

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144430.7 + phase: 0 /pseudo
(491 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
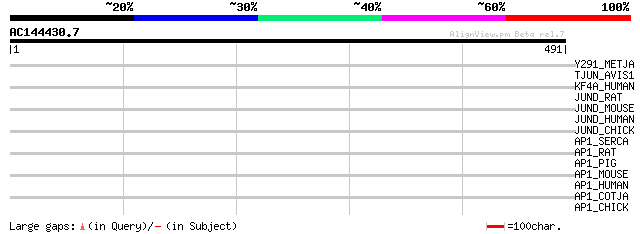
Score E
Sequences producing significant alignments: (bits) Value
Y291_METJA (Q57739) Probable signal recognition particle protein... 33 1.7
TJUN_AVIS1 (P05411) Transforming protein jun 32 4.9
KF4A_HUMAN (O95239) Chromosome-associated kinesin KIF4A (Chromok... 32 4.9
JUND_RAT (P52909) Transcription factor jun-D 31 6.4
JUND_MOUSE (P15066) Transcription factor jun-D 31 6.4
JUND_HUMAN (P17535) Transcription factor jun-D 31 6.4
JUND_CHICK (P27921) Transcription factor jun-D 31 6.4
AP1_SERCA (P54864) Transcription factor AP-1 (Proto-oncogene c-jun) 31 8.3
AP1_RAT (P17325) Transcription factor AP-1 (Activator protein 1)... 31 8.3
AP1_PIG (P56432) Transcription factor AP-1 (Activator protein 1)... 31 8.3
AP1_MOUSE (P05627) Transcription factor AP-1 (Activator protein ... 31 8.3
AP1_HUMAN (P05412) Transcription factor AP-1 (Activator protein ... 31 8.3
AP1_COTJA (P12981) Transcription factor AP-1 (Proto-oncogene c-jun) 31 8.3
AP1_CHICK (P18870) Transcription factor AP-1 (Proto-oncogene c-jun) 31 8.3
>Y291_METJA (Q57739) Probable signal recognition particle protein
MJ0291
Length = 409
Score = 33.1 bits (74), Expect = 1.7
Identities = 18/55 (32%), Positives = 31/55 (55%), Gaps = 3/55 (5%)
Query: 201 EVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLE 255
E K F LF+ K+P+K E+ + EE+K++ ++ P+ K+ KEE+ E
Sbjct: 35 EKSKISFTSLFK-KEPKKEEVKKEKAEEEIKKSEIVKTEPS--EKVEEAKEEIKE 86
>TJUN_AVIS1 (P05411) Transforming protein jun
Length = 296
Score = 31.6 bits (70), Expect = 4.9
Identities = 18/56 (32%), Positives = 32/56 (57%), Gaps = 1/56 (1%)
Query: 200 IEVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLE 255
I+ ++ + + RKR++ RI+R EE +T L QN L + N+L+E+V +
Sbjct: 218 IKAERKRMRNRIAASKSRKRKLERIARLEEKVKT-LKAQNSELASTANMLREQVAQ 272
>KF4A_HUMAN (O95239) Chromosome-associated kinesin KIF4A
(Chromokinesin)
Length = 1232
Score = 31.6 bits (70), Expect = 4.9
Identities = 14/57 (24%), Positives = 31/57 (53%)
Query: 199 LIEVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLE 255
L+ K+ K ++KQ R +E++++ R+ +Q L+ N + + N+L+ + E
Sbjct: 650 LMRQMKEDAEKFRQWKQKRDKEVIQLKERDRKRQYELLKLERNFQKQSNVLRRKTEE 706
>JUND_RAT (P52909) Transcription factor jun-D
Length = 341
Score = 31.2 bits (69), Expect = 6.4
Identities = 22/68 (32%), Positives = 38/68 (55%), Gaps = 4/68 (5%)
Query: 200 IEVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLEKIRT 259
I+ ++ + + RKR++ RISR EE +T L QN L + +LL+E+V +
Sbjct: 263 IKAERKRLRNRIAASKCRKRKLERISRLEEKVKT-LKSQNTELASTASLLREQVAQ---L 318
Query: 260 KRGILTRV 267
K+ +L+ V
Sbjct: 319 KQKVLSHV 326
>JUND_MOUSE (P15066) Transcription factor jun-D
Length = 341
Score = 31.2 bits (69), Expect = 6.4
Identities = 22/68 (32%), Positives = 38/68 (55%), Gaps = 4/68 (5%)
Query: 200 IEVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLEKIRT 259
I+ ++ + + RKR++ RISR EE +T L QN L + +LL+E+V +
Sbjct: 263 IKAERKRLRNRIAASKCRKRKLERISRLEEKVKT-LKSQNTELASTASLLREQVAQ---L 318
Query: 260 KRGILTRV 267
K+ +L+ V
Sbjct: 319 KQKVLSHV 326
>JUND_HUMAN (P17535) Transcription factor jun-D
Length = 347
Score = 31.2 bits (69), Expect = 6.4
Identities = 22/68 (32%), Positives = 38/68 (55%), Gaps = 4/68 (5%)
Query: 200 IEVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLEKIRT 259
I+ ++ + + RKR++ RISR EE +T L QN L + +LL+E+V +
Sbjct: 269 IKAERKRLRNRIAASKCRKRKLERISRLEEKVKT-LKSQNTELASTASLLREQVAQ---L 324
Query: 260 KRGILTRV 267
K+ +L+ V
Sbjct: 325 KQKVLSHV 332
>JUND_CHICK (P27921) Transcription factor jun-D
Length = 323
Score = 31.2 bits (69), Expect = 6.4
Identities = 22/68 (32%), Positives = 37/68 (54%), Gaps = 4/68 (5%)
Query: 200 IEVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLEKIRT 259
I+ ++ + + RKR++ RISR EE K L QN L + +LL+E+V +
Sbjct: 243 IKAERKRLRNRIAASKCRKRKLERISRLEE-KVKSLKSQNTELASTASLLREQVAQ---L 298
Query: 260 KRGILTRV 267
K+ +L+ V
Sbjct: 299 KQKVLSHV 306
>AP1_SERCA (P54864) Transcription factor AP-1 (Proto-oncogene c-jun)
Length = 314
Score = 30.8 bits (68), Expect = 8.3
Identities = 18/56 (32%), Positives = 32/56 (57%), Gaps = 1/56 (1%)
Query: 200 IEVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLE 255
I+ ++ + + RKR++ RI+R EE +T L QN L + N+L+E+V +
Sbjct: 236 IKAERKRMRNRIAASKCRKRKLERIARLEEKVKT-LKAQNSELASTANMLREQVAQ 290
>AP1_RAT (P17325) Transcription factor AP-1 (Activator protein 1)
(AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus
17 oncogene homolog)
Length = 334
Score = 30.8 bits (68), Expect = 8.3
Identities = 18/56 (32%), Positives = 32/56 (57%), Gaps = 1/56 (1%)
Query: 200 IEVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLE 255
I+ ++ + + RKR++ RI+R EE +T L QN L + N+L+E+V +
Sbjct: 256 IKAERKRMRNRIAASKCRKRKLERIARLEEKVKT-LKAQNSELASTANMLREQVAQ 310
>AP1_PIG (P56432) Transcription factor AP-1 (Activator protein 1)
(AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus
17 oncogene homolog)
Length = 331
Score = 30.8 bits (68), Expect = 8.3
Identities = 18/56 (32%), Positives = 32/56 (57%), Gaps = 1/56 (1%)
Query: 200 IEVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLE 255
I+ ++ + + RKR++ RI+R EE +T L QN L + N+L+E+V +
Sbjct: 253 IKAERKRMRNRIAASKCRKRKLERIARLEEKVKT-LKAQNSELASTANMLREQVAQ 307
>AP1_MOUSE (P05627) Transcription factor AP-1 (Activator protein 1)
(AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus
17 oncogene homolog) (Jun A) (AH119)
Length = 334
Score = 30.8 bits (68), Expect = 8.3
Identities = 18/56 (32%), Positives = 32/56 (57%), Gaps = 1/56 (1%)
Query: 200 IEVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLE 255
I+ ++ + + RKR++ RI+R EE +T L QN L + N+L+E+V +
Sbjct: 256 IKAERKRMRNRIAASKCRKRKLERIARLEEKVKT-LKAQNSELASTANMLREQVAQ 310
>AP1_HUMAN (P05412) Transcription factor AP-1 (Activator protein 1)
(AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus
17 oncogene homolog) (p39)
Length = 331
Score = 30.8 bits (68), Expect = 8.3
Identities = 18/56 (32%), Positives = 32/56 (57%), Gaps = 1/56 (1%)
Query: 200 IEVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLE 255
I+ ++ + + RKR++ RI+R EE +T L QN L + N+L+E+V +
Sbjct: 253 IKAERKRMRNRIAASKCRKRKLERIARLEEKVKT-LKAQNSELASTANMLREQVAQ 307
>AP1_COTJA (P12981) Transcription factor AP-1 (Proto-oncogene c-jun)
Length = 313
Score = 30.8 bits (68), Expect = 8.3
Identities = 18/56 (32%), Positives = 32/56 (57%), Gaps = 1/56 (1%)
Query: 200 IEVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLE 255
I+ ++ + + RKR++ RI+R EE +T L QN L + N+L+E+V +
Sbjct: 235 IKAERKRMRNRIAASKCRKRKLERIARLEEKVKT-LKAQNSELASTANMLREQVAQ 289
>AP1_CHICK (P18870) Transcription factor AP-1 (Proto-oncogene c-jun)
Length = 310
Score = 30.8 bits (68), Expect = 8.3
Identities = 18/56 (32%), Positives = 32/56 (57%), Gaps = 1/56 (1%)
Query: 200 IEVQKDQFNKLFRFKQPRKREILRISRREEVKQTGLIMQNPNLRTKLNLLKEEVLE 255
I+ ++ + + RKR++ RI+R EE +T L QN L + N+L+E+V +
Sbjct: 232 IKAERKRMRNRIAASKCRKRKLERIARLEEKVKT-LKAQNSELASTANMLREQVAQ 286
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.371 0.167 0.604
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,451,184
Number of Sequences: 164201
Number of extensions: 1358672
Number of successful extensions: 9433
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 13
Number of HSP's that attempted gapping in prelim test: 9431
Number of HSP's gapped (non-prelim): 15
length of query: 491
length of database: 59,974,054
effective HSP length: 114
effective length of query: 377
effective length of database: 41,255,140
effective search space: 15553187780
effective search space used: 15553187780
T: 11
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 68 (30.8 bits)
Medicago: description of AC144430.7


