

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143339.11 + phase: 0
(678 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
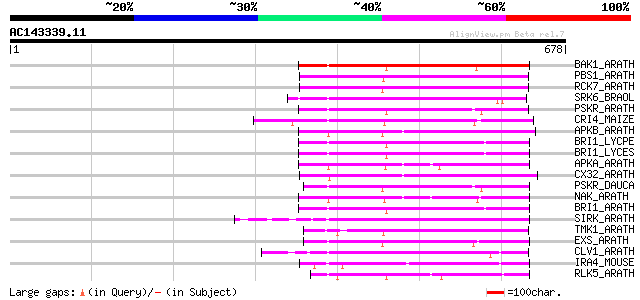
Score E
Sequences producing significant alignments: (bits) Value
BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated rec... 216 2e-55
PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC 2.7... 204 5e-52
RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase RLC... 194 5e-49
SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase rec... 194 6e-49
PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (... 188 5e-47
CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4 pr... 187 8e-47
APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor ... 184 7e-46
BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.... 183 1e-45
BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precurso... 183 1e-45
APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor ... 181 4e-45
CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx3... 180 1e-44
PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.... 179 2e-44
NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK ... 179 2e-44
BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC ... 179 2e-44
SIRK_ARATH (O64483) Senescence-induced receptor-like serine/thre... 176 1e-43
TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precur... 174 7e-43
EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase E... 171 6e-42
CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (... 166 2e-40
IRA4_MOUSE (Q8R4K2) Interleukin-1 receptor-associated kinase-4 (... 165 4e-40
RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC... 164 7e-40
>BAK1_ARATH (Q94F62) BRASSINOSTEROID INSENSITIVE 1-associated
receptor kinase 1 precursor (EC 2.7.1.37)
(BRI1-associated receptor kinase 1) (Somatic
embryogenesis receptor-like kinase 3)
Length = 615
Score = 216 bits (550), Expect = 2e-55
Identities = 121/291 (41%), Positives = 180/291 (61%), Gaps = 9/291 (3%)
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFAD-FENSERVFIN 411
+RF EL A++ F + +LGRGG+G+VYKG L+ G +VAVKR+ + + E F
Sbjct: 275 KRFSLRELQVASDNFSNKNILGRGGFGKVYKGRLAD-GTLVAVKRLKEERTQGGELQFQT 333
Query: 412 EVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS---LAWEVRYKV 468
EV +IS +HRNL++ G+C E LLV+ YM NGS+ + L +S L W R ++
Sbjct: 334 EVEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPESQPPLDWPKRQRI 393
Query: 469 ALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVV 528
ALG A L YLHD + ++HRD+K+AN+LLD +F +GDFG+AKL+D T V
Sbjct: 394 ALGSARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTTAVR 453
Query: 529 GTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQ----DGDFHVPLMNWVWGLY 584
GT G++APEY++ G++S+++D++ +G++ LEL TG+R F D V L++WV GL
Sbjct: 454 GTIGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVMLLDWVKGLL 513
Query: 585 VEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
E L D L + E++ L+ V L CT S+ ERPK EV+++L+
Sbjct: 514 KEKKLEALVDVDLQGNYKDEEVEQLIQVALLCTQSSPMERPKMSEVVRMLE 564
>PBS1_ARATH (Q9FE20) Serine/threonine-protein kinase PBS1 (EC
2.7.1.37) (AvrPphB susceptible protein 1)
Length = 456
Score = 204 bits (520), Expect = 5e-52
Identities = 120/287 (41%), Positives = 164/287 (56%), Gaps = 7/287 (2%)
Query: 355 FEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVR 414
F + EL AAT F D LG GG+G+VYKG L G++VAVK++ + R F+ EV
Sbjct: 74 FAFRELAAATMNFHPDTFLGEGGFGRVYKGRLDSTGQVVAVKQLDRNGLQGNREFLVEVL 133
Query: 415 IISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFG---DKKSLAWEVRYKVALG 471
++S L H NLV IG+C + D+ LLV+E+MP GSL+ HL DK++L W +R K+A G
Sbjct: 134 MLSLLHHPNLVNLIGYCADGDQRLLVYEFMPLGSLEDHLHDLPPDKEALDWNMRMKIAAG 193
Query: 472 VANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQ-RTDVVGT 530
A L +LHD A V++RD KS+N+LLD F KL DFG+AKL ++ T V+GT
Sbjct: 194 AAKGLEFLHDKANPPVIYRDFKSSNILLDEGFHPKLSDFGLAKLGPTGDKSHVSTRVMGT 253
Query: 531 YGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFH--VPLMNWVWGLYVE-G 587
YGY APEY G+ + +SD+YSFG+V LEL TGR+ H L+ W L+ +
Sbjct: 254 YGYCAPEYAMTGQLTVKSDVYSFGVVFLELITGRKAIDSEMPHGEQNLVAWARPLFNDRR 313
Query: 588 NLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVL 634
+ AD +L F + L V C RP +V+ L
Sbjct: 314 KFIKLADPRLKGRFPTRALYQALAVASMCIQEQAATRPLIADVVTAL 360
>RCK7_ARATH (Q9LQQ8) Putative serine/threonine-protein kinase
RLCKVII (EC 2.7.1.37)
Length = 423
Score = 194 bits (494), Expect = 5e-49
Identities = 112/287 (39%), Positives = 158/287 (55%), Gaps = 7/287 (2%)
Query: 355 FEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVR 414
F + EL AT F D LG GG+G+V+KG + L ++VA+K++ + R F+ EV
Sbjct: 91 FTFQELAEATGNFRSDCFLGEGGFGKVFKGTIEKLDQVVAIKQLDRNGVQGIREFVVEVL 150
Query: 415 IISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLF---GDKKSLAWEVRYKVALG 471
+S H NLV+ IG+C E D+ LLV+EYMP GSL+ HL KK L W R K+A G
Sbjct: 151 TLSLADHPNLVKLIGFCAEGDQRLLVYEYMPQGSLEDHLHVLPSGKKPLDWNTRMKIAAG 210
Query: 472 VANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQ-RTDVVGT 530
A L YLHD V++RD+K +N+LL D+ KL DFG+AK+ +T T V+GT
Sbjct: 211 AARGLEYLHDRMTPPVIYRDLKCSNILLGEDYQPKLSDFGLAKVGPSGDKTHVSTRVMGT 270
Query: 531 YGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVP--LMNWVWGLYVE-G 587
YGY AP+Y G+ + +SD+YSFG+V LEL TGR+ + L+ W L+ +
Sbjct: 271 YGYCAPDYAMTGQLTFKSDIYSFGVVLLELITGRKAIDNTKTRKDQNLVGWARPLFKDRR 330
Query: 588 NLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVL 634
N D L ++ V + L + C RP +V+ L
Sbjct: 331 NFPKMVDPLLQGQYPVRGLYQALAISAMCVQEQPTMRPVVSDVVLAL 377
>SRK6_BRAOL (Q09092) Putative serine/threonine-protein kinase
receptor precursor (EC 2.7.1.37) (S-receptor kinase)
(SRK)
Length = 849
Score = 194 bits (493), Expect = 6e-49
Identities = 113/304 (37%), Positives = 173/304 (56%), Gaps = 14/304 (4%)
Query: 340 FPKKFDLDKATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIF 399
F ++ ++ +P E +V AT F LG+GG+G VYKG L GK +AVKR+
Sbjct: 502 FSGEYKFEELELPL-IEMETVVKATENFSSCNKLGQGGFGIVYKGRLLD-GKEIAVKRLS 559
Query: 400 ADFENSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGD--K 457
F+NEV +I+RL H NLVQ +G C E DE +L++EY+ N SLD++LFG +
Sbjct: 560 KTSVQGTDEFMNEVTLIARLQHINLVQVLGCCIEGDEKMLIYEYLENLSLDSYLFGKTRR 619
Query: 458 KSLAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVD 517
L W R+ + GVA L YLH D+ ++HRD+K +N+LLD + K+ DFGMA++ +
Sbjct: 620 SKLNWNERFDITNGVARGLLYLHQDSRFRIIHRDLKVSNILLDKNMIPKISDFGMARIFE 679
Query: 518 -PMLRTQRTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGR--RFFQDGDFHV 574
VVGTYGY++PEY G S++SD++SFG++ LE+ +G+ R F + D+
Sbjct: 680 RDETEANTMKVVGTYGYMSPEYAMYGIFSEKSDVFSFGVIVLEIVSGKKNRGFYNLDYEN 739
Query: 575 PLMNWVWGLYVEGNLMCAAD----EKLNME---FDVSEMKSLLIVGLWCTHSNDKERPKA 627
L+++VW + EG + D + L+ + F E+ + +GL C + RP
Sbjct: 740 DLLSYVWSRWKEGRALEIVDPVIVDSLSSQPSIFQPQEVLKCIQIGLLCVQELAEHRPAM 799
Query: 628 YEVI 631
V+
Sbjct: 800 SSVV 803
>PSKR_ARATH (Q9ZVR7) Putative phytosulfokine receptor precursor (EC
2.7.1.37) (Phytosulfokine LRR receptor kinase)
Length = 1008
Score = 188 bits (477), Expect = 5e-47
Identities = 110/290 (37%), Positives = 160/290 (54%), Gaps = 12/290 (4%)
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINE 412
+ Y +L+ +TN FD ++G GG+G VYK L GK VA+K++ D ER F E
Sbjct: 720 KELSYDDLLDSTNSFDQANIIGCGGFGMVYKATLPD-GKKVAIKKLSGDCGQIEREFEAE 778
Query: 413 VRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS---LAWEVRYKVA 469
V +SR H NLV G+C +++ LL++ YM NGSLD L L W+ R ++A
Sbjct: 779 VETLSRAQHPNLVLLRGFCFYKNDRLLIYSYMENGSLDYWLHERNDGPALLKWKTRLRIA 838
Query: 470 LGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVG 529
G A L YLH+ + +LHRDIKS+N+LLD +F++ L DFG+A+L+ P TD+VG
Sbjct: 839 QGAAKGLLYLHEGCDPHILHRDIKSSNILLDENFNSHLADFGLARLMSPYETHVSTDLVG 898
Query: 530 TYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVP-----LMNWVWGLY 584
T GY+ PEY A+ + D+YSFG+V LEL T +R D P L++WV +
Sbjct: 899 TLGYIPPEYGQASVATYKGDVYSFGVVLLELLTDKR---PVDMCKPKGCRDLISWVVKMK 955
Query: 585 VEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVL 634
E D + + + EM +L + C N K+RP +++ L
Sbjct: 956 HESRASEVFDPLIYSKENDKEMFRVLEIACLCLSENPKQRPTTQQLVSWL 1005
>CRI4_MAIZE (O24585) Putative receptor protein kinase CRINKLY4
precursor (EC 2.7.1.-)
Length = 901
Score = 187 bits (475), Expect = 8e-47
Identities = 122/366 (33%), Positives = 192/366 (52%), Gaps = 29/366 (7%)
Query: 299 VVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKF-----------DLD 347
+ VA IV +++ L S+ + V+ K ++C E + + D++
Sbjct: 424 IFVAEIVFAVVLVLSVSVTTCLYVRHKLRHCQCSNRELRLAKSTAYSFRKDNMKIQPDME 483
Query: 348 KATIPR--RFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIF--ADFE 403
I R F Y EL AT GF +D +G+G + V+KG L G +VAVKR +D +
Sbjct: 484 DLKIRRAQEFSYEELEQATGGFSEDSQVGKGSFSCVFKGILRD-GTVVAVKRAIKASDVK 542
Query: 404 NSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGD----KKS 459
S + F NE+ ++SRL H +L+ +G+C + E LLV+E+M +GSL HL G KK
Sbjct: 543 KSSKEFHNELDLLSRLNHAHLLNLLGYCEDGSERLLVYEFMAHGSLYQHLHGKDPNLKKR 602
Query: 460 LAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPM 519
L W R +A+ A + YLH A V+HRDIKS+N+L+D D + ++ DFG++ L
Sbjct: 603 LNWARRVTIAVQAARGIEYLHGYACPPVIHRDIKSSNILIDEDHNARVADFGLSILGPAD 662
Query: 520 LRTQRTDV-VGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRF----FQDGDFHV 574
T +++ GT GYL PEY + +SD+YSFG+V LE+ +GR+ F++G+
Sbjct: 663 SGTPLSELPAGTLGYLDPEYYRLHYLTTKSDVYSFGVVLLEILSGRKAIDMQFEEGN--- 719
Query: 575 PLMNWVWGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVL 634
++ W L G++ D L+ D+ +K + V C K+RP +V L
Sbjct: 720 -IVEWAVPLIKAGDIFAILDPVLSPPSDLEALKKIASVACKCVRMRGKDRPSMDKVTTAL 778
Query: 635 QLEMAL 640
+ +AL
Sbjct: 779 EHALAL 784
>APKB_ARATH (P46573) Protein kinase APK1B, chloroplast precursor (EC
2.7.1.-)
Length = 412
Score = 184 bits (467), Expect = 7e-46
Identities = 111/306 (36%), Positives = 170/306 (55%), Gaps = 17/306 (5%)
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSY---------LGKIVAVKRIFADFE 403
+ F ++EL AAT F D +LG GG+G V+KG + G ++AVK++ D
Sbjct: 55 KSFTFAELKAATRNFRPDSVLGEGGFGSVFKGWIDEQTLTASKPGTGVVIAVKKLNQDGW 114
Query: 404 NSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLF---GDKKSL 460
+ ++ EV + + H NLV+ IG+C E + LLV+E+MP GSL+ HLF + L
Sbjct: 115 QGHQEWLAEVNYLGQFSHPNLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGSYFQPL 174
Query: 461 AWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPML 520
+W +R KVALG A L +LH +AE V++RD K++N+LLD++++ KL DFG+AK
Sbjct: 175 SWTLRLKVALGAAKGLAFLH-NAETSVIYRDFKTSNILLDSEYNAKLSDFGLAKDGPTGD 233
Query: 521 RTQ-RTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDG--DFHVPLM 577
++ T ++GTYGY APEY+ G + +SD+YS+G+V LE+ +GRR L+
Sbjct: 234 KSHVSTRIMGTYGYAAPEYLATGHLTTKSDVYSYGVVLLEVLSGRRAVDKNRPPGEQKLV 293
Query: 578 NWVWGLYV-EGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQL 636
W L + L D +L ++ + E + + L C K RP EV+ L+
Sbjct: 294 EWARPLLANKRKLFRVIDNRLQDQYSMEEACKVATLALRCLTFEIKLRPNMNEVVSHLEH 353
Query: 637 EMALPE 642
L E
Sbjct: 354 IQTLNE 359
>BRI1_LYCPE (Q8L899) Systemin receptor SR160 precursor (EC 2.7.1.37)
(Brassinosteroid LRR receptor kinase)
Length = 1207
Score = 183 bits (465), Expect = 1e-45
Identities = 107/290 (36%), Positives = 168/290 (57%), Gaps = 9/290 (3%)
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINE 412
R+ +++L+ ATNGF +D ++G GG+G VYK L G +VA+K++ +R F E
Sbjct: 874 RKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKD-GSVVAIKKLIHVSGQGDREFTAE 932
Query: 413 VRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS---LAWEVRYKVA 469
+ I ++ HRNLV +G+C +E LLV+EYM GSL+ L KK+ L W R K+A
Sbjct: 933 METIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKTGIKLNWPARRKIA 992
Query: 470 LGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPM-LRTQRTDVV 528
+G A L +LH + ++HRD+KS+NVLLD + ++ DFGMA+L+ M + +
Sbjct: 993 IGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLA 1052
Query: 529 GTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDF-HVPLMNWVWGLYVEG 587
GT GY+ PEY R S + D+YS+G+V LEL TG++ DF L+ WV L+ +G
Sbjct: 1053 GTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFGDNNLVGWV-KLHAKG 1111
Query: 588 NLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDK--ERPKAYEVIKVLQ 635
+ D +L E E++ L + + C +D+ +RP +V+ + +
Sbjct: 1112 KITDVFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAMFK 1161
>BRI1_LYCES (Q8GUQ5) Brassinosteroid LRR receptor kinase precursor (EC
2.7.1.37) (tBRI1) (Altered brassinolide sensitivity 1)
(Systemin receptor SR160)
Length = 1207
Score = 183 bits (464), Expect = 1e-45
Identities = 107/290 (36%), Positives = 167/290 (56%), Gaps = 9/290 (3%)
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINE 412
R+ +++L+ ATNGF +D ++G GG+G VYK L G +VA+K++ +R F E
Sbjct: 874 RKLTFADLLEATNGFHNDSLVGSGGFGDVYKAQLKD-GSVVAIKKLIHVSGQGDREFTAE 932
Query: 413 VRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKK---SLAWEVRYKVA 469
+ I ++ HRNLV +G+C +E LLV+EYM GSL+ L KK L W R K+A
Sbjct: 933 METIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDRKKIGIKLNWPARRKIA 992
Query: 470 LGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPM-LRTQRTDVV 528
+G A L +LH + ++HRD+KS+NVLLD + ++ DFGMA+L+ M + +
Sbjct: 993 IGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLA 1052
Query: 529 GTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDF-HVPLMNWVWGLYVEG 587
GT GY+ PEY R S + D+YS+G+V LEL TG++ DF L+ WV L+ +G
Sbjct: 1053 GTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKQPTDSADFGDNNLVGWV-KLHAKG 1111
Query: 588 NLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDK--ERPKAYEVIKVLQ 635
+ D +L E E++ L + + C +D+ +RP +V+ + +
Sbjct: 1112 KITDVFDRELLKEDASIEIELLQHLKVACACLDDRHWKRPTMIQVMAMFK 1161
>APKA_ARATH (Q06548) Protein kinase APK1A, chloroplast precursor (EC
2.7.1.-)
Length = 410
Score = 181 bits (460), Expect = 4e-45
Identities = 109/301 (36%), Positives = 171/301 (56%), Gaps = 21/301 (6%)
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSY---------LGKIVAVKRIFADFE 403
+ F ++EL +AT F D +LG GG+G V+KG + G ++AVK++ D
Sbjct: 54 KSFSFAELKSATRNFRPDSVLGEGGFGCVFKGWIDEKSLTASRPGTGLVIAVKKLNQDGW 113
Query: 404 NSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDK---KSL 460
+ ++ EV + + HR+LV+ IG+C E + LLV+E+MP GSL+ HLF + L
Sbjct: 114 QGHQEWLAEVNYLGQFSHRHLVKLIGYCLEDEHRLLVYEFMPRGSLENHLFRRGLYFQPL 173
Query: 461 AWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPML 520
+W++R KVALG A L +LH +E V++RD K++N+LLD++++ KL DFG+AK D +
Sbjct: 174 SWKLRLKVALGAAKGLAFLH-SSETRVIYRDFKTSNILLDSEYNAKLSDFGLAK--DGPI 230
Query: 521 RTQ---RTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDG--DFHVP 575
+ T V+GT+GY APEY+ G + +SD+YSFG+V LEL +GRR
Sbjct: 231 GDKSHVSTRVMGTHGYAAPEYLATGHLTTKSDVYSFGVVLLELLSGRRAVDKNRPSGERN 290
Query: 576 LMNWVWGLYV-EGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVL 634
L+ W V + + D +L ++ + E + + L C + K RP EV+ L
Sbjct: 291 LVEWAKPYLVNKRKIFRVIDNRLQDQYSMEEACKVATLSLRCLTTEIKLRPNMSEVVSHL 350
Query: 635 Q 635
+
Sbjct: 351 E 351
>CX32_ARATH (P27450) Probable serine/threonine-protein kinase Cx32,
chloroplast precursor (EC 2.7.1.37)
Length = 419
Score = 180 bits (457), Expect = 1e-44
Identities = 104/303 (34%), Positives = 170/303 (55%), Gaps = 14/303 (4%)
Query: 355 FEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYL---------GKIVAVKRIFADFENS 405
+ + +L AT F D MLG+GG+G+VY+G + G IVA+KR+ ++
Sbjct: 74 YNFLDLKTATKNFKPDSMLGQGGFGKVYRGWVDATTLAPSRVGSGMIVAIKRLNSESVQG 133
Query: 406 ERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVR 465
+ +EV + L HRNLV+ +G+C E ELLLV+E+MP GSL++HLF W++R
Sbjct: 134 FAEWRSEVNFLGMLSHRNLVKLLGYCREDKELLLVYEFMPKGSLESHLFRRNDPFPWDLR 193
Query: 466 YKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQ-R 524
K+ +G A L +LH ++ V++RD K++N+LLD+++ KL DFG+AKL ++
Sbjct: 194 IKIVIGAARGLAFLH-SLQREVIYRDFKASNILLDSNYDAKLSDFGLAKLGPADEKSHVT 252
Query: 525 TDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATG--RRFFQDGDFHVPLMNWVW- 581
T ++GTYGY APEY+ G +SD+++FG+V LE+ TG + L++W+
Sbjct: 253 TRIMGTYGYAAPEYMATGHLYVKSDVFAFGVVLLEIMTGLTAHNTKRPRGQESLVDWLRP 312
Query: 582 GLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALP 641
L + + D+ + ++ + + L C + K RP EV++VL+ L
Sbjct: 313 ELSNKHRVKQIMDKGIKGQYTTKVATEMARITLSCIEPDPKNRPHMKEVVEVLEHIQGLN 372
Query: 642 ELP 644
+P
Sbjct: 373 VVP 375
>PSKR_DAUCA (Q8LPB4) Phytosulfokine receptor precursor (EC 2.7.1.37)
(Phytosulfokine LRR receptor kinase)
Length = 1021
Score = 179 bits (455), Expect = 2e-44
Identities = 108/285 (37%), Positives = 155/285 (53%), Gaps = 12/285 (4%)
Query: 359 ELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISR 418
+++ +T+ F+ ++G GG+G VYK L G VA+KR+ D +R F EV +SR
Sbjct: 735 DILKSTSSFNQANIIGCGGFGLVYKATLPD-GTKVAIKRLSGDTGQMDREFQAEVETLSR 793
Query: 419 LIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLF---GDKKSLAWEVRYKVALGVANA 475
H NLV +G+C+ +++ LL++ YM NGSLD L SL W+ R ++A G A
Sbjct: 794 AQHPNLVHLLGYCNYKNDKLLIYSYMDNGSLDYWLHEKVDGPPSLDWKTRLRIARGAAEG 853
Query: 476 LRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLA 535
L YLH E +LHRDIKS+N+LL F L DFG+A+L+ P TD+VGT GY+
Sbjct: 854 LAYLHQSCEPHILHRDIKSSNILLSDTFVAHLADFGLARLILPYDTHVTTDLVGTLGYIP 913
Query: 536 PEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVP-----LMNWVWGLYVEGNLM 590
PEY A+ + D+YSFG+V LEL TGRR D P L++WV + E
Sbjct: 914 PEYGQASVATYKGDVYSFGVVLLELLTGRR---PMDVCKPRGSRDLISWVLQMKTEKRES 970
Query: 591 CAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
D + + EM +L + C N K RP +++ L+
Sbjct: 971 EIFDPFIYDKDHAEEMLLVLEIACRCLGENPKTRPTTQQLVSWLE 1015
>NAK_ARATH (P43293) Probable serine/threonine-protein kinase NAK (EC
2.7.1.37)
Length = 389
Score = 179 bits (455), Expect = 2e-44
Identities = 108/302 (35%), Positives = 174/302 (56%), Gaps = 23/302 (7%)
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSY---------LGKIVAVKRIFADFE 403
+ F SEL +AT F D ++G GG+G V+KG + G ++AVKR+ +
Sbjct: 54 KNFSLSELKSATRNFRPDSVVGEGGFGCVFKGWIDESSLAPSKPGTGIVIAVKRLNQEGF 113
Query: 404 NSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGD---KKSL 460
R ++ E+ + +L H NLV+ IG+C E++ LLV+E+M GSL+ HLF + L
Sbjct: 114 QGHREWLAEINYLGQLDHPNLVKLIGYCLEEEHRLLVYEFMTRGSLENHLFRRGTFYQPL 173
Query: 461 AWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPML 520
+W R ++ALG A L +LH +A+ V++RD K++N+LLD++++ KL DFG+A+ PM
Sbjct: 174 SWNTRVRMALGAARGLAFLH-NAQPQVIYRDFKASNILLDSNYNAKLSDFGLAR-DGPMG 231
Query: 521 RTQR--TDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQD----GDFHV 574
T V+GT GY APEY+ G S +SD+YSFG+V LEL +GRR G+ +
Sbjct: 232 DNSHVSTRVMGTQGYAAPEYLATGHLSVKSDVYSFGVVLLELLSGRRAIDKNQPVGEHN- 290
Query: 575 PLMNWVWG-LYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKV 633
L++W L + L+ D +L ++ ++ + ++ L C + K RP E++K
Sbjct: 291 -LVDWARPYLTNKRRLLRVMDPRLQGQYSLTRALKIAVLALDCISIDAKSRPTMNEIVKT 349
Query: 634 LQ 635
++
Sbjct: 350 ME 351
>BRI1_ARATH (O22476) BRASSINOSTEROID INSENSITIVE 1 precursor (EC
2.7.1.37) (AtBRI1) (Brassinosteroid LRR receptor kinase)
Length = 1196
Score = 179 bits (455), Expect = 2e-44
Identities = 107/290 (36%), Positives = 163/290 (55%), Gaps = 9/290 (3%)
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINE 412
R+ +++L+ ATNGF +D ++G GG+G VYK L G VA+K++ +R F+ E
Sbjct: 869 RKLTFADLLQATNGFHNDSLIGSGGFGDVYKAILKD-GSAVAIKKLIHVSGQGDREFMAE 927
Query: 413 VRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS---LAWEVRYKVA 469
+ I ++ HRNLV +G+C DE LLV+E+M GSL+ L KK+ L W R K+A
Sbjct: 928 METIGKIKHRNLVPLLGYCKVGDERLLVYEFMKYGSLEDVLHDPKKAGVKLNWSTRRKIA 987
Query: 470 LGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPM-LRTQRTDVV 528
+G A L +LH + ++HRD+KS+NVLLD + ++ DFGMA+L+ M + +
Sbjct: 988 IGSARGLAFLHHNCSPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVSTLA 1047
Query: 529 GTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDF-HVPLMNWVWGLYVEG 587
GT GY+ PEY R S + D+YS+G+V LEL TG+R DF L+ WV + +
Sbjct: 1048 GTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKRPTDSPDFGDNNLVGWV-KQHAKL 1106
Query: 588 NLMCAADEKLNMEFDVSEMKSL--LIVGLWCTHSNDKERPKAYEVIKVLQ 635
+ D +L E E++ L L V + C RP +V+ + +
Sbjct: 1107 RISDVFDPELMKEDPALEIELLQHLKVAVACLDDRAWRRPTMVQVMAMFK 1156
>SIRK_ARATH (O64483) Senescence-induced receptor-like
serine/threonine kinase precursor (FLG22-induced
receptor-like kinase 1)
Length = 876
Score = 176 bits (447), Expect = 1e-43
Identities = 116/365 (31%), Positives = 190/365 (51%), Gaps = 28/365 (7%)
Query: 275 LEDSNSTTSLVEGNDGKGSLKTVIVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEG-L 333
L DS S T K K ++ V+V +I+V L A + +R +K G L
Sbjct: 503 LSDSCSNT--------KKKNKNGYIIPLVVVGIIVVLLTAL----ALFRRFKKKQQRGTL 550
Query: 334 DEYGIPFPKKFDLDKATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIV 393
E P T R F+YSE+V TN F+ R++G+GG+G+VY G ++ G+ V
Sbjct: 551 GERNGPLK--------TAKRYFKYSEVVNITNNFE--RVIGKGGFGKVYHGVIN--GEQV 598
Query: 394 AVKRIFADFENSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHL 453
AVK + + + F EV ++ R+ H NL +G+C+E + ++L++EYM N +L +L
Sbjct: 599 AVKVLSEESAQGYKEFRAEVDLLMRVHHTNLTSLVGYCNEINHMVLIYEYMANENLGDYL 658
Query: 454 FGDKKS-LAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGM 512
G + L+WE R K++L A L YLH+ + ++HRD+K N+LL+ K+ DFG+
Sbjct: 659 AGKRSFILSWEERLKISLDAAQGLEYLHNGCKPPIVHRDVKPTNILLNEKLQAKMADFGL 718
Query: 513 AKLVDPMLRTQ-RTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGD 571
++ Q T V G+ GYL PEY + + +++SD+YS G+V LE+ TG+
Sbjct: 719 SRSFSVEGSGQISTVVAGSIGYLDPEYYSTRQMNEKSDVYSLGVVLLEVITGQPAIASSK 778
Query: 572 FH-VPLMNWVWGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEV 630
V + + V + G++ D++L +DV + + L CT +RP +V
Sbjct: 779 TEKVHISDHVRSILANGDIRGIVDQRLRERYDVGSAWKMSEIALACTEHTSAQRPTMSQV 838
Query: 631 IKVLQ 635
+ L+
Sbjct: 839 VMELK 843
>TMK1_ARATH (P43298) Putative receptor protein kinase TMK1 precursor
(EC 2.7.1.-)
Length = 942
Score = 174 bits (441), Expect = 7e-43
Identities = 102/293 (34%), Positives = 164/293 (55%), Gaps = 26/293 (8%)
Query: 360 LVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRI---------FADFENSERVFI 410
L + TN F D +LG GG+G VYKG L + G +AVKR+ FA+F++
Sbjct: 581 LRSVTNNFSSDNILGSGGFGVVYKGEL-HDGTKIAVKRMENGVIAGKGFAEFKS------ 633
Query: 411 NEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFG----DKKSLAWEVRY 466
E+ +++++ HR+LV +G+C + +E LLV+EYMP G+L HLF K L W+ R
Sbjct: 634 -EIAVLTKVRHRHLVTLLGYCLDGNEKLLVYEYMPQGTLSRHLFEWSEEGLKPLLWKQRL 692
Query: 467 KVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTD 526
+AL VA + YLH A Q +HRD+K +N+LL D K+ DFG+ +L + T
Sbjct: 693 TLALDVARGVEYLHGLAHQSFIHRDLKPSNILLGDDMRAKVADFGLVRLAPEGKGSIETR 752
Query: 527 VVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDG--DFHVPLMNWVWGLY 584
+ GT+GYLAPEY GR + + D+YSFG++ +EL TGR+ + + + L++W +Y
Sbjct: 753 IAGTFGYLAPEYAVTGRVTTKVDVYSFGVILMELITGRKSLDESQPEESIHLVSWFKRMY 812
Query: 585 V--EGNLMCAADEKLNM-EFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVL 634
+ E + A D +++ E ++ + ++ + C +RP + +L
Sbjct: 813 INKEASFKKAIDTTIDLDEETLASVHTVAELAGHCCAREPYQRPDMGHAVNIL 865
>EXS_ARATH (Q9LYN8) Leucine-rich repeat receptor protein kinase EXS
precursor (EC 2.7.1.37) (Extra sporogenous cells protein)
(EXCESS MICROSPOROCYTES1 protein)
Length = 1192
Score = 171 bits (433), Expect = 6e-42
Identities = 101/284 (35%), Positives = 153/284 (53%), Gaps = 9/284 (3%)
Query: 359 ELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISR 418
++V AT+ F ++G GG+G VYK L K VAVK++ R F+ E+ + +
Sbjct: 909 DIVEATDHFSKKNIIGDGGFGTVYKACLPG-EKTVAVKKLSEAKTQGNREFMAEMETLGK 967
Query: 419 LIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHL---FGDKKSLAWEVRYKVALGVANA 475
+ H NLV +G+C +E LLV+EYM NGSLD L G + L W R K+A+G A
Sbjct: 968 VKHPNLVSLLGYCSFSEEKLLVYEYMVNGSLDHWLRNQTGMLEVLDWSKRLKIAVGAARG 1027
Query: 476 LRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLA 535
L +LH ++HRDIK++N+LLD DF K+ DFG+A+L+ T + GT+GY+
Sbjct: 1028 LAFLHHGFIPHIIHRDIKASNILLDGDFEPKVADFGLARLISACESHVSTVIAGTFGYIP 1087
Query: 536 PEYINGGRASKESDMYSFGIVALELATGRR----FFQDGDFHVPLMNWVWGLYVEGNLMC 591
PEY RA+ + D+YSFG++ LEL TG+ F++ + L+ W +G +
Sbjct: 1088 PEYGQSARATTKGDVYSFGVILLELVTGKEPTGPDFKESE-GGNLVGWAIQKINQGKAVD 1146
Query: 592 AADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
D L + LL + + C +RP +V+K L+
Sbjct: 1147 VIDPLLVSVALKNSQLRLLQIAMLCLAETPAKRPNMLDVLKALK 1190
>CLV1_ARATH (Q9SYQ8) Receptor protein kinase CLAVATA1 precursor (EC
2.7.1.-)
Length = 980
Score = 166 bits (419), Expect = 2e-40
Identities = 106/338 (31%), Positives = 182/338 (53%), Gaps = 22/338 (6%)
Query: 308 ILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPRRFEY-SELVAATNG 366
I++ +IA+I G +++ + ++ ++ + + K T ++ ++ SE V
Sbjct: 641 IVITVIAAITGLILISVAIRQMNKKKNQKSLAW-------KLTAFQKLDFKSEDVLEC-- 691
Query: 367 FDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFA-DFENSERVFINEVRIISRLIHRNLV 425
++ ++G+GG G VY+G++ VA+KR+ S+ F E++ + R+ HR++V
Sbjct: 692 LKEENIIGKGGAGIVYRGSMPN-NVDVAIKRLVGRGTGRSDHGFTAEIQTLGRIRHRHIV 750
Query: 426 QFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS-LAWEVRYKVALGVANALRYLHDDAE 484
+ +G+ +D LL++EYMPNGSL L G K L WE R++VA+ A L YLH D
Sbjct: 751 RLLGYVANKDTNLLLYEYMPNGSLGELLHGSKGGHLQWETRHRVAVEAAKGLCYLHHDCS 810
Query: 485 QCVLHRDIKSANVLLDTDFSTKLGDFGMAK-LVDPMLRTQRTDVVGTYGYLAPEYINGGR 543
+LHRD+KS N+LLD+DF + DFG+AK LVD + + G+YGY+APEY +
Sbjct: 811 PLILHRDVKSNNILLDSDFEAHVADFGLAKFLVDGAASECMSSIAGSYGYIAPEYAYTLK 870
Query: 544 ASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVWGLYVE-------GNLMCAADEK 596
++SD+YSFG+V LEL G++ + V ++ WV E ++ D +
Sbjct: 871 VDEKSDVYSFGVVLLELIAGKKPVGEFGEGVDIVRWVRNTEEEITQPSDAAIVVAIVDPR 930
Query: 597 LNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVL 634
L + ++ + + + + C RP EV+ +L
Sbjct: 931 LT-GYPLTSVIHVFKIAMMCVEEEAAARPTMREVVHML 967
>IRA4_MOUSE (Q8R4K2) Interleukin-1 receptor-associated kinase-4 (EC
2.7.1.37) (IRAK-4)
Length = 459
Score = 165 bits (417), Expect = 4e-40
Identities = 107/295 (36%), Positives = 164/295 (55%), Gaps = 21/295 (7%)
Query: 355 FEYSELVAATNGFDDD------RMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENS--- 405
F + EL + TN FD+ +G GG+G VYKG ++ IVAVK++ A E S
Sbjct: 168 FSFHELKSITNNFDEQPASAGGNRMGEGGFGVVYKGCVN--NTIVAVKKLGAMVEISTEE 225
Query: 406 -ERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHL--FGDKKSLAW 462
++ F E+++++ H NLV+ +G+ + D L LV+ YMPNGSL L L+W
Sbjct: 226 LKQQFDQEIKVMATCQHENLVELLGFSSDSDNLCLVYAYMPNGSLLDRLSCLDGTPPLSW 285
Query: 463 EVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRT 522
R KVA G AN +R+LH++ +HRDIKSAN+LLD DF+ K+ DFG+A+ + +T
Sbjct: 286 HTRCKVAQGTANGIRFLHENHH---IHRDIKSANILLDKDFTAKISDFGLARASARLAQT 342
Query: 523 QRTD-VVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVW 581
T +VGT Y+APE + G + +SD+YSFG+V LEL TG + L++
Sbjct: 343 VMTSRIVGTTAYMAPEALR-GEITPKSDIYSFGVVLLELITGLAAVDENREPQLLLDIKE 401
Query: 582 GLY-VEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
+ E + DEK++ + D + ++++ C H RP +V ++LQ
Sbjct: 402 EIEDEEKTIEDYTDEKMS-DADPASVEAMYSAASQCLHEKKNRRPDIAKVQQLLQ 455
>RLK5_ARATH (P47735) Receptor-like protein kinase 5 precursor (EC
2.7.1.37)
Length = 999
Score = 164 bits (415), Expect = 7e-40
Identities = 102/285 (35%), Positives = 158/285 (54%), Gaps = 20/285 (7%)
Query: 368 DDDRMLGRGGYGQVYKGALSYLGKIVAVKRI----------FADFENSERVFINEVRIIS 417
D+ ++G G G+VYK L G++VAVK++ ++ + VF EV +
Sbjct: 684 DEKNVIGFGSSGKVYKVELRG-GEVVAVKKLNKSVKGGDDEYSSDSLNRDVFAAEVETLG 742
Query: 418 RLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS---LAWEVRYKVALGVAN 474
+ H+++V+ C D LLV+EYMPNGSL L GD+K L W R ++AL A
Sbjct: 743 TIRHKSIVRLWCCCSSGDCKLLVYEYMPNGSLADVLHGDRKGGVVLGWPERLRIALDAAE 802
Query: 475 ALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTD----VVGT 530
L YLH D ++HRD+KS+N+LLD+D+ K+ DFG+AK V M ++ + + G+
Sbjct: 803 GLSYLHHDCVPPIVHRDVKSSNILLDSDYGAKVADFGIAK-VGQMSGSKTPEAMSGIAGS 861
Query: 531 YGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVWGLYVEGNLM 590
GY+APEY+ R +++SD+YSFG+V LEL TG++ + WV + L
Sbjct: 862 CGYIAPEYVYTLRVNEKSDIYSFGVVLLELVTGKQPTDSELGDKDMAKWVCTALDKCGLE 921
Query: 591 CAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
D KL+++F E+ ++ +GL CT RP +V+ +LQ
Sbjct: 922 PVIDPKLDLKFK-EEISKVIHIGLLCTSPLPLNRPSMRKVVIMLQ 965
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 83,198,475
Number of Sequences: 164201
Number of extensions: 3759755
Number of successful extensions: 12499
Number of sequences better than 10.0: 1696
Number of HSP's better than 10.0 without gapping: 1170
Number of HSP's successfully gapped in prelim test: 526
Number of HSP's that attempted gapping in prelim test: 8398
Number of HSP's gapped (non-prelim): 1943
length of query: 678
length of database: 59,974,054
effective HSP length: 117
effective length of query: 561
effective length of database: 40,762,537
effective search space: 22867783257
effective search space used: 22867783257
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 69 (31.2 bits)
Medicago: description of AC143339.11


