

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144481.4 + phase: 0
(152 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
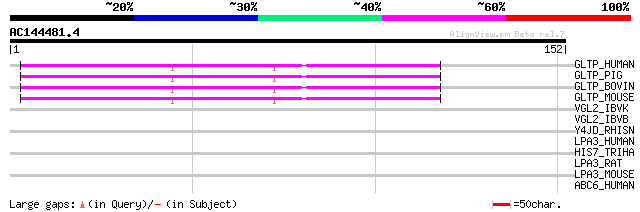
Score E
Sequences producing significant alignments: (bits) Value
GLTP_HUMAN (Q9NZD2) Glycolipid transfer protein (GLTP) 68 9e-12
GLTP_PIG (P68266) Glycolipid transfer protein (GLTP) 66 3e-11
GLTP_BOVIN (P68265) Glycolipid transfer protein (GLTP) 66 3e-11
GLTP_MOUSE (Q9JL62) Glycolipid transfer protein (GLTP) 63 3e-10
VGL2_IBVK (P12650) E2 glycoprotein precursor (Spike glycoprotein... 30 2.1
VGL2_IBVB (P11223) E2 glycoprotein precursor (Spike glycoprotein... 30 2.1
Y4JD_RHISN (P55504) Hypothetical 56.7 kDa protein y4jD 29 4.6
LPA3_HUMAN (O75145) Liprin-alpha 3 (Protein tyrosine phosphatase... 29 4.6
HIS7_TRIHA (P34041) Imidazoleglycerol-phosphate dehydratase (EC ... 28 6.0
LPA3_RAT (Q91Z79) Liprin-alpha 3 (Protein tyrosine phosphatase r... 28 7.8
LPA3_MOUSE (P60469) Liprin-alpha 3 (Protein tyrosine phosphatase... 28 7.8
ABC6_HUMAN (Q9NP58) ATP-binding cassette, sub-family B, member 6... 28 7.8
>GLTP_HUMAN (Q9NZD2) Glycolipid transfer protein (GLTP)
Length = 208
Score = 67.8 bits (164), Expect = 9e-12
Identities = 37/122 (30%), Positives = 66/122 (53%), Gaps = 8/122 (6%)
Query: 4 VKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAK-ASSSCTTGLLWLTRAMDFL 62
+K+DI GNI+++++ Y +NPAKF L ++++VE E A+ T L+WL R + F+
Sbjct: 44 IKADISGNITKIKAVYDTNPAKFRTLQNILEVEKEMYGAEWPKVGATLALMWLKRGLRFI 103
Query: 63 VAVFRNLIE------HADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFME 116
+++ + H + A T +Y LKK+HGW+ A+ AP + F++
Sbjct: 104 QVFLQSICDGERDENHPNLIRVNA-TKAYEMALKKYHGWIVQKIFQAALYAAPYKSDFLK 162
Query: 117 VV 118
+
Sbjct: 163 AL 164
>GLTP_PIG (P68266) Glycolipid transfer protein (GLTP)
Length = 208
Score = 66.2 bits (160), Expect = 3e-11
Identities = 36/122 (29%), Positives = 65/122 (52%), Gaps = 8/122 (6%)
Query: 4 VKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAK-ASSSCTTGLLWLTRAMDFL 62
+K+DI GNI+++++ Y +NP KF L ++++VE E A+ T L+WL R + F+
Sbjct: 44 IKADISGNITKIKAVYDTNPTKFRTLQNILEVEKEMYGAEWPKVGATLALMWLKRGLRFI 103
Query: 63 VAVFRNLIE------HADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFME 116
+++ + H + A T +Y LKK+HGW+ A+ AP + F++
Sbjct: 104 QVFLQSICDGERDENHPNLIRVNA-TKAYEMALKKYHGWIVQKIFQAALYAAPYKSDFLK 162
Query: 117 VV 118
+
Sbjct: 163 AL 164
>GLTP_BOVIN (P68265) Glycolipid transfer protein (GLTP)
Length = 208
Score = 66.2 bits (160), Expect = 3e-11
Identities = 36/122 (29%), Positives = 65/122 (52%), Gaps = 8/122 (6%)
Query: 4 VKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAK-ASSSCTTGLLWLTRAMDFL 62
+K+DI GNI+++++ Y +NP KF L ++++VE E A+ T L+WL R + F+
Sbjct: 44 IKADISGNITKIKAVYDTNPTKFRTLQNILEVEKEMYGAEWPKVGATLALMWLKRGLRFI 103
Query: 63 VAVFRNLIE------HADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFME 116
+++ + H + A T +Y LKK+HGW+ A+ AP + F++
Sbjct: 104 QVFLQSICDGERDENHPNLIRVNA-TKAYEMALKKYHGWIVQKIFQAALYAAPYKSDFLK 162
Query: 117 VV 118
+
Sbjct: 163 AL 164
>GLTP_MOUSE (Q9JL62) Glycolipid transfer protein (GLTP)
Length = 208
Score = 62.8 bits (151), Expect = 3e-10
Identities = 36/122 (29%), Positives = 64/122 (51%), Gaps = 8/122 (6%)
Query: 4 VKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAK-ASSSCTTGLLWLTRAMDFL 62
+K+DI GNI+++++ Y ++PAKF L ++++VE A+ T LLWL R + F+
Sbjct: 44 IKADISGNITKIKAVYDTDPAKFKTLQNILEVEKGMYGAEWPKVGATLALLWLKRGLRFI 103
Query: 63 VAVFRNLIE------HADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFME 116
+++ + H + A +Y LKK+HGWL A+ AP + F++
Sbjct: 104 QVFLQSICDGERDENHPNLIRVNA-NKAYEMALKKYHGWLVQKIFKAALYAAPYKSDFLK 162
Query: 117 VV 118
+
Sbjct: 163 AL 164
>VGL2_IBVK (P12650) E2 glycoprotein precursor (Spike glycoprotein)
(Peplomer protein) [Contains: Spike protein S1; Spike
protein S2]
Length = 1162
Score = 30.0 bits (66), Expect = 2.1
Identities = 16/53 (30%), Positives = 24/53 (45%)
Query: 30 YSLVQVEVETKTAKASSSCTTGLLWLTRAMDFLVAVFRNLIEHADWSMSQACT 82
Y++V + E+ A +SS CT G + R ++ WS SQ CT
Sbjct: 47 YAVVNISSESNNAGSSSGCTVGTIHGGRVVNASSIAMTAPSSGMAWSSSQFCT 99
>VGL2_IBVB (P11223) E2 glycoprotein precursor (Spike glycoprotein)
(Peplomer protein) [Contains: Spike protein S1; Spike
protein S2]
Length = 1162
Score = 30.0 bits (66), Expect = 2.1
Identities = 16/53 (30%), Positives = 24/53 (45%)
Query: 30 YSLVQVEVETKTAKASSSCTTGLLWLTRAMDFLVAVFRNLIEHADWSMSQACT 82
Y++V + E A +SS CT G++ R ++ WS SQ CT
Sbjct: 47 YAVVNISSEFNNAGSSSGCTVGIIHGGRVVNASSIAMTAPSSGMAWSSSQFCT 99
>Y4JD_RHISN (P55504) Hypothetical 56.7 kDa protein y4jD
Length = 511
Score = 28.9 bits (63), Expect = 4.6
Identities = 15/33 (45%), Positives = 21/33 (63%)
Query: 86 YKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
YKTL + G SS + +A LA R+KF++VV
Sbjct: 303 YKTLARHRGKSNSSPLRLAFCLAHARRKFVDVV 335
>LPA3_HUMAN (O75145) Liprin-alpha 3 (Protein tyrosine phosphatase
receptor type f polypeptide-interacting protein alpha 3)
(PTPRF-interacting protein alpha 3)
Length = 1194
Score = 28.9 bits (63), Expect = 4.6
Identities = 36/128 (28%), Positives = 54/128 (42%), Gaps = 16/128 (12%)
Query: 20 LSNPAKFNCLYSLVQ--VEVETKTAKASSSCTTGLLWLT-RAMDFLVAVFRNLIEHADWS 76
+SNP L +Q V + + +A ASS +TG +W+T M+ L A + + W
Sbjct: 885 ISNPLHRLKLRLAIQEMVSLTSPSAPASSRTSTGNVWMTHEEMESLTATTKPETKEISWE 944
Query: 77 MSQACTDSYYKTLKKWHG--WLASSTVTVAMMLAPDRKKFMEVVISMLTLSNFVLAFLL- 133
A D + +W G WL S + L R FME ++ L + L
Sbjct: 945 QILAYGDMNH----EWVGNDWLPS------LGLPQYRSYFMESLVDARMLDHLNKKELRG 994
Query: 134 SLKRITSF 141
LK + SF
Sbjct: 995 QLKMVDSF 1002
>HIS7_TRIHA (P34041) Imidazoleglycerol-phosphate dehydratase (EC
4.2.1.19) (IGPD)
Length = 208
Score = 28.5 bits (62), Expect = 6.0
Identities = 13/45 (28%), Positives = 24/45 (52%)
Query: 37 VETKTAKASSSCTTGLLWLTRAMDFLVAVFRNLIEHADWSMSQAC 81
+E +A AS + + ++ + + FL + L +HA WSM+ C
Sbjct: 41 LEASSAHASQTSKSQVISINTGIGFLDHMLHALAKHAGWSMALNC 85
>LPA3_RAT (Q91Z79) Liprin-alpha 3 (Protein tyrosine phosphatase
receptor type f polypeptide-interacting protein alpha 3)
(PTPRF-interacting protein alpha 3)
Length = 1192
Score = 28.1 bits (61), Expect = 7.8
Identities = 36/128 (28%), Positives = 53/128 (41%), Gaps = 16/128 (12%)
Query: 20 LSNPAKFNCLYSLVQ--VEVETKTAKASSSCTTGLLWLT-RAMDFLVAVFRNLIEHADWS 76
+SNP L +Q V + + +A ASS TG +W+T M+ L A + + W
Sbjct: 883 ISNPLHRLKLRLAIQEMVSLTSPSAPASSRTPTGNVWMTHEEMESLTAATKPETKEISWE 942
Query: 77 MSQACTDSYYKTLKKWHG--WLASSTVTVAMMLAPDRKKFMEVVISMLTLSNFVLAFLL- 133
A D + +W G WL S + L R FME ++ L + L
Sbjct: 943 QILAYGDMNH----EWVGNDWLPS------LGLPQYRSYFMESLVDARMLDHLNKKELRE 992
Query: 134 SLKRITSF 141
LK + SF
Sbjct: 993 QLKMVDSF 1000
>LPA3_MOUSE (P60469) Liprin-alpha 3 (Protein tyrosine phosphatase
receptor type f polypeptide-interacting protein alpha 3)
(PTPRF-interacting protein alpha 3)
Length = 1043
Score = 28.1 bits (61), Expect = 7.8
Identities = 36/128 (28%), Positives = 53/128 (41%), Gaps = 16/128 (12%)
Query: 20 LSNPAKFNCLYSLVQ--VEVETKTAKASSSCTTGLLWLT-RAMDFLVAVFRNLIEHADWS 76
+SNP L +Q V + + +A ASS TG +W+T M+ L A + + W
Sbjct: 734 ISNPLHRLKLRLAIQEMVSLTSPSAPASSRTPTGNVWMTHEEMESLTAATKPETKEISWE 793
Query: 77 MSQACTDSYYKTLKKWHG--WLASSTVTVAMMLAPDRKKFMEVVISMLTLSNFVLAFLL- 133
A D + +W G WL S + L R FME ++ L + L
Sbjct: 794 QILAYGDMNH----EWVGNDWLPS------LGLPQYRSYFMESLVDARMLDHLNKKELRG 843
Query: 134 SLKRITSF 141
LK + SF
Sbjct: 844 QLKMVDSF 851
>ABC6_HUMAN (Q9NP58) ATP-binding cassette, sub-family B, member 6,
mitochondrial precursor (Mitochondrial ABC transporter
3) (Mt-ABC transporter 3) (ABC transporter umat)
Length = 842
Score = 28.1 bits (61), Expect = 7.8
Identities = 28/97 (28%), Positives = 44/97 (44%), Gaps = 11/97 (11%)
Query: 51 GLLWLTRAMDFLVAVF-RNLI----EHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAM 105
GL+ L RA++ LV +F RN++ E A W+ S A T + Y LK G ST V+
Sbjct: 270 GLMGLERALNVLVPIFYRNIVNLLTEKAPWN-SLAWTVTSYVFLKFLQGGGTGSTGFVSN 328
Query: 106 MLAPDRKKFMEVVISMLTLSNFVLAFLLSLKRITSFW 142
+ + F+ + + T L L ++ W
Sbjct: 329 L-----RTFLWIRVQQFTSRRVELLIFSHLHELSLRW 360
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.327 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,089,982
Number of Sequences: 164201
Number of extensions: 457987
Number of successful extensions: 1576
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 6
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 1566
Number of HSP's gapped (non-prelim): 12
length of query: 152
length of database: 59,974,054
effective HSP length: 100
effective length of query: 52
effective length of database: 43,553,954
effective search space: 2264805608
effective search space used: 2264805608
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144481.4


