

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142143.3 + phase: 0 /pseudo
(212 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
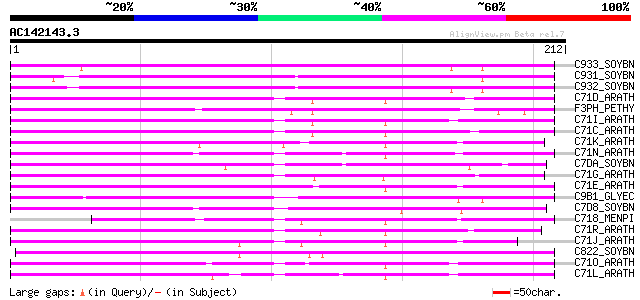
Score E
Sequences producing significant alignments: (bits) Value
C933_SOYBN (O81973) Cytochrome P450 93A3 (EC 1.14.-.-) (P450 CP5) 137 2e-32
C931_SOYBN (Q42798) Cytochrome P450 93A1 (EC 1.14.-.-) 129 5e-30
C932_SOYBN (Q42799) Cytochrome P450 93A2 (EC 1.14.-.-) 124 2e-28
C71D_ARATH (O49342) Cytochrome P450 71A13 (EC 1.14.-.-) 107 3e-23
F3PH_PETHY (Q9SBQ9) Flavonoid 3'-monooxygenase (EC 1.14.13.21) (... 106 3e-23
C71I_ARATH (Q9SAB6) Cytochrome P450 71A18 (EC 1.14.-.-) 104 1e-22
C71C_ARATH (O49340) Cytochrome P450 71A12 (EC 1.14.-.-) 104 1e-22
C71K_ARATH (Q9T0K2) Cytochrome P450 71A20 (EC 1.14.-.-) 104 2e-22
C71N_ARATH (Q9STL0) Cytochrome P450 71A23 (EC 1.14.-.-) 99 7e-21
C7DA_SOYBN (O48923) Cytochrome P450 71D10 (EC 1.14.-.-) 99 9e-21
C71G_ARATH (Q9FH66) Cytochrome P450 71A16 (EC 1.14.-.-) 99 9e-21
C71E_ARATH (P58045) Cytochrome P450 71A14 (EC 1.14.-.-) 99 9e-21
C9B1_GLYEC (P93149) Cytochrome P450 93B1 (EC 1.14.-.-) ((2S)-fla... 97 3e-20
C7D8_SOYBN (O81974) Cytochrome P450 71D8 (EC 1.14.-.-) (P450 CP7) 97 3e-20
C718_MENPI (Q42716) Cytochrome P450 71A8 (EC 1.14.-.-) 96 5e-20
C71R_ARATH (O65438) Cytochrome P450 71A27 (EC 1.14.-.-) 94 2e-19
C71J_ARATH (Q9T0K0) Cytochrome P450 71A19 (EC 1.14.-.-) 94 2e-19
C822_SOYBN (O81972) Cytochrome P450 82A2 (EC 1.14.-.-) (P450 CP4) 93 4e-19
C71O_ARATH (Q9STK9) Cytochrome P450 71A24 (EC 1.14.-.-) 93 4e-19
C71L_ARATH (Q9STL2) Cytochrome P450 71A21 (EC 1.14.-.-) 91 3e-18
>C933_SOYBN (O81973) Cytochrome P450 93A3 (EC 1.14.-.-) (P450 CP5)
Length = 510
Score = 137 bits (345), Expect = 2e-32
Identities = 76/215 (35%), Positives = 127/215 (58%), Gaps = 7/215 (3%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSY-FITAPYGPYWRFMKKICVTKLLSSSQLGRFMHI 59
+KTH+ FS RP + + L G F+ APYGPYW+FMKK+C+++LL L +F+ +
Sbjct: 88 LKTHEPAFSNRPANTVAVETLTYGFQDFLFAPYGPYWKFMKKLCMSELLGGHMLDQFLPV 147
Query: 60 REQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLV 119
R+ E +K +K +L + D G +F TL+NNI+ RM + T + + E ++ LV
Sbjct: 148 RQXETKKFIKRVLQKGISGEAVDFGGEFITLSNNIVSRMIVSQTSTTEDENEVEEMRKLV 207
Query: 120 REFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKE-HEERINTEDCQ 178
++ + K ++ + + L +FDL G+ K+L KI FD +L+ +K+ EER N +
Sbjct: 208 KDAAELSGKFNISDFVSFLKRFDLQGFNKRLEKIRDCFDTVLDRIIKQREEERRNKNETV 267
Query: 179 G-----DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
G DM+D+L + +E++E++L + +IKAF L
Sbjct: 268 GKREFKDMLDVLFDISEDESSEIKLNKENIKAFIL 302
>C931_SOYBN (Q42798) Cytochrome P450 93A1 (EC 1.14.-.-)
Length = 509
Score = 129 bits (324), Expect = 5e-30
Identities = 70/222 (31%), Positives = 124/222 (55%), Gaps = 20/222 (9%)
Query: 1 MKTHDLNFSYRPQFG--------SSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQ 52
+KTH++NFS RP S D+L F AP+GPYW+FMKK+C+++LLS
Sbjct: 86 LKTHEINFSNRPGQNVAVKGLAYDSQDFL-----FAFAPFGPYWKFMKKLCMSELLSGRM 140
Query: 53 LGRFMHIREQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDP 112
+ +F+ +R+QE ++ + + + D G + TL+NNI+ RM++ + N
Sbjct: 141 MDQFLPVRQQETKRFISRVFRKGVAGEAVDFGDELMTLSNNIVSRMTLSQKTSENDN-QA 199
Query: 113 EVIQCLVREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERI 172
E ++ LV + K ++ + + L FDL G+ +K+++ FD +++G +K+ +E
Sbjct: 200 EEMKKLVSNIAELMGKFNVSDFIWYLKPFDLQGFNRKIKETRDRFDVVVDGIIKQRQEER 259
Query: 173 NTEDCQG------DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
G DM+D+LL ++ +ENAE++L + +IKAF +
Sbjct: 260 RKNKETGTAKQFKDMLDVLLDMHEDENAEIKLDKKNIKAFIM 301
>C932_SOYBN (Q42799) Cytochrome P450 93A2 (EC 1.14.-.-)
Length = 502
Score = 124 bits (310), Expect = 2e-28
Identities = 69/214 (32%), Positives = 125/214 (58%), Gaps = 11/214 (5%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTH++NFS RP + +L ++ PYGP +F+KK+C+++LL L +F+ +R
Sbjct: 86 LKTHEINFSNRPGQNVAVQFLT----YVFGPYGPSVKFIKKLCMSELLGGRMLDQFLPVR 141
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
+QE +K +K +L + D G +F L+NNI+ RM+M T + E ++ LV
Sbjct: 142 QQETKKFIKRVLQKGIAGEAVDFGGEFMRLSNNIISRMTMNQTSSEDEK-QAEEMRMLVA 200
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKE-HEERINTEDCQG 179
+ + ++ + + L FDL G+ K++RK FD +L+ +K+ EER N ++ G
Sbjct: 201 DVAELMGTFNVSDFIWFLKPFDLQGFNKRIRKTRIRFDAVLDRIIKQREEERRNNKEIGG 260
Query: 180 -----DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
D++D+LL + ++++E++LT+ +IKAF +
Sbjct: 261 TRQFKDILDVLLDIGEDDSSEIKLTKENIKAFIM 294
>C71D_ARATH (O49342) Cytochrome P450 71A13 (EC 1.14.-.-)
Length = 497
Score = 107 bits (266), Expect = 3e-23
Identities = 62/211 (29%), Positives = 117/211 (55%), Gaps = 10/211 (4%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHD F+ RP+ + H + G + APYG YWR MK +C+ LL++ + F +R
Sbjct: 90 LKTHDHKFANRPRSKAVHGLMNGGRDVVFAPYGEYWRQMKSVCILNLLTNKMVESFEKVR 149
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEV--IQCL 118
E E+ +++ L S+ + S +L F TL +++ R+++G +K++ D ++
Sbjct: 150 EDEVNAMIEKLEKASSSSSSENLSELFITLPSDVTSRVALG----RKHSEDETARDLKKR 205
Query: 119 VREFMHVGAKLSMGEVLGPLGKFD-LFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDC 177
VR+ M + + +GE + L D + G+ K++++ F +++ ++EH E N
Sbjct: 206 VRQIMELLGEFPIGEYVPILAWIDGIRGFNNKIKEVSRGFSDLMDKVVQEHLEASND--- 262
Query: 178 QGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
+ D +DILL + +++N+ ++ RNDIK L
Sbjct: 263 KADFVDILLSIEKDKNSGFQVQRNDIKFMIL 293
>F3PH_PETHY (Q9SBQ9) Flavonoid 3'-monooxygenase (EC 1.14.13.21)
(Flavonoid 3'-hydroxylase) (Cytochrome P450 75B2)
Length = 512
Score = 106 bits (265), Expect = 3e-23
Identities = 70/217 (32%), Positives = 109/217 (49%), Gaps = 16/217 (7%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHD NFS RP + Y + APYGP WR ++KIC L S+ L F H+R
Sbjct: 90 LKTHDANFSSRPPNSGAEHMAYNYQDLVFAPYGPRWRMLRKICSVHLFSTKALDDFRHVR 149
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFK--KYNIDPEV--IQ 116
+ E++ L ++L S + LG T N L R+ +G F ++DP+ +
Sbjct: 150 QDEVKTLTRAL--ASAGQKPVKLGQLLNVCTTNALARVMLGKRVFADGSGDVDPQAAEFK 207
Query: 117 CLVREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTED 176
+V E M V ++G+ + L D+ G K++K+ FD L L+EH+ +I
Sbjct: 208 SMVVEMMVVAGVFNIGDFIPQLNWLDIQGVAAKMKKLHARFDAFLTDILEEHKGKI---- 263
Query: 177 CQGDMMDIL--LQVYRNENAE---VRLTRNDIKAFFL 208
G+M D+L L +N++A+ +LT +IKA L
Sbjct: 264 -FGEMKDLLSTLISLKNDDADNDGGKLTDTEIKALLL 299
>C71I_ARATH (Q9SAB6) Cytochrome P450 71A18 (EC 1.14.-.-)
Length = 497
Score = 104 bits (260), Expect = 1e-22
Identities = 61/211 (28%), Positives = 114/211 (53%), Gaps = 10/211 (4%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHDL F+ RP+ + H + G + PYG YWR MK +C+ LL++ + F +R
Sbjct: 90 LKTHDLKFANRPKSKAVHGLMNGGRDVVFGPYGEYWRQMKSVCILNLLTNKMVASFEKVR 149
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEV--IQCL 118
E+E+ +++ L S + + +L F TLT+++ R+S+G KKY D ++
Sbjct: 150 EEEVNAMMEKLEKASCSSSAENLSELFVTLTSDVTSRVSLG----KKYWEDETAGGLKKR 205
Query: 119 VREFMHVGAKLSMGEVLGPLGKFD-LFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDC 177
VR+ M + + +G+ + L D + G+ K+ ++ + ++E ++EH + +
Sbjct: 206 VRQIMELLREFPIGDYVPALAWIDRINGFNSKIVEVSRAYSDLMEKVVQEH---LEAGEH 262
Query: 178 QGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
+ D ++ILL + + +N ++ RNDIK L
Sbjct: 263 KADFVNILLSIEKEKNNGFKVQRNDIKFMIL 293
>C71C_ARATH (O49340) Cytochrome P450 71A12 (EC 1.14.-.-)
Length = 497
Score = 104 bits (260), Expect = 1e-22
Identities = 61/211 (28%), Positives = 116/211 (54%), Gaps = 10/211 (4%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHDL F+ RP+ + H + G + PYG YWR MK +C+ LL++ + F IR
Sbjct: 90 LKTHDLKFANRPRSKAVHGLMNGGRDVVFGPYGEYWRQMKSVCILNLLTNKMVASFEKIR 149
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEV--IQCL 118
E+EL +++K L S+ + S +L F TL +++ R+++G +K++ D ++
Sbjct: 150 EEELNEMIKKLEKASSSSSSENLSELFVTLPSDVTSRIALG----RKHSEDETARDLKKR 205
Query: 119 VREFMHVGAKLSMGEVLGPLGKFD-LFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDC 177
VR+ M + + +G+ + L D + G+ +++++ F +++ ++EH E N ++
Sbjct: 206 VRQIMELLGEFPIGDYVPALAWIDRINGFNARIKEVSQGFSDLMDKVVQEHLEAGNHKE- 264
Query: 178 QGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
D +DILL + ++ + R+DIK L
Sbjct: 265 --DFVDILLSIESEKSIGFQAQRDDIKFMIL 293
>C71K_ARATH (Q9T0K2) Cytochrome P450 71A20 (EC 1.14.-.-)
Length = 495
Score = 104 bits (259), Expect = 2e-22
Identities = 67/208 (32%), Positives = 112/208 (53%), Gaps = 9/208 (4%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MKTHDL + RP+ L G APYG YWR MK IC+ LL++ + + IR
Sbjct: 87 MKTHDLVCANRPKTKVVDKILSGGRDVAFAPYGEYWRQMKSICIQNLLNNKMVRSYEKIR 146
Query: 61 EQELEKLLKSL--LICSNENRSTDLGLDFTTLTNNILCRMSMGTT-CFKKYNIDPEVIQC 117
E+E++++++ L CS+ +L TLTN+I+CR+++G KK ID ++
Sbjct: 147 EEEIKRMIEKLEKASCSSSPSPVNLSQILMTLTNDIICRVALGRKYSGKKDGID---VEN 203
Query: 118 LVREFMHVGAKLSMGEVLGPLGKFD-LFGYGKKLRKIVGEFDKILEGFLKEHEERINTED 176
+VR F + + +GE + L D + G K+ + FD+ LE +KEHEE ++
Sbjct: 204 IVRTFAALLGEFPVGEYIPSLSWIDRIRGLDHKMEVVDKRFDEFLERVVKEHEEA--DKE 261
Query: 177 CQGDMMDILLQVYRNENAEVRLTRNDIK 204
+ D++D LL + ++ + L ++ +K
Sbjct: 262 TRSDLVDKLLTIQSDKTGQFELEKSALK 289
>C71N_ARATH (Q9STL0) Cytochrome P450 71A23 (EC 1.14.-.-)
Length = 483
Score = 99.0 bits (245), Expect = 7e-21
Identities = 61/209 (29%), Positives = 113/209 (53%), Gaps = 11/209 (5%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHD F+ RP+ LYK +APYG YWR MK + V LLS+ + F +R
Sbjct: 86 LKTHDRVFASRPRSKIFEKLLYKSRNMASAPYGEYWRQMKSVSVLHLLSNKMVRSFQDVR 145
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
++E+ ++++ I + ++ +L ++LTN+++CR+++G +KY + + + + R
Sbjct: 146 QEEITLMMET--IRKSSSKPVNLSKILSSLTNDVICRVALG----RKYGVGTDFKELIDR 199
Query: 121 EFMHVGAKLSMGEVLGPLGKFD-LFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQG 179
+G ++G + L D + G +L K +FDK+LE +++HE+ + +
Sbjct: 200 LMRQLGT-FTIGSYVPWLAWTDWVSGLEARLEKTANDFDKLLERIVQDHED---GDGDKT 255
Query: 180 DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
D +D+LL R+++ + R IKA L
Sbjct: 256 DFVDVLLAAQRDKSFGFDIDRLSIKAIVL 284
>C7DA_SOYBN (O48923) Cytochrome P450 71D10 (EC 1.14.-.-)
Length = 510
Score = 98.6 bits (244), Expect = 9e-21
Identities = 65/210 (30%), Positives = 111/210 (51%), Gaps = 12/210 (5%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MKTHDLNFS RP F S Y GS + + +G YWR ++KIC +LL++ ++ F IR
Sbjct: 102 MKTHDLNFSDRPDFVLSRIVSYNGSGIVFSQHGDYWRQLRKICTVELLTAKRVQSFRSIR 161
Query: 61 EQELEKLLKSLLICSNENRST--DLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCL 118
E+E+ +L+K + ++E + +L ++T I R + G KK I +
Sbjct: 162 EEEVAELVKKIAATASEEGGSIFNLTQSIYSMTFGIAARAAFG----KKSRYQQVFISNM 217
Query: 119 VREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINT---E 175
++ M +G S+ ++ F + G KL K+ D++L+ + EH+ R +
Sbjct: 218 HKQLMLLGG-FSVADLYPSSRVFQMMGATGKLEKVHRVTDRVLQDIIDEHKNRNRSSEER 276
Query: 176 DCQGDMMDILLQVYRNENAEVRLTRNDIKA 205
+ D++D+LL+ + +E RLT ++IKA
Sbjct: 277 EAVEDLVDVLLKF--QKESEFRLTDDNIKA 304
>C71G_ARATH (Q9FH66) Cytochrome P450 71A16 (EC 1.14.-.-)
Length = 497
Score = 98.6 bits (244), Expect = 9e-21
Identities = 64/207 (30%), Positives = 109/207 (51%), Gaps = 8/207 (3%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MKTHDL F+ RP S+H G + APYG YWR +K +C LLS+ + R
Sbjct: 89 MKTHDLKFANRPITKSAHKISNGGRDLVFAPYGEYWRNVKSLCTIHLLSNKMVQSSEKRR 148
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVI--QCL 118
E+E+ L+++L S + S +L T + ++I+ ++ +G KKY+ + I + +
Sbjct: 149 EEEITLLMETLEEASLSSSSVNLSKLITNMVSDIMGKVVLG----KKYSGEEGTIDVKTI 204
Query: 119 VREFMHVGAKLSMGEVLGPLGKF-DLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDC 177
+ F+ +GE + L + G KL KI +F +E L+EHE+ ++
Sbjct: 205 TKSFLDAVGLSPVGEYIPSLAWIGKITGSDGKLEKITKQFGDFIEKVLQEHEDTTADKET 264
Query: 178 QGDMMDILLQVYRNENAEVRLTRNDIK 204
D +D+LL + R+E A+ +L ++D+K
Sbjct: 265 P-DFVDMLLTIQRDETAQCQLDKSDLK 290
>C71E_ARATH (P58045) Cytochrome P450 71A14 (EC 1.14.-.-)
Length = 497
Score = 98.6 bits (244), Expect = 9e-21
Identities = 61/209 (29%), Positives = 110/209 (52%), Gaps = 5/209 (2%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MKTHDL + RPQ G + +PYG YWR +K +C+ LL+ ++ F +R
Sbjct: 90 MKTHDLKVANRPQLKVVEKIFNGGREMVFSPYGEYWRQIKSVCIVNLLNKKKVQSFEKVR 149
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
E+E+ ++++ + S+++ +L TLT+++ R+S+G K+ ++ IQ +R
Sbjct: 150 EEEISEMMERVEKASSDSSPLNLSELLLTLTSDVTSRVSLGRKYSKEESMSDFKIQ--MR 207
Query: 121 EFMHVGAKLSMGEVLGPLGKFD-LFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQG 179
+ + +GE + L D L G +K ++ F ++E L+EH + T+
Sbjct: 208 KITELVGGFPVGEYIPCLAWIDKLRGVDEKAEEVSKAFGDLMEKVLQEHLDA--TDKPTL 265
Query: 180 DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
D +D+LL + R+E V++ R+DIK L
Sbjct: 266 DFVDVLLSLERHERNGVQIRRSDIKFLIL 294
>C9B1_GLYEC (P93149) Cytochrome P450 93B1 (EC 1.14.-.-)
((2S)-flavanone 2-hydroxylase) (Licodione synthase)
(Flavone synthase II) (CYP GE-5)
Length = 523
Score = 97.1 bits (240), Expect = 3e-20
Identities = 61/220 (27%), Positives = 112/220 (50%), Gaps = 22/220 (10%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
++T++L F+ R + + Y+ S APYG YWRF+KK+ + +LL S + F H+R
Sbjct: 90 LQTNELAFNCRIESTAVKKLTYESSLAF-APYGDYWRFIKKLSMNELLGSRSINNFQHLR 148
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
QE +LL+ L + + ++ + LTNN++ M +G + E + +VR
Sbjct: 149 AQETHQLLRLLSNRARAFEAVNITEELLKLTNNVISIMMVG---------EAEEARDVVR 199
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEE------RINT 174
+ + + ++ + + K DL G+GK++ + FD ++E + + E+ R
Sbjct: 200 DVTEIFGEFNVSDFIWLFKKMDLQGFGKRIEDLFQRFDTLVERIISKREQTRKDRRRNGK 259
Query: 175 EDCQG------DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
+ QG D +DILL +EN+E+++ R IKA +
Sbjct: 260 KGEQGSGDGIRDFLDILLDCTEDENSEIKIQRVHIKALIM 299
>C7D8_SOYBN (O81974) Cytochrome P450 71D8 (EC 1.14.-.-) (P450 CP7)
Length = 504
Score = 97.1 bits (240), Expect = 3e-20
Identities = 68/213 (31%), Positives = 109/213 (50%), Gaps = 15/213 (7%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MKTHD++F RPQ + +Y + APYG YWR ++KIC +LLS+ ++ F HIR
Sbjct: 93 MKTHDVHFVQRPQLLAPQFMVYGATDIAFAPYGDYWRQIRKICTLELLSAKRVQSFSHIR 152
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
+ E +KL++S I S+ DL +L + R + G K N D + LVR
Sbjct: 153 QDENKKLIQS--IHSSAGSPIDLSGKLFSLLGTTVSRAAFG-----KENDDQDEFMSLVR 205
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGK-KLRKIVGEFDKILEGFLKEHEER-------I 172
+ + + + ++ L L K K+ + DKILE L++H E+
Sbjct: 206 KAITMTGGFEVDDMFPSLKPLHLLTRQKAKVEHVHQRADKILEDILRKHMEKRTRVKEGN 265
Query: 173 NTEDCQGDMMDILLQVYRNENAEVRLTRNDIKA 205
+E Q D++D+LL++ + + EV +T +IKA
Sbjct: 266 GSEAEQEDLVDVLLRLKESGSLEVPMTMENIKA 298
>C718_MENPI (Q42716) Cytochrome P450 71A8 (EC 1.14.-.-)
Length = 502
Score = 96.3 bits (238), Expect = 5e-20
Identities = 59/181 (32%), Positives = 104/181 (56%), Gaps = 12/181 (6%)
Query: 32 YGPYWRFMKKICVTKLLSSSQLGRFMHIREQELEKLLKSLLICSNENRSTDLGLDFTTLT 91
YG YWR +K ICV +LLS+ ++ F +RE+E E L+K + + + + +L FT LT
Sbjct: 131 YGEYWRQLKTICVVQLLSNKRVQSFRSVREEETELLMKKI---GDSSGNVNLSHMFTQLT 187
Query: 92 NNILCRMSMGTTCFKKYNI---DPEVIQCLVREFMHVGAKLSMGEVLGPLGKFD-LFGYG 147
N+++CR ++G +KY + E ++REF+ + +S+G+ + L + + G+
Sbjct: 188 NDVVCRSAIG----RKYGAGDENGEKFLEILREFLELLGAISIGDFVPSLWWINRINGFD 243
Query: 148 KKLRKIVGEFDKILEGFLKEHEERINTEDCQGDMMDILLQVYRNENAEVRLTRNDIKAFF 207
+++ +I E D+ LE + E E + + +DILL++YRN +A V + R+ IKA
Sbjct: 244 RRVDRIAKEMDEFLEKVIHERLEN-PAAKAEENFVDILLEIYRNNSAGVSIDRDSIKAII 302
Query: 208 L 208
L
Sbjct: 303 L 303
>C71R_ARATH (O65438) Cytochrome P450 71A27 (EC 1.14.-.-)
Length = 499
Score = 94.4 bits (233), Expect = 2e-19
Identities = 60/206 (29%), Positives = 108/206 (52%), Gaps = 9/206 (4%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MKTHDL F+ RP+ + + ++ G I PYG W+ MK + V LL++ + F ++R
Sbjct: 90 MKTHDLKFANRPKSKAINIFMEGGRDIIFGPYGEDWKSMKSLGVVHLLNNKMVRSFENLR 149
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQC--L 118
E+E++ + + L S+ + S +L TLTN+I+CR+++G +KYN + I L
Sbjct: 150 EEEIKVMTEKLEEASSSSSSVNLSKLLMTLTNDIICRITLG----RKYNEEEGGIDIKNL 205
Query: 119 VREFMHVGAKLSMGEVLGPLGKFD-LFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDC 177
V K G+ + L D + G K++ I + D L+ ++EH + + E
Sbjct: 206 VMTSSEFFGKFFFGDFIPSLAWIDWISGIDDKMKDINNKLDCFLDSMVQEHVDADHKE-- 263
Query: 178 QGDMMDILLQVYRNENAEVRLTRNDI 203
D +D+LL + +++ + R+D+
Sbjct: 264 PSDFIDMLLLIQKDKTKRFKFDRSDL 289
>C71J_ARATH (Q9T0K0) Cytochrome P450 71A19 (EC 1.14.-.-)
Length = 490
Score = 94.4 bits (233), Expect = 2e-19
Identities = 61/199 (30%), Positives = 109/199 (54%), Gaps = 11/199 (5%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KT+D+ + RP+ L G APYG YW+ MK IC+ LLS+ + + IR
Sbjct: 90 LKTYDVICANRPKTKVIDKILRGGRDVAFAPYGEYWKQMKSICIQNLLSNKMVRSYKKIR 149
Query: 61 EQELEKLLKSLLICSNENRSTDLGLD--FTTLTNNILCRMSMGTTCFKKYNI--DPEVIQ 116
E E++ +++ + S+ + + + L F TLTN+I+CR ++G +KY+ D ++
Sbjct: 150 EDEIKLMIEKVENASSCSPPSPVNLSQLFMTLTNDIICRAALG----RKYSSKEDGIDVE 205
Query: 117 CLVREFMHVGAKLSMGEVLGPLGKFD-LFGYGKKLRKIVGEFDKILEGFLKEHEERINTE 175
+VR F + + +GE + L D + G K+ ++ FD+ LE +KEHE+ +
Sbjct: 206 NIVRAFSALVGEFPIGEYIPSLSWIDKIRGQDHKMEEVDKRFDEFLERVVKEHEDA--NK 263
Query: 176 DCQGDMMDILLQVYRNENA 194
D + D++D LL + +++A
Sbjct: 264 DTRSDLVDTLLTIQSDKSA 282
>C822_SOYBN (O81972) Cytochrome P450 82A2 (EC 1.14.-.-) (P450 CP4)
Length = 522
Score = 93.2 bits (230), Expect = 4e-19
Identities = 62/216 (28%), Positives = 105/216 (47%), Gaps = 10/216 (4%)
Query: 3 THDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIREQ 62
T+D+ S P S++ Y S + APYGPYWR ++KI +++ LS S++ + H+R
Sbjct: 98 TNDIAVSSLPDLISANLLCYNRSMIVVAPYGPYWRQLRKILMSEFLSPSRVEQLHHVRVS 157
Query: 63 ELEKLLKSLLICSNENRSTDLGLD-------FTTLTNNILCRMSMGTTCFKKYNIDPE-V 114
E++ + L N++ G F+ L N++ RM G F D E
Sbjct: 158 EVQSSITELFRDWRSNKNVQSGFATVELKQWFSLLVFNMILRMVCGKRYFSASTSDDEKA 217
Query: 115 IQCL--VREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERI 172
+C+ V EF+ + A ++G+ + L FD GY +R+ E D+I+ +L EH ++
Sbjct: 218 NRCVKAVDEFVRLAATFTVGDAIPYLRWFDFGGYENDMRETGKELDEIIGEWLDEHRQKR 277
Query: 173 NTEDCQGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
+ D+M +LL + + E IK+F L
Sbjct: 278 KMGENVQDLMSVLLSLLEGKTIEGMNVDIVIKSFVL 313
>C71O_ARATH (Q9STK9) Cytochrome P450 71A24 (EC 1.14.-.-)
Length = 486
Score = 93.2 bits (230), Expect = 4e-19
Identities = 60/209 (28%), Positives = 106/209 (50%), Gaps = 11/209 (5%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHD F+ RP+ Y G APYG YWR MK +CV L S+ + F +R
Sbjct: 88 LKTHDRVFASRPRSKIFDKIFYNGRDVALAPYGEYWRQMKSVCVLHLFSNKMVRSFRDVR 147
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
++E+ +++ + I S + +L LTNN++CR+++G +KY + + L++
Sbjct: 148 QEEISLMIEKIRISS--SLRINLSEILVNLTNNVICRVALG----RKYGGKTD-FKDLMK 200
Query: 121 EFMHVGAKLSMGEVLGPLGKFD-LFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQG 179
+ + S+G + L D + G +L KI + D+ LE +++H ++ + +
Sbjct: 201 RLTRLLGEFSVGSYVSWLAWIDWIRGLDGQLIKISNDLDEFLERVVQDH---VDGDGHKN 257
Query: 180 DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
D +D LL + R ++ + R IKA L
Sbjct: 258 DFVDFLLTIEREKSVGFEIDRLSIKAIIL 286
>C71L_ARATH (Q9STL2) Cytochrome P450 71A21 (EC 1.14.-.-)
Length = 490
Score = 90.5 bits (223), Expect = 3e-18
Identities = 62/211 (29%), Positives = 115/211 (54%), Gaps = 15/211 (7%)
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHD F+ RP+ Y G APYG YWR +K +CV +LLS+ + F ++R
Sbjct: 89 LKTHDRVFASRPRSKLFEKLFYDGRDVAFAPYGEYWRQIKSVCVLRLLSNKMVTSFRNVR 148
Query: 61 EQELEKLLKSLLICSN--ENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCL 118
++E+ +++ + S+ N S LG +LTN+++ R+++G +KY+ + + + +
Sbjct: 149 QEEISLMMEKIQKSSSLQVNVSELLG----SLTNDVISRIALG----RKYSGETDSKELM 200
Query: 119 VREFMHVGAKLSMGEVLGPLGKFD-LFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDC 177
R M +G + S+G + LG D + G +L K + D+ LE +++H ++ +
Sbjct: 201 KRLMMLMG-EFSVGTYVPWLGWIDWISGLDGQLNKTGNDLDEFLEKVVQDH---VDGDGQ 256
Query: 178 QGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
+ D +D+LL++ R ++ + R IKA L
Sbjct: 257 RTDFVDVLLRIQREKSIGFEIDRLCIKAIVL 287
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.142 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,902,252
Number of Sequences: 164201
Number of extensions: 1020868
Number of successful extensions: 3246
Number of sequences better than 10.0: 136
Number of HSP's better than 10.0 without gapping: 110
Number of HSP's successfully gapped in prelim test: 26
Number of HSP's that attempted gapping in prelim test: 3053
Number of HSP's gapped (non-prelim): 139
length of query: 212
length of database: 59,974,054
effective HSP length: 106
effective length of query: 106
effective length of database: 42,568,748
effective search space: 4512287288
effective search space used: 4512287288
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC142143.3


