

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144345.5 - phase: 0
(131 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
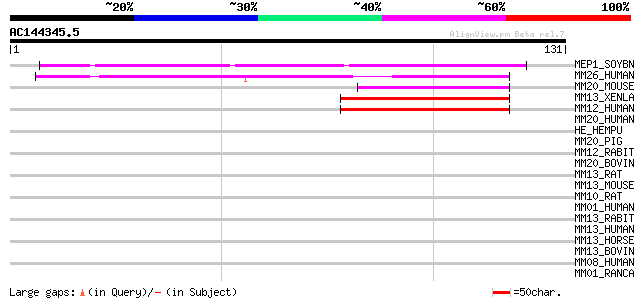
Score E
Sequences producing significant alignments: (bits) Value
MEP1_SOYBN (P29136) Metalloendoproteinase 1 precursor (EC 3.4.24... 71 4e-13
MM26_HUMAN (Q9NRE1) Matrix metalloproteinase-26 precursor (EC 3.... 44 6e-05
MM20_MOUSE (P57748) Matrix metalloproteinase-20 precursor (EC 3.... 40 9e-04
MM13_XENLA (Q10835) Collagenase 3 precursor (EC 3.4.24.-) (Matri... 40 9e-04
MM12_HUMAN (P39900) Macrophage metalloelastase precursor (EC 3.4... 40 9e-04
MM20_HUMAN (O60882) Matrix metalloproteinase-20 precursor (EC 3.... 40 0.001
HE_HEMPU (P91953) Hatching enzyme precursor (EC 3.4.24.12) (HE) ... 39 0.002
MM20_PIG (P79287) Matrix metalloproteinase-20 precursor (EC 3.4.... 39 0.002
MM12_RABIT (P79227) Macrophage metalloelastase precursor (EC 3.4... 39 0.003
MM20_BOVIN (O18767) Matrix metalloproteinase-20 precursor (EC 3.... 38 0.003
MM13_RAT (P23097) Collagenase 3 precursor (EC 3.4.24.-) (Matrix ... 38 0.003
MM13_MOUSE (P33435) Collagenase 3 precursor (EC 3.4.24.-) (Matri... 38 0.003
MM10_RAT (P07152) Stromelysin-2 precursor (EC 3.4.24.22) (Matrix... 38 0.004
MM01_HUMAN (P03956) Interstitial collagenase precursor (EC 3.4.2... 38 0.004
MM13_RABIT (O62806) Collagenase 3 precursor (EC 3.4.24.-) (Matri... 37 0.006
MM13_HUMAN (P45452) Collagenase 3 precursor (EC 3.4.24.-) (Matri... 37 0.006
MM13_HORSE (O18927) Collagenase 3 precursor (EC 3.4.24.-) (Matri... 37 0.006
MM13_BOVIN (O77656) Collagenase 3 precursor (EC 3.4.24.-) (Matri... 37 0.006
MM08_HUMAN (P22894) Neutrophil collagenase precursor (EC 3.4.24.... 37 0.006
MM01_RANCA (Q11133) Interstitial collagenase precursor (EC 3.4.2... 37 0.008
>MEP1_SOYBN (P29136) Metalloendoproteinase 1 precursor (EC 3.4.24.-)
(SMEP1)
Length = 305
Score = 71.2 bits (173), Expect = 4e-13
Identities = 46/115 (40%), Positives = 61/115 (53%), Gaps = 3/115 (2%)
Query: 8 AFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTVIKLGSNVNSGFIR 67
AF++W+ + F ETTSY+ ANIKI F + N+ D G ++ G
Sbjct: 171 AFSKWTPVVNIA-FQETTSYETANIKILFASKNHGDPYPFDGPGG-ILGHAFAPTDGRCH 228
Query: 68 LIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAILP 122
A +YWV D S FDLE+ A+H+IGHLLGL HS D +MYP+I P
Sbjct: 229 FDADEYWVASGD-VTKSPVTSAFDLESVAVHEIGHLLGLGHSSDLRAIMYPSIPP 282
>MM26_HUMAN (Q9NRE1) Matrix metalloproteinase-26 precursor (EC
3.4.24.-) (MMP-26) (Matrilysin-2) (Endometase)
Length = 261
Score = 43.9 bits (102), Expect = 6e-05
Identities = 32/114 (28%), Positives = 52/114 (45%), Gaps = 13/114 (11%)
Query: 7 NAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTV--IKLGSNVNSG 64
NA + WS T ++ + DA+IK+ F+ + DG G + L ++ N G
Sbjct: 126 NAVSIWSNVTPLI--FQQVQNGDADIKVSFWQWAHEDGWPFDGPGGILGHAFLPNSGNPG 183
Query: 65 FIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
+ +++W Y +L A H+IGH LGL HS ++ +MYP
Sbjct: 184 VVHFDKNEHWSASDTGY---------NLFLVATHEIGHSLGLQHSGNQSSIMYP 228
>MM20_MOUSE (P57748) Matrix metalloproteinase-20 precursor (EC
3.4.24.-) (MMP-20) (Enamel metalloproteinase)
(Enamelysin)
Length = 482
Score = 40.0 bits (92), Expect = 9e-04
Identities = 17/36 (47%), Positives = 21/36 (58%)
Query: 83 WSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
W+ F+L T A H+ GH LGL HS D +MYP
Sbjct: 210 WTMGTNGFNLFTVAAHEFGHALGLGHSTDPSALMYP 245
>MM13_XENLA (Q10835) Collagenase 3 precursor (EC 3.4.24.-) (Matrix
metalloproteinase-13) (MMP-13) (Fragment)
Length = 469
Score = 40.0 bits (92), Expect = 9e-04
Identities = 18/40 (45%), Positives = 25/40 (62%)
Query: 79 DNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
D+ M+S N ++L A H+ GH LGLDHS D +M+P
Sbjct: 201 DDEMFSTDNKGYNLFVVAAHEFGHALGLDHSRDPGSLMFP 240
>MM12_HUMAN (P39900) Macrophage metalloelastase precursor (EC
3.4.24.65) (HME) (Matrix metalloproteinase-12) (MMP-12)
(Macrophage elastase) (ME)
Length = 470
Score = 40.0 bits (92), Expect = 9e-04
Identities = 17/40 (42%), Positives = 26/40 (64%)
Query: 79 DNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
++ W+ +G +L A+H+IGH LGL HS D + VM+P
Sbjct: 199 EDEFWTTHSGGTNLFLTAVHEIGHSLGLGHSSDPKAVMFP 238
>MM20_HUMAN (O60882) Matrix metalloproteinase-20 precursor (EC
3.4.24.-) (MMP-20) (Enamel metalloproteinase)
(Enamelysin)
Length = 483
Score = 39.7 bits (91), Expect = 0.001
Identities = 17/36 (47%), Positives = 21/36 (58%)
Query: 83 WSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
W+ F+L T A H+ GH LGL HS D +MYP
Sbjct: 211 WTMGTNGFNLFTVAAHEFGHALGLAHSTDPSALMYP 246
>HE_HEMPU (P91953) Hatching enzyme precursor (EC 3.4.24.12) (HE)
(HEZ) (Envelysin) (Sea-urchin-hatching proteinase)
Length = 591
Score = 39.3 bits (90), Expect = 0.002
Identities = 33/116 (28%), Positives = 47/116 (40%), Gaps = 10/116 (8%)
Query: 3 NVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTVIKLGSNVN 62
N R AF W + L F E +I+I F + + DG+ G V+
Sbjct: 201 NELRRAFQVWDDVSS-LTFREVVDSSSVDIRIKFGSYEHGDGISFDGQGG-VLAHAFLPR 258
Query: 63 SGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
+G S+ W + T N +L A H+ GH LGL HS + +MYP
Sbjct: 259 NGDAHFDDSERWTIGT--------NSGTNLFQVAAHEFGHSLGLYHSDVQSALMYP 306
>MM20_PIG (P79287) Matrix metalloproteinase-20 precursor (EC
3.4.24.-) (MMP-20) (Enamel metalloproteinase)
(Enamelysin)
Length = 483
Score = 38.9 bits (89), Expect = 0.002
Identities = 17/36 (47%), Positives = 21/36 (58%)
Query: 83 WSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
W+ F+L T A H+ GH LGL HS D +MYP
Sbjct: 211 WTMGMNGFNLFTVAAHEFGHALGLAHSTDPSALMYP 246
>MM12_RABIT (P79227) Macrophage metalloelastase precursor (EC
3.4.24.65) (MME) (Matrix metalloproteinase-12) (MMP-12)
Length = 464
Score = 38.5 bits (88), Expect = 0.003
Identities = 16/40 (40%), Positives = 26/40 (65%)
Query: 79 DNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
++ +WS +L A+H++GH LGLDHS D + +M+P
Sbjct: 194 EDEIWSKSYKGTNLFLVAVHELGHALGLDHSNDPKAIMFP 233
>MM20_BOVIN (O18767) Matrix metalloproteinase-20 precursor (EC
3.4.24.-) (MMP-20) (Enamel metalloproteinase)
(Enamelysin)
Length = 481
Score = 38.1 bits (87), Expect = 0.003
Identities = 16/36 (44%), Positives = 21/36 (57%)
Query: 83 WSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
W+ F+L T A H+ GH LGL HS D +M+P
Sbjct: 209 WTMGTNGFNLFTVAAHEFGHALGLAHSTDPSALMFP 244
>MM13_RAT (P23097) Collagenase 3 precursor (EC 3.4.24.-) (Matrix
metalloproteinase-13) (MMP-13) (UMRCASE) (Fragment)
Length = 466
Score = 38.1 bits (87), Expect = 0.003
Identities = 16/40 (40%), Positives = 25/40 (62%)
Query: 79 DNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
D+ W+ + ++L A H++GH LGLDHS D +M+P
Sbjct: 198 DDETWTSSSKGYNLFIVAAHELGHSLGLDHSKDPGALMFP 237
>MM13_MOUSE (P33435) Collagenase 3 precursor (EC 3.4.24.-) (Matrix
metalloproteinase-13) (MMP-13)
Length = 472
Score = 38.1 bits (87), Expect = 0.003
Identities = 16/40 (40%), Positives = 25/40 (62%)
Query: 79 DNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
D+ W+ + ++L A H++GH LGLDHS D +M+P
Sbjct: 204 DDETWTSSSKGYNLFIVAAHELGHSLGLDHSKDPGALMFP 243
>MM10_RAT (P07152) Stromelysin-2 precursor (EC 3.4.24.22) (Matrix
metalloproteinase-10) (MMP-10) (Transin-2) (SL-2)
(Transformation-associated protein 34A)
Length = 476
Score = 37.7 bits (86), Expect = 0.004
Identities = 18/40 (45%), Positives = 24/40 (60%)
Query: 79 DNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
D+ WS +L A H++GH LGL HS +KE +MYP
Sbjct: 199 DDEKWSLGPSGTNLFLVAAHELGHSLGLFHSNNKESLMYP 238
>MM01_HUMAN (P03956) Interstitial collagenase precursor (EC
3.4.24.7) (Matrix metalloproteinase-1) (MMP-1)
(Fibroblast collagenase)
Length = 469
Score = 37.7 bits (86), Expect = 0.004
Identities = 17/37 (45%), Positives = 23/37 (61%), Gaps = 2/37 (5%)
Query: 85 WQNG--EFDLETAAMHQIGHLLGLDHSFDKEYVMYPA 119
W N E++L A H++GH LGL HS D +MYP+
Sbjct: 203 WTNNFREYNLHRVAAHELGHSLGLSHSTDIGALMYPS 239
>MM13_RABIT (O62806) Collagenase 3 precursor (EC 3.4.24.-) (Matrix
metalloproteinase-13) (MMP-13)
Length = 471
Score = 37.4 bits (85), Expect = 0.006
Identities = 16/40 (40%), Positives = 24/40 (60%)
Query: 79 DNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
D+ W+ + ++L A H+ GH LGLDHS D +M+P
Sbjct: 203 DDETWTSSSKGYNLFLVAAHEFGHSLGLDHSKDPGALMFP 242
>MM13_HUMAN (P45452) Collagenase 3 precursor (EC 3.4.24.-) (Matrix
metalloproteinase-13) (MMP-13)
Length = 471
Score = 37.4 bits (85), Expect = 0.006
Identities = 16/40 (40%), Positives = 24/40 (60%)
Query: 79 DNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
D+ W+ + ++L A H+ GH LGLDHS D +M+P
Sbjct: 203 DDETWTSSSKGYNLFLVAAHEFGHSLGLDHSKDPGALMFP 242
>MM13_HORSE (O18927) Collagenase 3 precursor (EC 3.4.24.-) (Matrix
metalloproteinase-13) (MMP-13)
Length = 472
Score = 37.4 bits (85), Expect = 0.006
Identities = 16/40 (40%), Positives = 24/40 (60%)
Query: 79 DNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
D+ W+ + ++L A H+ GH LGLDHS D +M+P
Sbjct: 204 DDETWTSSSKGYNLFLVAAHEFGHSLGLDHSKDPGALMFP 243
>MM13_BOVIN (O77656) Collagenase 3 precursor (EC 3.4.24.-) (Matrix
metalloproteinase-13) (MMP-13)
Length = 471
Score = 37.4 bits (85), Expect = 0.006
Identities = 16/40 (40%), Positives = 24/40 (60%)
Query: 79 DNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
D+ W+ + ++L A H+ GH LGLDHS D +M+P
Sbjct: 203 DDETWTSSSKGYNLFLVAAHEFGHSLGLDHSKDPGALMFP 242
>MM08_HUMAN (P22894) Neutrophil collagenase precursor (EC 3.4.24.34)
(Matrix metalloproteinase-8) (MMP-8) (PMNL collagenase)
(PMNL-CL)
Length = 467
Score = 37.4 bits (85), Expect = 0.006
Identities = 32/118 (27%), Positives = 51/118 (43%), Gaps = 19/118 (16%)
Query: 6 RNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINY-----IDGVDDVVVGDTVIKLGSN 60
++AF WS + ++ S +A+I I FY ++ DG + ++ + G
Sbjct: 134 KDAFELWSVASPLI--FTRISQGEADINIAFYQRDHGDNSPFDGPNGILAH--AFQPGQG 189
Query: 61 VNSGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
+ G A + W + NY +L A H+ GH LGL HS D +MYP
Sbjct: 190 IG-GDAHFDAEETWTNTSANY---------NLFLVAAHEFGHSLGLAHSSDPGALMYP 237
>MM01_RANCA (Q11133) Interstitial collagenase precursor (EC
3.4.24.7) (Matrix metalloproteinase-1) (MMP-1) (TC1)
Length = 384
Score = 37.0 bits (84), Expect = 0.008
Identities = 16/39 (41%), Positives = 23/39 (58%)
Query: 83 WSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAIL 121
W+ +++L A H++GH LGL HS D +MYP L
Sbjct: 174 WTKNFQDYNLYRVAAHELGHSLGLSHSTDIGALMYPTYL 212
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.138 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,019,086
Number of Sequences: 164201
Number of extensions: 652259
Number of successful extensions: 1595
Number of sequences better than 10.0: 104
Number of HSP's better than 10.0 without gapping: 84
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 1506
Number of HSP's gapped (non-prelim): 108
length of query: 131
length of database: 59,974,054
effective HSP length: 107
effective length of query: 24
effective length of database: 42,404,547
effective search space: 1017709128
effective search space used: 1017709128
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC144345.5


