + +Q R
Sbjct: 168 DLLSL----GSPRPLPVRWTKAR---------GFYVEQLRVVEFGSLEALMELLQMGLSR 214
Query: 267 RIVKSTLCNDRSSRSHCMVILDV---------------PTVGGRLMLVDMAGSENIEQAG 311
R S N SSRSH ++ L + P VGG+L VD+AGSE + G
Sbjct: 215 RRSSSHTLNQASSRSHALLTLHISRPTSQQVPPVDLGEPPVGGKLCFVDLAGSEKVAATG 274
Query: 312 QTGFEAKMQTAKINQGNIALKRVVESIANGD---SHVPFRDSKLTMLLQDSFEDDKSKIL 368
G + ++ IN+ +AL + + + +H+PFRDSKLT LL DS + L
Sbjct: 275 SQG-QLMLEANSINRSLLALGHCISLLLDPQRKQNHIPFRDSKLTKLLADSL-GGRGVTL 332
Query: 369 MILCASPDPKEIHKTISTLEYGAKAKCIV---RGPHTP-VKEEDSSSTVILGSRIAAMDE 424
M+ C SP + + +T+STL Y ++A+ I +GP +P VK ++
Sbjct: 333 MVACVSPSAQCLPETLSTLRYASRAQRITTRPQGPKSPGVKPPQQ------------VEN 380
Query: 425 FIMKLQMENKLREKERNEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQ 484
+++LQ EN+ + ++ H TAP + ++ R+L
Sbjct: 381 ELLRLQEENRHLRFQLDQMH-----------------TTAPGAH---GARMAWAQRNLYG 420
Query: 485 ELEKKLEECQRMTNEFVEL----ERKRMEERILQQQEEVEILRKRL 526
L++ + E +R+ E +L + R E+R+L QQ V L +RL
Sbjct: 421 MLQEFMLENERLRKEMRQLRSSRDLARAEQRVLAQQ--VHDLERRL 464
>K13B_HUMAN (Q9NQT8) Kinesin-like protein KIF13B (Kinesin-like
protein GAKIN)
Length = 1826
Score = 124 bits (312), Expect = 7e-28
Identities = 116/397 (29%), Positives = 179/397 (44%), Gaps = 60/397 (15%)
Query: 113 DLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAGIVYRALRDILGDGDT 172
D+ +K E+ + G I YG TGSGKS+TM G + Q G++ R + T
Sbjct: 77 DIVFKCLGENILQNAFDGYNACIFAYGQTGSGKSYTMMGTADQPGLIPRLCSGLF--ERT 134
Query: 173 DSEGSDGDSSKGFCLGVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASK 232
E ++ S K V+V+ +EIYNE++ DLL G S+
Sbjct: 135 QKEENEEQSFK---------VEVSYMEIYNEKVRDLLDPKG------------SRQTLKV 173
Query: 233 VKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVP-- 290
+ V+G + ++ I + + K R V +T N+ SSRSH ++ + +
Sbjct: 174 REHSVLGPYVDGLSKLAATSYKDIESLMSEGNKSRTVAATNMNEESSRSHAVLKITLTHT 233
Query: 291 -------TVG---GRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIA- 339
T G G+L LVD+AGSE + G G K + + IN+ L V+ ++A
Sbjct: 234 LYDAKSGTSGEKVGKLSLVDLAGSERATKTGAAGDRLK-EGSNINESLTTLGLVISALAD 292
Query: 340 -----NGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAK 394
N + VP+RDS LT LL+DS SK M+ SP +T+STL Y +AK
Sbjct: 293 QSAGKNKNKFVPYRDSVLTWLLKDSL-GGNSKTAMVATVSPAADNYDETLSTLRYADRAK 351
Query: 395 CIVRGPHTPVKEEDSSSTVI--LGSRIAAMDEFIMKLQMENKLREKERNEAHKKL----- 447
IV + V ED ++ +I L + + E + K + K+R E +KL
Sbjct: 352 HIV---NNAVVNEDPNARIIRDLREEVEKLREQLTKAEAMKSPELKDRLEESEKLIQEMT 408
Query: 448 ------MKKEEEIAALRTK-VETAPASEEEINLKVNE 477
++K EEIA R K +E+ S + +KV +
Sbjct: 409 VTWEEKLRKTEEIAQERQKQLESLGISLQSSGIKVGD 445
Score = 32.3 bits (72), Expect = 4.7
Identities = 31/129 (24%), Positives = 57/129 (44%), Gaps = 12/129 (9%)
Query: 416 GSRIAAMDEFIMKLQMENKLREKERNEAHKKLMKKEEEIAALRTKVETAPASEEEINLKV 475
G RI + +L + K ++ ER + + K E + +++ S E++ +V
Sbjct: 529 GDRILWGNNHFFRLNLPKKKKKAEREDEDQDPSMKNENSSE---QLDVDGDSSSEVSSEV 585
Query: 476 NERTRHLRQELEKKL----EECQRMTN----EFVELERKRMEERILQQQEEVEILRKRLE 527
N + + E+ K + Q + N + E +R +E + L + E+E LR+RL
Sbjct: 586 NFNYEYAQMEVTMKALGSNDPMQSILNSLEQQHEEEKRSALERQRLMYEHELEQLRRRLS 645
Query: 528 EIELQLCSS 536
E Q C S
Sbjct: 646 P-EKQNCRS 653
>KF2C_CRIGR (P70096) Kinesin-like protein KIF2C (Mitotic
centromere-associated kinesin) (MCAK) (Kinesin-like
protein 6)
Length = 718
Score = 124 bits (311), Expect = 9e-28
Identities = 119/471 (25%), Positives = 192/471 (40%), Gaps = 81/471 (17%)
Query: 98 DFTLDGVSVSEEEELDLFYKKFVESRINGVKLGDKCTIMMYGPTGSGKSHTMFGCSKQAG 157
DF D + +E + Y+ + + G K T YG TGSGK+HTM G
Sbjct: 306 DFAFDETASNE-----VVYRFTARPLVQTIFEGGKATCFAYGQTGSGKTHTMGG------ 354
Query: 158 IVYRALRDILGDGDTDSEGSDGDSSKGFCL--------GVRTFVQVTVLEIYNEEIYDLL 209
D+ G S+G +S+ L + V VT EIYN +++DLL
Sbjct: 355 -------DLSGKSQNTSKGIYAMASRDVFLLKSQPRYRNLNLEVYVTFFEIYNGKVFDLL 407
Query: 210 STNGGGGGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISG------NEAGKISKEIQKV 263
+ K KL V+ + +K + G N A + K +
Sbjct: 408 N--------------------KKAKLRVL-EDSKQQVQVVGLQEYLVNCADDVIKMLNMG 446
Query: 264 EKRRIVKSTLCNDRSSRSHCMVILDVPTVG---GRLMLVDMAGSENIEQAGQTGFEAKMQ 320
R T N SSRSH + + G G+ LVD+AG+E + +M+
Sbjct: 447 SACRTSGQTFANSNSSRSHACFQILLRAKGRLHGKFSLVDLAGNERGADTSSADRQTRME 506
Query: 321 TAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEI 380
A+IN+ +ALK + ++ +H PFR+SKLT +L+DSF + S+ MI SP
Sbjct: 507 GAEINKSLLALKECIRALGQNKAHTPFRESKLTQVLRDSFIGENSRTCMIAMISPGISSC 566
Query: 381 HKTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQMENKLREKER 440
T++TL Y + K + PH+ + E +QME + E
Sbjct: 567 EYTLNTLRYADRVKEL--SPHSGLSGE-------------------QPIQMETEEMEASS 605
Query: 441 NEAHKKL-MKKEEEIAALRTKVETAPASEEEINLKVNERTRHLRQELEKKLEECQRMTNE 499
N + K+EEE+++ + A + E+ + E R + Q+ LE +
Sbjct: 606 NGTSLAVNFKEEEELSSQMSSFNEAMSQIRELEERAMEELREIIQQGPGWLELSEMTDQP 665
Query: 500 FVELER--KRMEERILQQQEEVEILRKRLEEIELQLCSSSKQERKDENESK 548
+LE + E + QQ + LR+ ++ + + + +Q K N K
Sbjct: 666 DYDLETFVNKAESALTQQTKHFSALREVIKALRVAM-QLEEQASKQMNSKK 715
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.312 0.131 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 92,388,547
Number of Sequences: 164201
Number of extensions: 4411514
Number of successful extensions: 40479
Number of sequences better than 10.0: 1335
Number of HSP's better than 10.0 without gapping: 311
Number of HSP's successfully gapped in prelim test: 1066
Number of HSP's that attempted gapping in prelim test: 30947
Number of HSP's gapped (non-prelim): 5786
length of query: 754
length of database: 59,974,054
effective HSP length: 118
effective length of query: 636
effective length of database: 40,598,336
effective search space: 25820541696
effective search space used: 25820541696
T: 11
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 70 (31.6 bits)
Medicago: description of AC144481.6


