

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143341.6 + phase: 0
(683 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
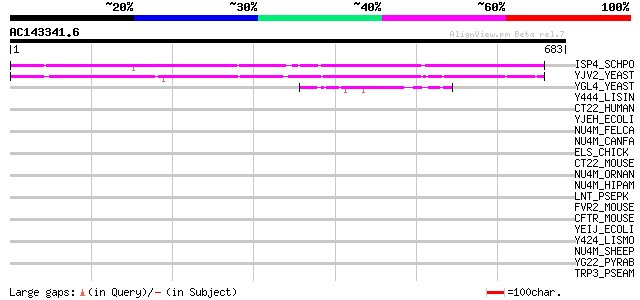
Score E
Sequences producing significant alignments: (bits) Value
ISP4_SCHPO (P40900) Sexual differentiation process protein isp4 464 e-130
YJV2_YEAST (P40897) Hypothetical 91.6 kDa protein in HXT8-CRT1 i... 415 e-115
YGL4_YEAST (P53134) Hypothetical 80.0 kDa protein in SNF4-TAF60 ... 45 5e-04
Y444_LISIN (Q92EL4) Putative sugar uptake protein lin0444 34 1.1
CT22_HUMAN (Q8N2K0) Protein C20orf22 33 2.5
YJEH_ECOLI (P39277) Hypothetical protein yjeH 32 4.2
NU4M_FELCA (P48916) NADH-ubiquinone oxidoreductase chain 4 (EC 1... 32 4.2
NU4M_CANFA (Q9ZZ58) NADH-ubiquinone oxidoreductase chain 4 (EC 1... 32 4.2
ELS_CHICK (P07916) Elastin precursor (Tropoelastin) (Fragment) 32 4.2
CT22_MOUSE (Q99LR1) Protein C20orf22 homolog 32 4.2
NU4M_ORNAN (Q36458) NADH-ubiquinone oxidoreductase chain 4 (EC 1... 32 5.5
NU4M_HIPAM (Q9ZZY2) NADH-ubiquinone oxidoreductase chain 4 (EC 1... 32 5.5
LNT_PSEPK (Q88DN4) Apolipoprotein N-acyltransferase (EC 2.3.1.-)... 32 5.5
FVR2_MOUSE (Q91X85) Feline leukemia virus subgroup C receptor-re... 32 5.5
CFTR_MOUSE (P26361) Cystic fibrosis transmembrane conductance re... 32 5.5
YEIJ_ECOLI (P33021) Hypothetical transport protein yeiJ 32 7.2
Y424_LISMO (Q8Y9U6) Putative sugar uptake protein lmo0424 32 7.2
NU4M_SHEEP (O78755) NADH-ubiquinone oxidoreductase chain 4 (EC 1... 32 7.2
YG22_PYRAB (Q9UY86) Hypothetical UPF0290 protein PYRAB16220 31 9.4
TRP3_PSEAM (O93267) Trypsinogen-like protein 3 precursor 31 9.4
>ISP4_SCHPO (P40900) Sexual differentiation process protein isp4
Length = 785
Score = 464 bits (1195), Expect = e-130
Identities = 234/662 (35%), Positives = 374/662 (56%), Gaps = 20/662 (3%)
Query: 1 MNQFLGYKTNPLKITSVSAQIITLPLGKLMAATLPIKTIQVPFTSLTFSLNPGSFSVKEH 60
+N F + + ++ + ++++ P ++ P + ++ L F+ PG F+VKEH
Sbjct: 108 VNMFFSLRNPTVTLSVLISELLAYPALQIWDLIFPDREFRIG--RLKFNFKPGPFNVKEH 165
Query: 61 VLISIFATSGSSGVYAINIITIVKGFYHRNIHPIAAYLLALSTQILGYGWAGIFRRFLVD 120
LI + ++ Y+ +II + Y + L L+TQ++GYG AG+ RR LV
Sbjct: 166 ALIVVMSSVSFGNAYSTDIILAQRVHYKQRFGFGYEICLTLATQLIGYGLAGLSRRLLVR 225
Query: 121 SPYMWWPEILVQVSLFRAFHEKEKRPKGGT----SRLQFFFLVFVASFAYYIVPGYFFQA 176
M WP LVQ +L + H K+ R S +FF VF+ASF + P Y FQA
Sbjct: 226 PASMLWPVNLVQCTLIKTLHRKDLRNAVANGWRISPFRFFLYVFIASFIWNWFPSYIFQA 285
Query: 177 ISTVSFFCLIWKDSITAQQIGSGMKGLGIGSFGLDWNTVAGFLGSPLAVPGFAIINIMAG 236
+S ++ I +S T QI G+ I DWN ++ ++ SPL P A++NI+ G
Sbjct: 286 LSLFAWVTWIRPNSPTVNQIFGESTGISILPMTFDWNQISAYILSPLMAPADALMNILLG 345
Query: 237 FVLYMYVLIPIAYWNNVYDAKKFPLISSHTFDSTGATYNITRILNTKTFDIDMESYNNYS 296
+L+ +++ P + N + P+ SS D G +YN+TRIL TK D+++Y NYS
Sbjct: 346 VILFFWIVTPALNFTNTWYGDYLPISSSGIIDHFGNSYNVTRIL-TKDATFDLDAYQNYS 404
Query: 297 KIYLSVTFAFQYGLSFAALTATISHVVLFHGEMIVQMWKKTTTSLKNQLGDVHTRIMKKN 356
I++S T+A +GLSFA++T+ I HV+L+HG+ I + D+H ++MK
Sbjct: 405 PIFMSTTYALAFGLSFASITSVIFHVILYHGKEIYDRLRDPPAP------DIHEKLMKA- 457
Query: 357 YEQVPEWWFVAILILMVMMALLACEGFGKQLQLPWWGILLSLSIALVFTLPIGVIEATTS 416
Y++VP +W++++ + M + +G + + PWW I++ + + V+ +PIG+++A T+
Sbjct: 458 YDEVPFYWYLSVFLAFFGMMMGTI--YGWKTETPWWVIIVGVIFSAVWFIPIGIVQAITN 515
Query: 417 ARTGLNVITELVIGFTYPGKPLANVAFKTYGHTSMVQALGFLGDFKLGHYMKIPPKSMFI 476
+ GLNV TE ++G+ YPG+PLA + FKT G+ +M Q L F D K GHYMK+PP+ MF
Sbjct: 516 IQLGLNVFTEFIVGYMYPGRPLAMMIFKTVGYITMTQGLAFAADLKFGHYMKLPPRIMFY 575
Query: 477 VQLVGTVVSLSVHFGTAWWLLTSIENICDESLLPKGSPWTCPGDDAFYNASIIWGVVGPK 536
Q++ T+ S V G W L +I+N+C + +TCP F+N+S+IWGV+GPK
Sbjct: 576 TQMIATIWSCFVQIGVLDWALGNIDNVCQAD---QPDNYTCPNATVFFNSSVIWGVIGPK 632
Query: 537 RMFTNDGVYPGMNWFFLIGLIAPVPVWLLSIKFPNHKWIKLINIPIIIVGASTIPPRRSV 596
RMF+ Y G+ +F+L G++ + W L K+P KW +N P+I G IPP V
Sbjct: 633 RMFSGKNTYTGLQYFWLAGVLGTILFWALWKKWP-QKWWGQLNGPLIFGGTGYIPPATPV 691
Query: 597 NYITWGIVGMFFNFYVYRKFRVWWARHTYILSAGLDAGVAFMGLVLYFSLQSYGIFGPTW 656
NY+ W +G+FFN+Y+ + F WW ++ + LSA LD G ++L+F LQ + P W
Sbjct: 692 NYLAWSGIGLFFNYYLKKIFADWWQKYNFTLSAALDTGTQLSVIILFFCLQLPMVNFPDW 751
Query: 657 WG 658
WG
Sbjct: 752 WG 753
>YJV2_YEAST (P40897) Hypothetical 91.6 kDa protein in HXT8-CRT1
intergenic region
Length = 799
Score = 415 bits (1067), Expect = e-115
Identities = 227/667 (34%), Positives = 376/667 (56%), Gaps = 29/667 (4%)
Query: 1 MNQFLGYKTNPLKITSVSAQIITLPLGKLMAATLPIKTIQVPFTSLTFSLNPGSFSVKEH 60
+NQF + L+I + AQ++ P+G+++A K +VPF F LNPG F+ KEH
Sbjct: 123 VNQFFSLRYPSLEINFLVAQVVCYPIGRILALLPDWKCSKVPF----FDLNPGPFTKKEH 178
Query: 61 VLISIFATSGSSGVYAINIITIVKGFYHRNIHPIAAYLLALSTQILGYGWAGIFRRFLVD 120
+++I SS YA+ I+ FY+ ++ +LL ++Q++GYG AG+ RR++V+
Sbjct: 179 AVVTIAVALTSSTAYAMYILNAQGSFYNMKLNVGYQFLLVWTSQMIGYGAAGLTRRWVVN 238
Query: 121 SPYMWWPEILVQVSLFRAFHEK--EKRPKGGTS--RLQFFFLVFVASFAYYIVPGYFFQA 176
WP+ L+ VSLF + H + EK G + R +FF +V + SF +Y VPG+ F
Sbjct: 239 PASSIWPQTLISVSLFDSLHSRKVEKTVANGWTMPRYRFFLIVLIGSFIWYWVPGFLFTG 298
Query: 177 ISTVSFFCLIW----KDSITAQQIGSGMKGLGIGSFGLDWNTVAGFL-GSPLAVPGFAII 231
+S F ++W + + A I GLG D+ V+ + GS A P +
Sbjct: 299 LSY--FNVILWGSKTRHNFIANTIFGTQSGLGALPITFDYTQVSQAMSGSVFATPFYVSA 356
Query: 232 NIMAGFVLYMYVLIPIAYWNNVYDAKKFPLISSHTFDSTGATYNITRILNTKTFDIDMES 291
N A +++ +++P Y+ N + AK P+IS T+D+T YN+T+ILN + + I++E
Sbjct: 357 NTYASVLIFFVIVLPCLYFTNTWYAKYMPVISGSTYDNTQNKYNVTKILN-EDYSINLEK 415
Query: 292 YNNYSKIYLSVTFAFQYGLSFAALTATISHVVLFHGEMIVQMWKKTTTSLKNQLGDVHTR 351
Y YS +++ ++ Y L+FAA+ A H +L+HG+ IV +K KN D+H R
Sbjct: 416 YKEYSPVFVPFSYLLSYALNFAAVIAVFVHCILYHGKDIVAKFKDR----KNGGTDIHMR 471
Query: 352 IMKKNYEQVPEWWFVAILILMVMMALLACEGFGKQLQLPWWGILLSLSIALVFTLPIGVI 411
I KNY+ P+WW++ + I+M+ + +A F + P W ++++ I+LV +P G++
Sbjct: 472 IYSKNYKDCPDWWYLLLQIVMIGLGFVAVCCF--DTKFPAWAFVIAILISLVNFIPQGIL 529
Query: 412 EATTSARTGLNVITELVIGFTYPGKPLANVAFKTYGHTSMVQALGFLGDFKLGHYMKIPP 471
EA T+ GLN+ITEL+ G+ P +P+AN+ FK YG M Q L D KL YMK+ P
Sbjct: 530 EAMTNQHVGLNIITELICGYMLPLRPMANLLFKLYGFIVMRQGLNLSRDLKLAMYMKVSP 589
Query: 472 KSMFIVQLVGTVVSLSVHFGTAWWLLTSIENICDESLLPKGSPWTCPGDDAFYNASIIWG 531
+ +F VQ+ T++S V+ G W++ +I+ +C P G +TC +NASIIW
Sbjct: 590 RLIFAVQIYATIISGMVNVGVQEWMMHNIDGLCTTD-QPNG--FTCANGRTVFNASIIWS 646
Query: 532 VVGPKRMFTNDGVYPGMNWFFLIGLIAPVPVWLLSIKFPNHKWIKLINIPIIIVGASTIP 591
+ PK +F++ +Y + WFFLIGL+ P+ V+ + KFP K+ K I+ P+ G IP
Sbjct: 647 L--PKYLFSSGRIYNPLMWFFLIGLLFPLAVYAVQWKFPKFKFAKHIHTPVFFTGPGNIP 704
Query: 592 PRRSVNYITWGIVGMFFNFYVYRKFRVWWARHTYILSAGLDAGVAFMGLVLYFSLQSYGI 651
P NY + + N + +++R W+ ++ +++ AG++AGVA ++++ +Q Y
Sbjct: 705 PSTPYNYSLFFAMSFCLNL-IRKRWRAWFNKYNFVMGAGVEAGVAISVVIIFLCVQ-YPG 762
Query: 652 FGPTWWG 658
+WWG
Sbjct: 763 GKLSWWG 769
>YGL4_YEAST (P53134) Hypothetical 80.0 kDa protein in SNF4-TAF60
intergenic region
Length = 725
Score = 45.4 bits (106), Expect = 5e-04
Identities = 46/195 (23%), Positives = 85/195 (43%), Gaps = 30/195 (15%)
Query: 357 YEQVPEWWFVAILILMVMMALLACEGFGKQLQLPWWGILLSLSIALVFTLPIGVI---EA 413
Y V V+ +I +V + L FG Q+ +P + I+ +L +AL ++ +G+ E
Sbjct: 445 YTTVISGCLVSSIICIVSIIYL----FGIQV-IPLYAIITALILALFLSI-LGIRALGET 498
Query: 414 TTSARTGLNVITELVIGFTYP----GKPLANVAFKTYGHTSMVQALGFLGDFKLGHYMKI 469
+ +G+ I++L+ F P G L NV S QA + D K GH +
Sbjct: 499 DLNPVSGIGKISQLIFAFIIPRDRPGSVLMNVVSGGIAEASAQQAGDLMQDLKTGHLLGA 558
Query: 470 PPKSMFIVQLVGTVVSLSVHFGTAWWLLTSIENICDESLLPKGSPWTCPGDDAFYNASII 529
P++ F QL+G S+ +L+S +C + ++ P + +++
Sbjct: 559 SPRAQFCAQLIGACWSI---------ILSSFMYLCYNKV------YSIPSEQFRIPTAVV 603
Query: 530 WGVVGPKRMFTNDGV 544
W + R+ T G+
Sbjct: 604 W--IDCARLVTGKGL 616
>Y444_LISIN (Q92EL4) Putative sugar uptake protein lin0444
Length = 285
Score = 34.3 bits (77), Expect = 1.1
Identities = 66/312 (21%), Positives = 127/312 (40%), Gaps = 60/312 (19%)
Query: 94 IAAYLLALSTQILGYGWAGIFRRFLVDSPYMWWPEILVQVSLFRAFHEKEKRPKGGTSRL 153
++ YL+AL +LG+G+ I +P E L+ S+ S L
Sbjct: 1 MSIYLIAL-LPVLGWGFMPIIANLRKSTP----EEQLLGTSI---------------SAL 40
Query: 154 QFFFLVF--------VASFAYYIVPGYFFQAISTVSFFCLIWKDSITAQQIGSGMKGLGI 205
F F++F V SF V G F+ + F + A I +G + +G
Sbjct: 41 LFAFILFWILSPEITVLSFIVSFVSGIFWSFGQLLQFKGIAASSVAKAMPISNGTQLVGA 100
Query: 206 GSFGL----DWNTVAGFLGSPLAVPGFAIINIMAGFV-----------LYMY-VLIPIAY 249
F + +W TV + +AV I +M GF ++Y ++I ++
Sbjct: 101 TLFAVLVFQEWQTVTAVIIGVVAVILILIGVVMTGFQKRGNHITESVSFHVYGIVILSSF 160
Query: 250 WNNVYDAKKFPLISSHTFDSTGATYNITRILNTKTFDIDMESYNNYSKIYLSVTFAFQYG 309
+ +Y ++++ FD TG + + + + T I + + +VTF G
Sbjct: 161 FLTLY------VVTNQLFDVTGFSIILPQAIGMLTCAIGINLAKKTAISRKNVTFNLMTG 214
Query: 310 LSFAALTATISHVVLFHGEMIVQMWKKTTTSLKNQLGDVHT---RIMKKNYEQVPEWWFV 366
LS+ +I+++ +F ++ + T+ S+ V T ++ K + EW F+
Sbjct: 215 LSW-----SIANLGMFLATAVLGV--ATSFSISQACVIVATIGGILIFKQKKSPLEWTFI 267
Query: 367 AILILMVMMALL 378
IL++M+ ++
Sbjct: 268 LSGILLIMVGVV 279
>CT22_HUMAN (Q8N2K0) Protein C20orf22
Length = 398
Score = 33.1 bits (74), Expect = 2.5
Identities = 17/47 (36%), Positives = 26/47 (55%), Gaps = 6/47 (12%)
Query: 545 YPGMNWFFLIGLIAPVPVWLLSIKFPNHKWIKLINIPIIIVGASTIP 591
+PG +WFFL P+ IKF N + +K I+ P++I+ A P
Sbjct: 294 FPGFDWFFLD------PITSSGIKFANDENVKHISCPLLILHAEDDP 334
>YJEH_ECOLI (P39277) Hypothetical protein yjeH
Length = 418
Score = 32.3 bits (72), Expect = 4.2
Identities = 40/152 (26%), Positives = 67/152 (43%), Gaps = 20/152 (13%)
Query: 358 EQVPEWWFVAILILMVMMALLACEGFGKQLQLPW--WGILLSLSIALVFTLPIGVIEATT 415
E+V W F++++ + + AL GFG Q W W +LL+ L IG A++
Sbjct: 85 ERVTGWLFLSVIPVGLPAALQIAAGFG-QAMFGWHSWQLLLAELGTLALVWYIGTRGASS 143
Query: 416 SARTG---LNVITELVIGFTYPG--KPLANVAFKTYGH---TSMVQALG-----FLGDFK 462
SA +I L++ + G KP AN+ F G+ T + AL F+G
Sbjct: 144 SANLQTVIAGLIVALIVAIWWAGDIKP-ANIPFPAPGNIELTGLFAALSVMFWCFVGLEA 202
Query: 463 LGHY---MKIPPKSMFIVQLVGTVVSLSVHFG 491
H K P + ++G +++ V++G
Sbjct: 203 FAHLASEFKNPERDFPRALMIGLLLAGLVYWG 234
>NU4M_FELCA (P48916) NADH-ubiquinone oxidoreductase chain 4 (EC
1.6.5.3)
Length = 459
Score = 32.3 bits (72), Expect = 4.2
Identities = 28/118 (23%), Positives = 57/118 (47%), Gaps = 13/118 (11%)
Query: 387 LQLPWWGILLSLSIALVFTLPIGVIEATTSARTGLNVITELV-IGFTYPGKPLANVAFKT 445
+ L WG++++ SI L T +I ++ + L ++ L+ ++Y G +A
Sbjct: 262 MMLSLWGMVMTSSICLRQTDLKSLIAYSSVSHMALVIVAVLIQTPWSYMGATALMIA--- 318
Query: 446 YGHTSMVQALGFLGDFKLGHYMKIPPKSMFIVQLVGTVVSLSVHFGTAWWLLTSIENI 503
+G TS + L +Y ++ ++M + + + T++ L AWWLL S+ N+
Sbjct: 319 HGLTSSM-----LFCLANSNYERVHSRTMILARGLQTILPLMA----AWWLLASLANL 367
>NU4M_CANFA (Q9ZZ58) NADH-ubiquinone oxidoreductase chain 4 (EC
1.6.5.3)
Length = 459
Score = 32.3 bits (72), Expect = 4.2
Identities = 29/118 (24%), Positives = 57/118 (47%), Gaps = 13/118 (11%)
Query: 387 LQLPWWGILLSLSIALVFTLPIGVIEATTSARTGLNVITELV-IGFTYPGKPLANVAFKT 445
+ L WG++++ SI L T +I ++ + L ++ L+ ++Y G +A
Sbjct: 262 MMLSLWGMIMTSSICLRQTDLKSLIAYSSVSHMALVIVAVLIQTPWSYMGATALMIA--- 318
Query: 446 YGHTSMVQALGFLGDFKLGHYMKIPPKSMFIVQLVGTVVSLSVHFGTAWWLLTSIENI 503
+G TS + L +Y +I ++M + + + T++ L AWWLL S+ N+
Sbjct: 319 HGLTSSM-----LFCLANSNYERIHSRTMILARGLQTLLPLMA----AWWLLASLTNL 367
>ELS_CHICK (P07916) Elastin precursor (Tropoelastin) (Fragment)
Length = 750
Score = 32.3 bits (72), Expect = 4.2
Identities = 18/42 (42%), Positives = 23/42 (53%), Gaps = 6/42 (14%)
Query: 197 GSGMKGLGIGSFGLDWNTVAGFLGSPLAVPGFAIINIMAGFV 238
G G+ GLG+G G AG LG+ + VPGF + I G V
Sbjct: 675 GLGVGGLGVGGLG------AGGLGAGVGVPGFGVSPIFPGGV 710
>CT22_MOUSE (Q99LR1) Protein C20orf22 homolog
Length = 398
Score = 32.3 bits (72), Expect = 4.2
Identities = 17/47 (36%), Positives = 26/47 (55%), Gaps = 6/47 (12%)
Query: 545 YPGMNWFFLIGLIAPVPVWLLSIKFPNHKWIKLINIPIIIVGASTIP 591
+PG +WFFL P+ IKF N + +K I+ P++I+ A P
Sbjct: 294 FPGFDWFFLD------PITSSGIKFANDENMKHISCPLLILHAEDDP 334
>NU4M_ORNAN (Q36458) NADH-ubiquinone oxidoreductase chain 4 (EC
1.6.5.3)
Length = 460
Score = 32.0 bits (71), Expect = 5.5
Identities = 31/140 (22%), Positives = 61/140 (43%), Gaps = 24/140 (17%)
Query: 373 VMMALLACEGFGKQLQLPW-----WGILLSLSIALVFTLPIGVIEATTSARTGLNVITEL 427
++ ++ E K + P+ WG++++ SI L T +I ++ + GL V L
Sbjct: 243 ILRIIIILEPISKFMAYPFIIMATWGMIMTSSICLRQTDLKSMIAYSSVSHMGLVVAASL 302
Query: 428 VIGFTYPGKPLANVAFKTYGHTSMVQALGFLGDFKL----GHYMKIPPKSMFIVQLVGTV 483
+ + G T+M+ A G +Y +I ++M +V+ + V
Sbjct: 303 I-----------QTPWGFMGATAMMIAHGLTSSMLFCLANTNYERIHSRTMLLVRGLQMV 351
Query: 484 VSLSVHFGTAWWLLTSIENI 503
+ L ++WWLL S+ N+
Sbjct: 352 LPLM----SSWWLLASLANL 367
>NU4M_HIPAM (Q9ZZY2) NADH-ubiquinone oxidoreductase chain 4 (EC
1.6.5.3)
Length = 459
Score = 32.0 bits (71), Expect = 5.5
Identities = 29/118 (24%), Positives = 57/118 (47%), Gaps = 13/118 (11%)
Query: 387 LQLPWWGILLSLSIALVFTLPIGVIEATTSARTGLNVITELV-IGFTYPGKPLANVAFKT 445
+ L WG++++ SI L T +I ++ + L ++ L+ ++Y G +A
Sbjct: 262 IMLSLWGMIMTSSICLRQTDLKSLIAYSSVSHMALVIVAILIQTPWSYMGATALMIA--- 318
Query: 446 YGHTSMVQALGFLGDFKLGHYMKIPPKSMFIVQLVGTVVSLSVHFGTAWWLLTSIENI 503
+G TS + L +Y +I ++M + + + T++ L AWWLL S+ N+
Sbjct: 319 HGLTSSM-----LFCLANSNYERIHSRTMILARGLQTLLPLMA----AWWLLASLTNL 367
>LNT_PSEPK (Q88DN4) Apolipoprotein N-acyltransferase (EC 2.3.1.-)
(ALP N-acyltransferase)
Length = 505
Score = 32.0 bits (71), Expect = 5.5
Identities = 27/114 (23%), Positives = 51/114 (44%), Gaps = 20/114 (17%)
Query: 557 IAPVPVWLLSIKFPNHKWIKLINIPIIIVGASTIPPRRSVNYITWGIVGMFFNFYVYRKF 616
+AP +W L++ ++I ++ +G + PR+++ W G +F F +Y
Sbjct: 26 LAPFDIWPLAV----------LSIAVLYLGLRELSPRQAM----WR--GWWFGFGLYGA- 68
Query: 617 RVWWARHTYILSAGLDAGVAFMGLVLYFSLQSYGIFGPTW-WG--LEADHCPLA 667
WW + G +A + L+ +F+ ++ PTW W L + PLA
Sbjct: 69 GTWWIYVSMNTYGGASPLLAIVLLLAFFAALAWFFALPTWLWARWLRRNEAPLA 122
>FVR2_MOUSE (Q91X85) Feline leukemia virus subgroup C
receptor-related protein 2 (Calcium-chelate transporter)
(CCT)
Length = 551
Score = 32.0 bits (71), Expect = 5.5
Identities = 52/265 (19%), Positives = 96/265 (35%), Gaps = 53/265 (20%)
Query: 205 IGSFGLDWNTVAGFLGSPLAVPGF-----------AIINIMAGFVLYMYVLIPIAYWNNV 253
+ FG GFL P+ VP + I+ G ++++L+ I +
Sbjct: 240 VAVFGNQLGIAIGFLVPPVLVPNIKDPEKLAYHISIMFYIIGGVATFLFILVIIVFKEKP 299
Query: 254 YDAKKFPLISSHTFDSTGATY--NITRILNTKTFDIDMESYNNYSKIYLSVTFAFQYGLS 311
S+ +T A+Y +I R+ + N + + L +T+ G +
Sbjct: 300 KHPPSRAQSLSYALATTDASYLSSIVRL------------FKNLNFVLLVITYGLNAG-A 346
Query: 312 FAALTATISHVVLFH--GEMIVQMWKKTTTSLKNQLGDVHTRIMKKNYEQVPEWWFVAIL 369
F AL+ ++ +V+ H GE + T + G + + I + E V +
Sbjct: 347 FYALSTLLNRMVILHFPGEEVNAGRIGLTIVIAGMFGAMISGIWLDKSKTYKETTLVVYI 406
Query: 370 ILMVMMALLACEGFGKQLQLPWWGILLSLSIALVFT--LPIGVIEATTSARTGLNVITEL 427
+ +V M + F L W + + ++ T LP+G E
Sbjct: 407 MTLVGMVVYT---FTLNLNHLWIVFITAGTLGFFMTGYLPLGF---------------EF 448
Query: 428 VIGFTYP-----GKPLANVAFKTYG 447
+ FTYP L NV+ + +G
Sbjct: 449 AVEFTYPESEGVSSGLLNVSAQVFG 473
>CFTR_MOUSE (P26361) Cystic fibrosis transmembrane conductance
regulator (CFTR) (cAMP-dependent chloride channel)
Length = 1476
Score = 32.0 bits (71), Expect = 5.5
Identities = 13/39 (33%), Positives = 23/39 (58%)
Query: 156 FFLVFVASFAYYIVPGYFFQAISTVSFFCLIWKDSITAQ 194
FF+VF++ Y ++ G + I T FC++ + S+T Q
Sbjct: 315 FFVVFLSVLPYTVINGIVLRKIFTTISFCIVLRMSVTRQ 353
>YEIJ_ECOLI (P33021) Hypothetical transport protein yeiJ
Length = 416
Score = 31.6 bits (70), Expect = 7.2
Identities = 27/112 (24%), Positives = 49/112 (43%), Gaps = 17/112 (15%)
Query: 17 VSAQIITLPLGKLMAATLPIKT--IQVPFTSLTFSLNPGSFSVKEHVLISIFATSGSSGV 74
++A ++ +P G L A L T QV F +L+F+ P ++ ++ ++GV
Sbjct: 203 LAASLMAIPGGILFARLLSPATESSQVSFNNLSFTETPPKSIIEAAATGAMTGLKIAAGV 262
Query: 75 YAI-----NIITIVKG----------FYHRNIHPIAAYLLALSTQILGYGWA 111
+ II ++ G F H ++ I YLLA ++G W+
Sbjct: 263 ATVVMAFVAIIALINGIIGGVGGWFGFEHASLESILGYLLAPLAWVMGVDWS 314
>Y424_LISMO (Q8Y9U6) Putative sugar uptake protein lmo0424
Length = 285
Score = 31.6 bits (70), Expect = 7.2
Identities = 64/312 (20%), Positives = 126/312 (39%), Gaps = 60/312 (19%)
Query: 94 IAAYLLALSTQILGYGWAGIFRRFLVDSPYMWWPEILVQVSLFRAFHEKEKRPKGGTSRL 153
++ YL+AL +LG+G+ I +P E L+ S+ S L
Sbjct: 1 MSIYLIAL-LPVLGWGFMPIIANLRKSTP----EEQLLGTSI---------------SAL 40
Query: 154 QFFFLVF--------VASFAYYIVPGYFFQAISTVSFFCLIWKDSITAQQIGSGMKGLGI 205
F ++F V SF + G F+ + F + A I +G + +G
Sbjct: 41 LFALILFWILSPEITVLSFIVSFISGIFWSFGQLLQFKGIAASSVAKAMPISNGTQLVGA 100
Query: 206 GSFGL----DWNTVAGFLGSPLAVPGFAIINIMAGFV-----------LYMY-VLIPIAY 249
F + +W TV + +AV I +M GF ++Y ++I ++
Sbjct: 101 TLFAVLVFREWQTVTAVIIGVVAVILILIGVVMTGFQKRGNHITESVSFHVYGIVILSSF 160
Query: 250 WNNVYDAKKFPLISSHTFDSTGATYNITRILNTKTFDIDMESYNNYSKIYLSVTFAFQYG 309
+ +Y ++++ FD TG + + + + T I + + +VTF G
Sbjct: 161 FLTLY------VVTNQLFDVTGFSIILPQAIGMLTCAIGINLAKKTAISRKNVTFNLMTG 214
Query: 310 LSFAALTATISHVVLFHGEMIVQMWKKTTTSLKNQLGDVHT---RIMKKNYEQVPEWWFV 366
LS+ +I+++ +F ++ + T+ S+ V T ++ K + EW F+
Sbjct: 215 LSW-----SIANLGMFLATAVLGV--ATSFSISQACVIVATIGGILIFKQKKSPLEWTFI 267
Query: 367 AILILMVMMALL 378
IL++M+ ++
Sbjct: 268 LSGILLIMVGVV 279
>NU4M_SHEEP (O78755) NADH-ubiquinone oxidoreductase chain 4 (EC
1.6.5.3)
Length = 459
Score = 31.6 bits (70), Expect = 7.2
Identities = 28/118 (23%), Positives = 57/118 (47%), Gaps = 13/118 (11%)
Query: 387 LQLPWWGILLSLSIALVFTLPIGVIEATTSARTGLNVITELV-IGFTYPGKPLANVAFKT 445
+ L WG++++ SI L T +I ++ + L ++ L+ ++Y G +A
Sbjct: 262 IMLSLWGMIMTSSICLRQTDLKSLIAYSSVSHMALVIVAILIQTPWSYMGATALMIA--- 318
Query: 446 YGHTSMVQALGFLGDFKLGHYMKIPPKSMFIVQLVGTVVSLSVHFGTAWWLLTSIENI 503
+G TS + L +Y ++ ++M + + + T++ L AWWLL S+ N+
Sbjct: 319 HGLTSSM-----LFCLANSNYERVHSRTMILARGLQTLLPLMA----AWWLLASLTNL 367
>YG22_PYRAB (Q9UY86) Hypothetical UPF0290 protein PYRAB16220
Length = 171
Score = 31.2 bits (69), Expect = 9.4
Identities = 27/96 (28%), Positives = 43/96 (44%), Gaps = 11/96 (11%)
Query: 161 VASFAYYIVPGYFFQAISTVSFFCLIWKDSITAQQIGSGMKGL-----GIGSFGLD-WNT 214
V + + + PGY+ V L+ ++ IGS +K G + GLD W
Sbjct: 65 VGTIQHLMFPGYYGSLKLAVGVAFLLSLGALVGDLIGSFIKRRLNMPRGYPAVGLDQW-- 122
Query: 215 VAGFLGSPLAVPGFAIINIMAGFVLYMYVLIPIAYW 250
GFL S L + + I G VL++ V+ P+ +W
Sbjct: 123 --GFLISALCF-AYPLRTIPTGEVLFLLVVTPVIHW 155
>TRP3_PSEAM (O93267) Trypsinogen-like protein 3 precursor
Length = 256
Score = 31.2 bits (69), Expect = 9.4
Identities = 19/52 (36%), Positives = 26/52 (49%), Gaps = 1/52 (1%)
Query: 451 MVQALGFLGDFKLGHYMKIPPKSM-FIVQLVGTVVSLSVHFGTAWWLLTSIE 501
+V ALG G LG Y + PP S + V L +S S WW++TS +
Sbjct: 5 LVLALGLAGASPLGEYKECPPHSRPWQVNLHDGKMSCSGALIDRWWIVTSFD 56
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.327 0.142 0.460
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 80,987,545
Number of Sequences: 164201
Number of extensions: 3509700
Number of successful extensions: 9780
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 20
Number of HSP's that attempted gapping in prelim test: 9749
Number of HSP's gapped (non-prelim): 33
length of query: 683
length of database: 59,974,054
effective HSP length: 117
effective length of query: 566
effective length of database: 40,762,537
effective search space: 23071595942
effective search space used: 23071595942
T: 11
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 69 (31.2 bits)
Medicago: description of AC143341.6


