

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
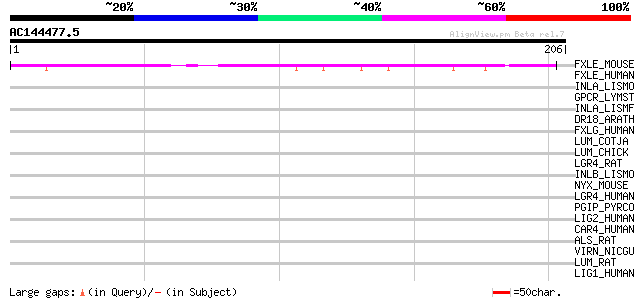
Query= AC144477.5 - phase: 1 /pseudo
(206 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
FXLE_MOUSE (Q8BID8) F-box/LRR-repeat protein 14 (F-box and leuci... 50 5e-06
FXLE_HUMAN (Q8N1E6) F-box/LRR-repeat protein 14 (F-box and leuci... 40 0.003
INLA_LISMO (P25146) Internalin A precursor 40 0.005
GPCR_LYMST (P46023) G-protein coupled receptor GRL101 precursor 40 0.005
INLA_LISMF (Q723K6) Internalin A precursor 39 0.008
DR18_ARATH (O64789) Putative disease resistance protein At1g61310 38 0.019
FXLG_HUMAN (Q8N461) F-box/LRR-repeat protein 16 (F-box and leuci... 37 0.042
LUM_COTJA (Q9DE67) Lumican precursor (Keratan sulfate proteoglyc... 36 0.055
LUM_CHICK (P51890) Lumican precursor (Keratan sulfate proteoglyc... 36 0.055
LGR4_RAT (Q9Z2H4) Leucine-rich repeat-containing G protein-coupl... 36 0.055
INLB_LISMO (P25147) Internalin B precursor 36 0.055
NYX_MOUSE (P83503) Nyctalopin precursor 36 0.072
LGR4_HUMAN (Q9BXB1) Leucine-rich repeat-containing G protein-cou... 36 0.072
PGIP_PYRCO (Q05091) Polygalacturonase inhibitor precursor (Polyg... 35 0.094
LIG2_HUMAN (O94898) Leucine-rich repeats and immunoglobulin-like... 35 0.12
CAR4_HUMAN (Q9Y239) Caspase recruitment domain protein 4 (Nod1 p... 35 0.12
ALS_RAT (P35859) Insulin-like growth factor binding protein comp... 35 0.16
VIRN_NICGU (Q40392) TMV resistance protein N 34 0.27
LUM_RAT (P51886) Lumican precursor (Keratan sulfate proteoglycan... 34 0.27
LIG1_HUMAN (Q96JA1) Leucine-rich repeats and immunoglobulin-like... 34 0.27
>FXLE_MOUSE (Q8BID8) F-box/LRR-repeat protein 14 (F-box and
leucine-rich repeat protein 14)
Length = 400
Score = 49.7 bits (117), Expect = 5e-06
Identities = 63/213 (29%), Positives = 105/213 (48%), Gaps = 23/213 (10%)
Query: 1 LQKLSTLNVEGC----SITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFL 56
LQ+L +LN+ C + ++ AA CL L + L D L+H+ L
Sbjct: 168 LQRLKSLNLRSCRHLSDVGIGHLAGMTRSAAEGCLGLEQLTLQDCQKLTDLSLKHISRGL 227
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTE-VGNSGIRYIS 115
L+ +L+L+ F I D GL++L+ + L+SL L + + ++GI +++
Sbjct: 228 TGLR-----LLNLS-------FCGGISDAGLLHLSHMGSLRSLNLRSCDNISDTGIMHLA 275
Query: 116 -GLNKLEDLNLSFTS-VTDNGLKRLL-GLTNLKSLNLDARQITDAGLANLT-SLSGLITL 171
G +L L++SF V D L + GL LKSL+L + I+D G+ + + GL TL
Sbjct: 276 MGSLRLSGLDVSFCDKVGDQSLAYIAQGLDGLKSLSLCSCHISDDGINRMVRQMHGLRTL 335
Query: 172 DLFG-ARITDSGTTYLRCMYIYQLQAIVVFLCS 203
++ RITD G L ++ QL I ++ C+
Sbjct: 336 NIGQCVRITDKGLE-LIAEHLSQLTGIDLYGCT 367
Score = 39.7 bits (91), Expect = 0.005
Identities = 51/188 (27%), Positives = 75/188 (39%), Gaps = 49/188 (26%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQV 58
L L LN+ C I+ A ++S + +L LNL C +SD G ++
Sbjct: 227 LTGLRLLNLSFCGGISDAGLLHLSHMGSLRSLNLRSCDNISDTGIMHLAMGS-------- 278
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLT-GLTLLKSLVLSDTEVGNSGI-RYISG 116
L L+ L F K+GD+ L + GL LKSL L + + GI R +
Sbjct: 279 --------LRLSGLDVS--FCDKVGDQSLAYIAQGLDGLKSLSLCSCHISDDGINRMVRQ 328
Query: 117 LNKLEDLNLS---------------------------FTSVTDNGLKRLLGLTNLKSLNL 149
++ L LN+ T +T GL+R+ L LK LNL
Sbjct: 329 MHGLRTLNIGQCVRITDKGLELIAEHLSQLTGIDLYGCTRITKRGLERITQLPCLKVLNL 388
Query: 150 DARQITDA 157
Q+TD+
Sbjct: 389 GLWQMTDS 396
>FXLE_HUMAN (Q8N1E6) F-box/LRR-repeat protein 14 (F-box and
leucine-rich repeat protein 14)
Length = 418
Score = 40.4 bits (93), Expect = 0.003
Identities = 55/207 (26%), Positives = 81/207 (38%), Gaps = 49/207 (23%)
Query: 1 LQKLSTLNVEGCS-ITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQV 58
L L LN+ C I+ A ++S + +L LNL C +SD G ++
Sbjct: 227 LTGLRLLNLSFCGGISDAGLLHLSHMGSLRSLNLRSCDNISDTGIMHLAMGS-------- 278
Query: 59 LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLT-GLTLLKSLVLSDTEVGNSGI-RYISG 116
L L+ L F K+GD+ L + GL LKSL L + + GI R +
Sbjct: 279 --------LRLSGLDVS--FCDKVGDQSLAYIAQGLDGLKSLSLCSCHISDDGINRMVRQ 328
Query: 117 LNKLEDLNLS---------------------------FTSVTDNGLKRLLGLTNLKSLNL 149
++ L LN+ T +T GL+R+ L LK LNL
Sbjct: 329 MHGLRTLNIGQCVRITDKGLELIAEHLSQLTGIDLYGCTRITKRGLERITQLPCLKVLNL 388
Query: 150 DARQITDAGLANLTSLSGLITLDLFGA 176
Q+TD+ S L T+ G+
Sbjct: 389 GLWQMTDSEKEARGDFSPLFTVRTRGS 415
>INLA_LISMO (P25146) Internalin A precursor
Length = 800
Score = 39.7 bits (91), Expect = 0.005
Identities = 34/95 (35%), Positives = 53/95 (55%), Gaps = 15/95 (15%)
Query: 87 LVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKS 146
L NLTGLTL + + + N ++ LN+LE LS +++D + L GLT+L+
Sbjct: 141 LTNLTGLTLFNNQITDIDPLKN-----LTNLNRLE---LSSNTISD--ISALSGLTSLQQ 190
Query: 147 LNLDARQITD-AGLANLTSLSGLITLDLFGARITD 180
L+ Q+TD LANLT+L LD+ +++D
Sbjct: 191 LSF-GNQVTDLKPLANLTTLE---RLDISSNKVSD 221
Score = 38.9 bits (89), Expect = 0.008
Identities = 36/101 (35%), Positives = 52/101 (50%), Gaps = 10/101 (9%)
Query: 90 LTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNL 149
L LT L+ L +S +V S I ++ L LE L + ++D + L LTNL L+L
Sbjct: 203 LANLTTLERLDISSNKV--SDISVLAKLTNLESLIATNNQISD--ITPLGILTNLDELSL 258
Query: 150 DARQITDAGLANLTSLSGLITLDLFGARITD----SGTTYL 186
+ Q+ D G L SL+ L LDL +I++ SG T L
Sbjct: 259 NGNQLKDIG--TLASLTNLTDLDLANNQISNLAPLSGLTKL 297
Score = 32.7 bits (73), Expect = 0.61
Identities = 35/110 (31%), Positives = 55/110 (49%), Gaps = 15/110 (13%)
Query: 86 GLVNLTGLTLLKSLV---LSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLT 142
G+ ++ G+ L +L S+ ++ + I + L KL D+ ++ + D + L LT
Sbjct: 87 GIKSIDGVEYLNNLTQINFSNNQL--TDITPLKNLTKLVDILMNNNQIAD--ITPLANLT 142
Query: 143 NLKSLNLDARQITDAG-LANLTSLSGLITLDLFGARITD----SGTTYLR 187
NL L L QITD L NLT+L+ L+L I+D SG T L+
Sbjct: 143 NLTGLTLFNNQITDIDPLKNLTNLN---RLELSSNTISDISALSGLTSLQ 189
>GPCR_LYMST (P46023) G-protein coupled receptor GRL101 precursor
Length = 1115
Score = 39.7 bits (91), Expect = 0.005
Identities = 34/105 (32%), Positives = 49/105 (46%), Gaps = 8/105 (7%)
Query: 82 IGDEGLVNLTGLTLLKS-------LVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNG 134
IGD L+NLT T + L LS + I + KL LNL+ ++T
Sbjct: 565 IGDN-LLNLTSTTFSATYYDKVTYLDLSRNHLTEIPIYSFQNMWKLTHLNLADNNITSLK 623
Query: 135 LKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARIT 179
LLGL+NLK L+++ +I +S+ L LDL R+T
Sbjct: 624 NGSLLGLSNLKQLHINGNKIETIEEDTFSSMIHLTVLDLSNQRLT 668
Score = 30.4 bits (67), Expect = 3.0
Identities = 25/94 (26%), Positives = 40/94 (41%)
Query: 89 NLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLN 148
+L GL+ LK L ++ ++ S + L L+LS +T GL + LN
Sbjct: 626 SLLGLSNLKQLHINGNKIETIEEDTFSSMIHLTVLDLSNQRLTHVYKNMFKGLKQITVLN 685
Query: 149 LDARQITDAGLANLTSLSGLITLDLFGARITDSG 182
+ QI +L+ + +DL G I D G
Sbjct: 686 ISRNQINSIDNGAFNNLANVRLIDLSGNVIKDIG 719
>INLA_LISMF (Q723K6) Internalin A precursor
Length = 800
Score = 38.9 bits (89), Expect = 0.008
Identities = 36/101 (35%), Positives = 52/101 (50%), Gaps = 10/101 (9%)
Query: 90 LTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNL 149
L LT L+ L +S +V S I ++ L LE L + ++D + L LTNL L+L
Sbjct: 203 LANLTTLERLDISSNKV--SDISVLAKLTNLESLIATNNQISD--ITPLGILTNLDELSL 258
Query: 150 DARQITDAGLANLTSLSGLITLDLFGARITD----SGTTYL 186
+ Q+ D G L SL+ L LDL +I++ SG T L
Sbjct: 259 NGNQLKDIG--TLASLTNLTDLDLANNQISNLAPLSGLTKL 297
Score = 38.9 bits (89), Expect = 0.008
Identities = 34/95 (35%), Positives = 53/95 (55%), Gaps = 15/95 (15%)
Query: 87 LVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKS 146
L NLTGLTL + + + N ++ LN+LE LS +++D + L GLT+L+
Sbjct: 141 LSNLTGLTLFNNQITDIDPLKN-----LTNLNRLE---LSSNTISD--ISALSGLTSLQQ 190
Query: 147 LNLDARQITD-AGLANLTSLSGLITLDLFGARITD 180
L+ Q+TD LANLT+L LD+ +++D
Sbjct: 191 LSF-GNQVTDLKPLANLTTLE---RLDISSNKVSD 221
Score = 32.0 bits (71), Expect = 1.0
Identities = 35/110 (31%), Positives = 55/110 (49%), Gaps = 15/110 (13%)
Query: 86 GLVNLTGLTLLKSLV---LSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLT 142
G+ ++ GL L +L S+ ++ + I + L KL D+ ++ + D + L L+
Sbjct: 87 GIKSIDGLEYLNNLTQINFSNNQL--TDITPLKDLTKLVDILMNNNQIAD--ITPLANLS 142
Query: 143 NLKSLNLDARQITDAG-LANLTSLSGLITLDLFGARITD----SGTTYLR 187
NL L L QITD L NLT+L+ L+L I+D SG T L+
Sbjct: 143 NLTGLTLFNNQITDIDPLKNLTNLN---RLELSSNTISDISALSGLTSLQ 189
>DR18_ARATH (O64789) Putative disease resistance protein At1g61310
Length = 925
Score = 37.7 bits (86), Expect = 0.019
Identities = 40/122 (32%), Positives = 56/122 (45%), Gaps = 11/122 (9%)
Query: 87 LVNLTG-----LTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTD--NGLKRLL 139
L NL+G + L L LSD N ISGL L+ L+LSFT + GLK L
Sbjct: 558 LKNLSGEFIRYMQKLVVLDLSDNRDFNELPEQISGLVSLQYLDLSFTRIEQLPVGLKELK 617
Query: 140 GLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSGTTYLRCMYIYQLQAIVV 199
LT L L AR + +G++ L SL L L G+++ + + LQ + +
Sbjct: 618 KLTFL-DLAYTARLCSISGISRLLSLR---VLSLLGSKVHGDASVLKELQQLENLQDLAI 673
Query: 200 FL 201
L
Sbjct: 674 TL 675
>FXLG_HUMAN (Q8N461) F-box/LRR-repeat protein 16 (F-box and
leucine-rich repeat protein 16) (cg14134)
Length = 479
Score = 36.6 bits (83), Expect = 0.042
Identities = 56/206 (27%), Positives = 97/206 (46%), Gaps = 24/206 (11%)
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQV- 58
+Q + L + GC+ + S A + L+++ C ++DD S L L +
Sbjct: 217 MQGVVRLELSGCNDFTEAGLWSSLSARITSLSVSDCINVADDAIAAISQLLPNLAELSLQ 276
Query: 59 -----------LQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLT-GLTLLKSLVLSD-TE 105
A +G H RL + +I + G+VN+ L L +L LS ++
Sbjct: 277 AYHVTDTALAYFTARQGHSTHTLRLLS----CWEITNHGVVNVVHSLPNLTALSLSGCSK 332
Query: 106 VGNSGIRYIS-GLNKLEDLNLSFTS-VTDNGLKRLL-GLTNLKSLNLD-ARQITDAGLAN 161
V + G+ ++ L KL L+LS+ +TD L+ + L L+ L LD +ITD GL+
Sbjct: 333 VTDDGVELVAENLRKLRSLDLSWCPRITDMALEYVACDLHRLEELVLDRCVRITDTGLSY 392
Query: 162 LTSLSGLITLDL-FGARITDSGTTYL 186
L+++S L +L L + ++ D G +L
Sbjct: 393 LSTMSSLRSLYLRWCCQVQDFGLKHL 418
Score = 32.7 bits (73), Expect = 0.61
Identities = 37/98 (37%), Positives = 53/98 (53%), Gaps = 10/98 (10%)
Query: 100 VLSDTEVGNSGI-RYISGLNKLEDLNLSFTS-VTDNGLKRLL-GLTNLKSLNLD-ARQIT 155
+LS E+ N G+ + L L L+LS S VTD+G++ + L L+SL+L +IT
Sbjct: 301 LLSCWEITNHGVVNVVHSLPNLTALSLSGCSKVTDDGVELVAENLRKLRSLDLSWCPRIT 360
Query: 156 DAGL----ANLTSLSGLITLDLFGARITDSGTTYLRCM 189
D L +L L L+ LD RITD+G +YL M
Sbjct: 361 DMALEYVACDLHRLEELV-LDRC-VRITDTGLSYLSTM 396
>LUM_COTJA (Q9DE67) Lumican precursor (Keratan sulfate proteoglycan
lumican) (KSPG lumican)
Length = 343
Score = 36.2 bits (82), Expect = 0.055
Identities = 34/111 (30%), Positives = 51/111 (45%), Gaps = 27/111 (24%)
Query: 85 EGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVT--DNGLKRLL--- 139
EGLVNLT + L + + +D+ G GLN L L+LSF +T GL L
Sbjct: 161 EGLVNLTVIHLQNNQLKADSISG-----AFKGLNSLLYLDLSFNQLTKLPTGLPHSLLML 215
Query: 140 ----------------GLTNLKSLNLDARQITDAGL-ANLTSLSGLITLDL 173
G L+ L L ++TD+G+ N+ +++ L+ LDL
Sbjct: 216 YFDNNQISNIPDEYFQGFKTLQYLRLSHNKLTDSGIPGNVFNITSLVELDL 266
>LUM_CHICK (P51890) Lumican precursor (Keratan sulfate proteoglycan
lumican) (KSPG lumican)
Length = 343
Score = 36.2 bits (82), Expect = 0.055
Identities = 34/111 (30%), Positives = 51/111 (45%), Gaps = 27/111 (24%)
Query: 85 EGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVT--DNGLKRLL--- 139
EGLVNLT + L + + +D+ G GLN L L+LSF +T GL L
Sbjct: 161 EGLVNLTVIHLQNNQLKTDSISG-----AFKGLNSLLYLDLSFNQLTKLPTGLPHSLLML 215
Query: 140 ----------------GLTNLKSLNLDARQITDAGL-ANLTSLSGLITLDL 173
G L+ L L ++TD+G+ N+ +++ L+ LDL
Sbjct: 216 YFDNNQISNIPDEYFQGFKTLQYLRLSHNKLTDSGIPGNVFNITSLVELDL 266
>LGR4_RAT (Q9Z2H4) Leucine-rich repeat-containing G protein-coupled
receptor 4 precursor
Length = 951
Score = 36.2 bits (82), Expect = 0.055
Identities = 26/84 (30%), Positives = 39/84 (45%)
Query: 90 LTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNL 149
L+GL LK L L + ++ I GL+ L+ L L +T GL L+ L L
Sbjct: 101 LSGLKELKVLTLQNNQLRTVPSEAIHGLSALQSLRLDANHITSVPEDSFEGLVQLRHLWL 160
Query: 150 DARQITDAGLANLTSLSGLITLDL 173
D +T+ + L++L L L L
Sbjct: 161 DDNSLTEVPVRPLSNLPTLQALTL 184
Score = 32.7 bits (73), Expect = 0.61
Identities = 25/85 (29%), Positives = 39/85 (45%), Gaps = 2/85 (2%)
Query: 89 NLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLN 148
NLTG L+SL L+ T++ + L L+LS+ ++ D L G L+ ++
Sbjct: 314 NLTGTVHLESLTLTGTKISSIPDDLCQNQKMLRTLDLSYNNIRD--LPSFNGCRALEEIS 371
Query: 149 LDARQITDAGLANLTSLSGLITLDL 173
L QI+ L+ L LDL
Sbjct: 372 LQRNQISLIKENTFQGLTSLRILDL 396
Score = 29.3 bits (64), Expect = 6.7
Identities = 19/60 (31%), Positives = 32/60 (52%)
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQIT 155
L+ L L+ ++ + +SGL +L+ L L + + + GL+ L+SL LDA IT
Sbjct: 83 LEELQLAGNDLSLIHPKALSGLKELKVLTLQNNQLRTVPSEAIHGLSALQSLRLDANHIT 142
>INLB_LISMO (P25147) Internalin B precursor
Length = 630
Score = 36.2 bits (82), Expect = 0.055
Identities = 32/100 (32%), Positives = 57/100 (57%), Gaps = 6/100 (6%)
Query: 66 VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNL 125
++HL +L++ KI D + L+ LT L +L L D ++ S I ++GL KL++L L
Sbjct: 160 LVHLPQLESLYLGNNKITD--ITVLSRLTKLDTLSLEDNQI--SDIVPLAGLTKLQNLYL 215
Query: 126 SFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSL 165
S ++D L+ L GL NL L L +++ + + + ++L
Sbjct: 216 SKNHISD--LRALAGLKNLDVLELFSQECLNKPINHQSNL 253
>NYX_MOUSE (P83503) Nyctalopin precursor
Length = 476
Score = 35.8 bits (81), Expect = 0.072
Identities = 26/84 (30%), Positives = 37/84 (43%)
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQIT 155
L+ L+L+D + GL +L LNL ++ L L+ L LD IT
Sbjct: 227 LEDLLLNDNLLATLPAAAFRGLRRLRTLNLGGNALGSVARAWFSDLAELELLYLDRNSIT 286
Query: 156 DAGLANLTSLSGLITLDLFGARIT 179
+LSGL+ L L G R+T
Sbjct: 287 FVEEGAFQNLSGLLALHLNGNRLT 310
>LGR4_HUMAN (Q9BXB1) Leucine-rich repeat-containing G
protein-coupled receptor 4 precursor (G protein-coupled
receptor 48)
Length = 951
Score = 35.8 bits (81), Expect = 0.072
Identities = 26/84 (30%), Positives = 39/84 (45%)
Query: 90 LTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNL 149
L+GL LK L L + ++ I GL+ L+ L L +T GL L+ L L
Sbjct: 101 LSGLKELKVLTLQNNQLKTVPSEAIRGLSALQSLRLDANHITSVPEDSFEGLVQLRHLWL 160
Query: 150 DARQITDAGLANLTSLSGLITLDL 173
D +T+ + L++L L L L
Sbjct: 161 DDNSLTEVPVHPLSNLPTLQALTL 184
Score = 32.3 bits (72), Expect = 0.79
Identities = 25/85 (29%), Positives = 37/85 (43%), Gaps = 2/85 (2%)
Query: 89 NLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLN 148
NLTG L+SL L+ T++ + L L+LS+ ++ D L G L+ ++
Sbjct: 314 NLTGTVHLESLTLTGTKISSIPNNLCQEQKMLRTLDLSYNNIRD--LPSFNGCHALEEIS 371
Query: 149 LDARQITDAGLANLTSLSGLITLDL 173
L QI L L LDL
Sbjct: 372 LQRNQIYQIKEGTFQGLISLRILDL 396
Score = 29.6 bits (65), Expect = 5.1
Identities = 19/60 (31%), Positives = 32/60 (52%)
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQIT 155
L+ L L+ ++ + +SGL +L+ L L + + + GL+ L+SL LDA IT
Sbjct: 83 LEELQLAGNDLSFIHPKALSGLKELKVLTLQNNQLKTVPSEAIRGLSALQSLRLDANHIT 142
>PGIP_PYRCO (Q05091) Polygalacturonase inhibitor precursor
(Polygalacturonase-inhibiting protein)
Length = 330
Score = 35.4 bits (80), Expect = 0.094
Identities = 29/75 (38%), Positives = 38/75 (50%), Gaps = 8/75 (10%)
Query: 89 NLTG--------LTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLG 140
NLTG L LKSL LS T + S ++S L L L+LSF ++T L
Sbjct: 106 NLTGPIQPAIAKLKGLKSLRLSWTNLSGSVPDFLSQLKNLTFLDLSFNNLTGAIPSSLSE 165
Query: 141 LTNLKSLNLDARQIT 155
L NL +L LD ++T
Sbjct: 166 LPNLGALRLDRNKLT 180
>LIG2_HUMAN (O94898) Leucine-rich repeats and immunoglobulin-like
domains protein 2 precursor (LIG-2)
Length = 1065
Score = 35.0 bits (79), Expect = 0.12
Identities = 26/85 (30%), Positives = 38/85 (44%), Gaps = 3/85 (3%)
Query: 92 GLTLLKSLVLSD---TEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLN 148
GL+LL+ L L D T + + R++S L L+ N + ++ + GLT+L L
Sbjct: 333 GLSLLERLNLGDNRVTHIADGVFRFLSNLQTLDLRNNEISWAIEDASEAFAGLTSLTKLI 392
Query: 149 LDARQITDAGLANLTSLSGLITLDL 173
L QI L L LDL
Sbjct: 393 LQGNQIKSITKKAFIGLESLEHLDL 417
Score = 35.0 bits (79), Expect = 0.12
Identities = 25/84 (29%), Positives = 39/84 (45%)
Query: 90 LTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNL 149
L GL +L+ L +S + +L +L+LS+ +T +GL+ L+ LNL
Sbjct: 283 LYGLRMLQQLYVSQNAIERISPDAWEFCQRLSELDLSYNQLTRLDESAFVGLSLLERLNL 342
Query: 150 DARQITDAGLANLTSLSGLITLDL 173
++T LS L TLDL
Sbjct: 343 GDNRVTHIADGVFRFLSNLQTLDL 366
Score = 30.8 bits (68), Expect = 2.3
Identities = 40/155 (25%), Positives = 59/155 (37%), Gaps = 26/155 (16%)
Query: 3 KLSTLNVEGCSIT---AACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVL 59
+L LN+ IT A CF+ +S+ + LN NR + K P LQ L
Sbjct: 168 QLKYLNLSNNRITTLEAGCFDNLSSSLLVVKLNRNRMSMIPPKIFKL-------PHLQFL 220
Query: 60 QA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNK 119
+ R + + EGL GL L+SL + + GLN
Sbjct: 221 ELKRNRIKIV---------------EGLT-FQGLDSLRSLKMQRNGISKLKDGAFFGLNN 264
Query: 120 LEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQI 154
+E+L L ++T L GL L+ L + I
Sbjct: 265 MEELELEHNNLTRVNKGWLYGLRMLQQLYVSQNAI 299
>CAR4_HUMAN (Q9Y239) Caspase recruitment domain protein 4 (Nod1
protein)
Length = 953
Score = 35.0 bits (79), Expect = 0.12
Identities = 33/98 (33%), Positives = 46/98 (46%), Gaps = 9/98 (9%)
Query: 98 SLVLSDTEVGNSGIRYISG-LNKLEDLNLSFTSVTDNGLKRLL-GLTNLK---SLNLDAR 152
+L L + + + G+R + ++L L LS +TD G+K L LT K L L
Sbjct: 707 ALDLDNNNLNDYGVRELQPCFSRLTVLRLSVNQITDGGVKVLSEELTKYKIVTYLGLYNN 766
Query: 153 QITDAGLANLTSL----SGLITLDLFGARITDSGTTYL 186
QITD G +T + GL L L +IT G YL
Sbjct: 767 QITDVGARYVTKILDECKGLTHLKLGKNKITSEGGKYL 804
>ALS_RAT (P35859) Insulin-like growth factor binding protein complex
acid labile chain precursor (ALS)
Length = 603
Score = 34.7 bits (78), Expect = 0.16
Identities = 29/94 (30%), Positives = 43/94 (44%), Gaps = 6/94 (6%)
Query: 86 GLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLK 145
GL N+ + L + + S E R GL+KL L+L + + L GL+ L+
Sbjct: 360 GLFNVAVMNLSGNCLRSLPE------RVFQGLDKLHSLHLEHSCLGHVRLHTFAGLSGLR 413
Query: 146 SLNLDARQITDAGLANLTSLSGLITLDLFGARIT 179
L L I+ +L LS L+ LDL R+T
Sbjct: 414 RLFLRDNSISSIEEQSLAGLSELLELDLTTNRLT 447
Score = 33.9 bits (76), Expect = 0.27
Identities = 25/92 (27%), Positives = 44/92 (47%)
Query: 82 IGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGL 141
+G L GL+ L+ L L D + + + ++GL++L +L+L+ +T + GL
Sbjct: 398 LGHVRLHTFAGLSGLRRLFLRDNSISSIEEQSLAGLSELLELDLTTNRLTHLPRQLFQGL 457
Query: 142 TNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
+L+ L L Q+T L L LD+
Sbjct: 458 GHLEYLLLSYNQLTTLSAEVLGPLQRAFWLDI 489
Score = 32.7 bits (73), Expect = 0.61
Identities = 23/84 (27%), Positives = 38/84 (44%)
Query: 92 GLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDA 151
GL L L L+ + + R L+ LE+L L + G + GL L+ L L+
Sbjct: 288 GLLGLHVLRLAHNAIASLRPRTFKDLHFLEELQLGHNRIRQLGERTFEGLGQLEVLTLND 347
Query: 152 RQITDAGLANLTSLSGLITLDLFG 175
QIT+ + + L + ++L G
Sbjct: 348 NQITEVRVGAFSGLFNVAVMNLSG 371
Score = 29.6 bits (65), Expect = 5.1
Identities = 27/78 (34%), Positives = 33/78 (41%)
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQIT 155
L SL LS +G GL+ L DLNL + S+ GL NL L L ++T
Sbjct: 148 LASLSLSSNLLGRLEEGLFQGLSHLWDLNLGWNSLVVLPDTVFQGLGNLHELVLAGNKLT 207
Query: 156 DAGLANLTSLSGLITLDL 173
A L L LDL
Sbjct: 208 YLQPALFCGLGELRELDL 225
>VIRN_NICGU (Q40392) TMV resistance protein N
Length = 1144
Score = 33.9 bits (76), Expect = 0.27
Identities = 23/59 (38%), Positives = 37/59 (61%), Gaps = 2/59 (3%)
Query: 116 GLNKLEDLNLSFTSVTDNGLKRLLG-LTNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
GL+ LE LNLS+ ++ D GL +G L++LK L+L +R + +++ L L +LDL
Sbjct: 831 GLHSLEYLNLSYCNLIDGGLPEEIGSLSSLKKLDL-SRNNFEHLPSSIAQLGALQSLDL 888
>LUM_RAT (P51886) Lumican precursor (Keratan sulfate proteoglycan
lumican) (KSPG lumican)
Length = 338
Score = 33.9 bits (76), Expect = 0.27
Identities = 37/104 (35%), Positives = 52/104 (49%), Gaps = 11/104 (10%)
Query: 81 KIGD-EGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLL 139
K+G +GLVNLT + L + L + V S + GL LE L+LSF ++ K
Sbjct: 151 KLGSFDGLVNLTFIYLQHNQ-LKEEAVSAS----LKGLKSLEYLDLSFNQMS----KLPA 201
Query: 140 GL-TNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSG 182
GL T+L +L LD +IT+ +GL L L + DSG
Sbjct: 202 GLPTSLLTLYLDNNKITNIPDEYFNRFTGLQYLRLSHNELADSG 245
>LIG1_HUMAN (Q96JA1) Leucine-rich repeats and immunoglobulin-like
domains protein 1 precursor (LIG-1)
Length = 1093
Score = 33.9 bits (76), Expect = 0.27
Identities = 27/85 (31%), Positives = 38/85 (43%)
Query: 89 NLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLN 148
+L GLT L L LS+ + + S KL +L LSF ++T + L L++L L
Sbjct: 278 SLYGLTALHQLHLSNNSIARIHRKGWSFCQKLHELVLSFNNLTRLDEESLAELSSLSVLR 337
Query: 149 LDARQITDAGLANLTSLSGLITLDL 173
L I+ L L LDL
Sbjct: 338 LSHNSISHIAEGAFKGLRSLRVLDL 362
Score = 30.8 bits (68), Expect = 2.3
Identities = 18/61 (29%), Positives = 29/61 (47%)
Query: 91 TGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLD 150
+GL L L L ++ + R SGL LE LNL ++ + + NLK L++
Sbjct: 379 SGLDSLSKLTLFGNKIKSVAKRAFSGLEGLEHLNLGGNAIRSVQFDAFVKMKNLKELHIS 438
Query: 151 A 151
+
Sbjct: 439 S 439
Score = 30.0 bits (66), Expect = 3.9
Identities = 28/89 (31%), Positives = 40/89 (44%), Gaps = 9/89 (10%)
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQI- 154
L LVLS + ++ L+ L L LS S++ GL +L+ L+LD +I
Sbjct: 309 LHELVLSFNNLTRLDEESLAELSSLSVLRLSHNSISHIAEGAFKGLRSLRVLDLDHNEIS 368
Query: 155 -----TDAGLANLTSLSGLITLDLFGARI 178
T + L SLS L LFG +I
Sbjct: 369 GTIEDTSGAFSGLDSLS---KLTLFGNKI 394
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.335 0.148 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,633,363
Number of Sequences: 164201
Number of extensions: 686506
Number of successful extensions: 2948
Number of sequences better than 10.0: 73
Number of HSP's better than 10.0 without gapping: 26
Number of HSP's successfully gapped in prelim test: 47
Number of HSP's that attempted gapping in prelim test: 2782
Number of HSP's gapped (non-prelim): 182
length of query: 206
length of database: 59,974,054
effective HSP length: 105
effective length of query: 101
effective length of database: 42,732,949
effective search space: 4316027849
effective search space used: 4316027849
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC144477.5


