

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142143.13 - phase: 0
(259 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
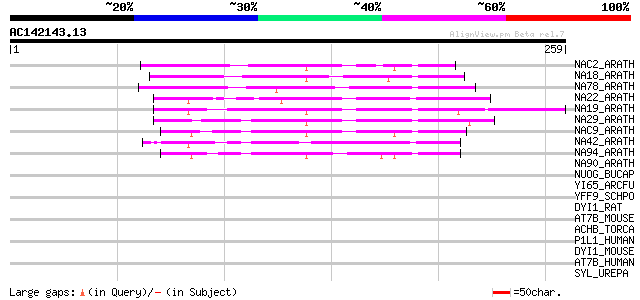
Score E
Sequences producing significant alignments: (bits) Value
NAC2_ARATH (Q39013) NAC-domain containing protein 2 (ANAC002) 65 2e-10
NA18_ARATH (Q9ZNU2) NAC-domain containing protein 18 (ANAC018) (... 59 2e-08
NA78_ARATH (Q84K00) NAC-domain containing protein 78 (ANAC078) 56 8e-08
NA22_ARATH (Q84TE6) NAC-domain containing protein 21/22 (ANAC021... 55 2e-07
NA19_ARATH (Q9C932) NAC-domain containing protein 19 (ANAC019) (... 53 6e-07
NA29_ARATH (O49255) NAC-domain containing protein 29 (ANAC029) (... 51 3e-06
NAC9_ARATH (Q9ZVH0) Putative NAC-domain containing protein 9 (AN... 49 9e-06
NA42_ARATH (Q9SK55) Putative NAC-domain containing protein 42 (A... 47 4e-05
NA94_ARATH (Q9FIW5) Putative NAC-domain containing protein 94 (A... 44 3e-04
NA90_ARATH (Q9FMR3) NAC-domain containing protein 90 (ANAC090) 40 0.004
NUOG_BUCAP (Q8K9Y2) NADH-quinone oxidoreductase chain G (EC 1.6.... 34 0.40
YI65_ARCFU (O28414) Hypothetical protein AF1865 32 1.2
YFF9_SCHPO (O14066) Hypothetical serine-rich protein C1687.09 in... 32 1.5
DYI1_RAT (Q63100) Dynein intermediate chain 1, cytosolic (DH IC-... 30 5.8
AT7B_MOUSE (Q64446) Copper-transporting ATPase 2 (EC 3.6.3.4) (C... 30 5.8
ACHB_TORCA (P02712) Acetylcholine receptor protein, beta chain p... 30 5.8
P1L1_HUMAN (Q8TDX9) Polycystic kidney disease 1-like 1 protein (... 30 7.6
DYI1_MOUSE (O88485) Dynein intermediate chain 1, cytosolic (DH I... 30 7.6
AT7B_HUMAN (P35670) Copper-transporting ATPase 2 (EC 3.6.3.4) (C... 30 7.6
SYL_UREPA (Q9PQC0) Leucyl-tRNA synthetase (EC 6.1.1.4) (Leucine-... 29 9.9
>NAC2_ARATH (Q39013) NAC-domain containing protein 2 (ANAC002)
Length = 289
Score = 65.1 bits (157), Expect = 2e-10
Identities = 47/150 (31%), Positives = 69/150 (45%), Gaps = 22/150 (14%)
Query: 62 INELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKL 121
++EL LP G +F P D+E++ H + P+I E + P +L
Sbjct: 1 MSELLQLPPGFRFHPTDEELVMHYLCRKCASQSIAVPIIAEI--------DLYKYDPWEL 52
Query: 122 PGVKKDGQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMK-- 178
PG+ G+ +F P + Y G+R R + W TG +P+ G+ K
Sbjct: 53 PGLALYGEKEWYFFSPRDRKYPNGSRPNRSAGSGY------WKATGADKPI---GLPKPV 103
Query: 179 GFKKILVLYTNYGRQKKPEKTNWVMHQYHL 208
G KK LV Y G+ K EKTNW+MH+Y L
Sbjct: 104 GIKKALVFYA--GKAPKGEKTNWIMHEYRL 131
>NA18_ARATH (Q9ZNU2) NAC-domain containing protein 18 (ANAC018) (NO
APICAL MERISTEM protein) (AtNAM)
Length = 320
Score = 58.5 bits (140), Expect = 2e-08
Identities = 44/152 (28%), Positives = 65/152 (41%), Gaps = 21/152 (13%)
Query: 66 PGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVK 125
P LP G +F P D+E++ H + VP +I + + P +LP
Sbjct: 15 PNLPPGFRFHPTDEELVIHYLKRKADSVPLPVAII--------ADVDLYKFDPWELPAKA 66
Query: 126 KDGQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDG----VMKGF 180
G+ +F P + Y G R R + W TG +PVI G G
Sbjct: 67 SFGEQEWYFFSPRDRKYPNGARPNR------AATSGYWKATGTDKPVISTGGGGSKKVGV 120
Query: 181 KKILVLYTNYGRQKKPEKTNWVMHQYHLGSNE 212
KK LV Y+ G+ K K++W+MH+Y L N+
Sbjct: 121 KKALVFYS--GKPPKGVKSDWIMHEYRLTDNK 150
>NA78_ARATH (Q84K00) NAC-domain containing protein 78 (ANAC078)
Length = 567
Score = 56.2 bits (134), Expect = 8e-08
Identities = 43/160 (26%), Positives = 69/160 (42%), Gaps = 19/160 (11%)
Query: 61 GINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEK 120
G + L G +F P D+E++++ + + + P I I + P
Sbjct: 2 GRGSVTSLAPGFRFHPTDEELVRYYLKRKVCNKPFKFDAISV--------TDIYKSEPWD 53
Query: 121 LPG---VKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVM 177
LP +K +FF K Y+ G++ R + + W TGK R + +
Sbjct: 54 LPDKSKLKSRDLEWYFFSMLDKKYSNGSKTNRATE------KGYWKTTGKDREIRNGSRV 107
Query: 178 KGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDG 217
G KK LV + GR + E+TNWVMH+Y L + +K G
Sbjct: 108 VGMKKTLVYHK--GRAPRGERTNWVMHEYRLSDEDLKKAG 145
>NA22_ARATH (Q84TE6) NAC-domain containing protein 21/22 (ANAC021)
(ANAC022)
Length = 324
Score = 54.7 bits (130), Expect = 2e-07
Identities = 44/159 (27%), Positives = 73/159 (45%), Gaps = 18/159 (11%)
Query: 68 LPAGVKFDPNDQEIL-QHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVK- 125
LP G +F P D E++ +L + L + + PL+ ++ + C P +P +
Sbjct: 19 LPPGFRFHPKDDELVCDYLMRRSLHNNHR-PPLV-----LIQVDLNKC--EPWDIPKMAC 70
Query: 126 KDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILV 185
G+ +F+ + + Y TG R R T W TGK R ++ G + G +K LV
Sbjct: 71 VGGKDWYFYSQRDRKYATGLRTNRATATGY------WKATGKDRTILRKGKLVGMRKTLV 124
Query: 186 LYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSKK 224
Y GR + KT+WVMH++ L + + L K+
Sbjct: 125 FY--QGRAPRGRKTDWVMHEFRLQGSHHPPNHSLSSPKE 161
>NA19_ARATH (Q9C932) NAC-domain containing protein 19 (ANAC019)
(ANAC) (Abscicic-acid-responsive NAC)
Length = 317
Score = 53.1 bits (126), Expect = 6e-07
Identities = 55/198 (27%), Positives = 84/198 (41%), Gaps = 24/198 (12%)
Query: 68 LPAGVKFDPNDQEIL-QHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKK 126
LP G +F P D+E++ Q+L K +F L E + P LP
Sbjct: 14 LPPGFRFYPTDEELMVQYLCRKAAGY---------DFSLQLIAEIDLYKFDPWVLPNKAL 64
Query: 127 DGQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILV 185
G+ +F P + Y G+R R + W TG + + +G G KK LV
Sbjct: 65 FGEKEWYFFSPRDRKYPNGSRPNRVAGSGY------WKATGTDKIISTEGQRVGIKKALV 118
Query: 186 LYTNYGRQKKPEKTNWVMHQYHL----GSNEEEKDGELVVSKKILEYLSSSERRLACNSV 241
Y G+ K KTNW+MH+Y L N K + V+ +I + SS+++++ N +
Sbjct: 119 FYI--GKAPKGTKTNWIMHEYRLIEPSRRNGSTKLDDWVLC-RIYKKQSSAQKQVYDNGI 175
Query: 242 IRFMYSSVNFFLSLMASS 259
S N S +SS
Sbjct: 176 ANAREFSNNGTSSTTSSS 193
>NA29_ARATH (O49255) NAC-domain containing protein 29 (ANAC029)
(NAC2) (NAC-LIKE, ACTIVATED BY AP3/PI protein) (NAP)
Length = 268
Score = 50.8 bits (120), Expect = 3e-06
Identities = 43/163 (26%), Positives = 71/163 (43%), Gaps = 20/163 (12%)
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKD 127
LP G +F P D+E++ + L + P IP ++ I P +LP +
Sbjct: 9 LPPGFRFHPTDEELIVYY----LRNQTMSKPCPVSIIPEVD----IYKFDPWQLPEKTEF 60
Query: 128 GQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILVL 186
G+ +F P + Y G R R + W TG + + G KK LV
Sbjct: 61 GENEWYFFSPRERKYPNGVRPNRAAVSGY------WKATGTDKAIHSGSSNVGVKKALVF 114
Query: 187 YTNYGRQKKPEKTNWVMHQYHLGSNEE---EKDGELVVSKKIL 226
Y GR K KT+W+MH+Y L + + +++G + + + +L
Sbjct: 115 YK--GRPPKGIKTDWIMHEYRLHDSRKASTKRNGSMRLDEWVL 155
>NAC9_ARATH (Q9ZVH0) Putative NAC-domain containing protein 9
(ANAC009)
Length = 418
Score = 49.3 bits (116), Expect = 9e-06
Identities = 46/147 (31%), Positives = 65/147 (43%), Gaps = 21/147 (14%)
Query: 71 GVKFDPNDQEILQ-HLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKDGQ 129
G +F P D+E++ +L+ KV + +PL E I L+ I P LP G+
Sbjct: 19 GFRFHPTDEELVSFYLKRKV-----QHNPLSIELIRQLD----IYKYDPWDLPKFAMTGE 69
Query: 130 IRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMK--GFKKILVL 186
+F+ P + Y +R R W TG RP+ K G KK LV
Sbjct: 70 KEWYFYCPRDRKYRNSSRPNRVTGAGF------WKATGTDRPIYSSEGNKCIGLKKSLVF 123
Query: 187 YTNYGRQKKPEKTNWVMHQYHLGSNEE 213
Y GR K KT+W+MH++ L S E
Sbjct: 124 YK--GRAAKGVKTDWMMHEFRLPSLSE 148
>NA42_ARATH (Q9SK55) Putative NAC-domain containing protein 42
(ANAC042)
Length = 275
Score = 47.4 bits (111), Expect = 4e-05
Identities = 48/149 (32%), Positives = 65/149 (43%), Gaps = 19/149 (12%)
Query: 63 NELPGLPAGVKFDPNDQEIL-QHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKL 121
NE P LP G +F P D+E+L +L KV KL E I ++ I P L
Sbjct: 15 NEAP-LP-GFRFHPTDEELLGYYLRRKVENKTIKL-----ELIKQID----IYKYDPWDL 63
Query: 122 PGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFK 181
P V G+ +F G + R V+ + W TG +PV + G K
Sbjct: 64 PRVSSVGEKEWYFF-----CMRGRKYRNSVRPNRVTGSGFWKATGIDKPVYSNLDCVGLK 118
Query: 182 KILVLYTNYGRQKKPEKTNWVMHQYHLGS 210
K LV Y G K KT+W+MH++ L S
Sbjct: 119 KSLVYY--LGSAGKGTKTDWMMHEFRLPS 145
>NA94_ARATH (Q9FIW5) Putative NAC-domain containing protein 94
(ANAC094)
Length = 337
Score = 44.3 bits (103), Expect = 3e-04
Identities = 44/144 (30%), Positives = 64/144 (43%), Gaps = 21/144 (14%)
Query: 71 GVKFDPNDQEILQ-HLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKDGQ 129
G +F P D+E++ +L+ KVL L + I ++ I P LP + G+
Sbjct: 23 GFRFHPTDEELVSFYLKRKVLHK-----SLPFDLIKKVD----IYKYDPWDLPKLAAMGE 73
Query: 130 IRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVI-IDGVMK-GFKKILVL 186
+F+ P + Y TR R W TG RP+ +D G KK LV
Sbjct: 74 KEWYFYCPRDRKYRNSTRPNRVT------GGGFWKATGTDRPIYSLDSTRCIGLKKSLVF 127
Query: 187 YTNYGRQKKPEKTNWVMHQYHLGS 210
Y GR K KT+W+MH++ L S
Sbjct: 128 YR--GRAAKGVKTDWMMHEFRLPS 149
>NA90_ARATH (Q9FMR3) NAC-domain containing protein 90 (ANAC090)
Length = 235
Score = 40.4 bits (93), Expect = 0.004
Identities = 38/144 (26%), Positives = 60/144 (41%), Gaps = 22/144 (15%)
Query: 71 GVKFDPNDQEILQ-----HLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVK 125
G +F P ++E++ LE + + ++ P++D F + +H + GV+
Sbjct: 8 GFRFYPTEEELVSFYLRNQLEGRSDDSMHRVIPVLDVF--------EVEPSHLPNVAGVR 59
Query: 126 KDGQIRH-FFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVII-DGVMKGFKKI 183
G FF P + R+ R + W TG PV D M G KK
Sbjct: 60 CRGDAEQWFFFVPRQE-----REARGGRPSRTTGSGYWKATGSPGPVFSKDNKMIGAKKT 114
Query: 184 LVLYTNYGRQKKPEKTNWVMHQYH 207
+V YT G+ KT W M++YH
Sbjct: 115 MVFYT--GKAPTGRKTKWKMNEYH 136
>NUOG_BUCAP (Q8K9Y2) NADH-quinone oxidoreductase chain G (EC
1.6.99.5) (NADH dehydrogenase I, chain G) (NDH-1, chain
G)
Length = 910
Score = 33.9 bits (76), Expect = 0.40
Identities = 27/90 (30%), Positives = 44/90 (48%), Gaps = 7/90 (7%)
Query: 176 VMKGFKKILVLYTNYGRQKKPEKTNW-VMHQYHLGSNEEEKDGELVVSKKI-LEY----- 228
+ K KKIL +Y ++ +K ++ W V+ YH+ NEE ++ + I LEY
Sbjct: 787 LFKSNKKILDIYFHFSPKKFIKEKYWYVIPYYHIFGNEELTQYSSIIKQNIPLEYVLISE 846
Query: 229 LSSSERRLACNSVIRFMYSSVNFFLSLMAS 258
L E L NS++ F + +F L + S
Sbjct: 847 LDGLELGLKKNSIVEFNCLNQDFRLPIRLS 876
>YI65_ARCFU (O28414) Hypothetical protein AF1865
Length = 321
Score = 32.3 bits (72), Expect = 1.2
Identities = 23/71 (32%), Positives = 35/71 (48%), Gaps = 13/71 (18%)
Query: 178 KGFKKILVLYTN----YGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSKKILEYLSSSE 233
KG ++LV++T+ GR+K K+NW + L EEK V K I++YL
Sbjct: 57 KGIPEVLVVFTSRDVIEGRKKLEYKSNW----FSLSGGSEEK-----VEKPIVKYLKKLF 107
Query: 234 RRLACNSVIRF 244
R + N + F
Sbjct: 108 RHIEKNFNLEF 118
>YFF9_SCHPO (O14066) Hypothetical serine-rich protein C1687.09 in
chromosome I
Length = 1379
Score = 32.0 bits (71), Expect = 1.5
Identities = 18/60 (30%), Positives = 28/60 (46%), Gaps = 2/60 (3%)
Query: 95 KLHPLIDEFIPTLEGENGICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTD 154
K+ P + E P L+ + I P L +K + +HF+HR Y +GT R+ D
Sbjct: 1164 KIFPAVHENPPPLKTKQSIAKPSPHTLRVPQK--KHKHFYHRDDGKYKSGTNLRKSETVD 1221
>DYI1_RAT (Q63100) Dynein intermediate chain 1, cytosolic (DH IC-1)
(Cytoplasmic dynein intermediate chain 1)
Length = 643
Score = 30.0 bits (66), Expect = 5.8
Identities = 17/65 (26%), Positives = 27/65 (41%)
Query: 28 ESPLVTDDETKPTEVRTLTCPSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEA 87
++PL T + E + P GH+ E ++Q E P + Q+IL E
Sbjct: 169 QTPLATHQSEEDEEDEEMVEPKVGHDSELENQDKKQETKEAPPRELTEEEKQQILHSEEF 228
Query: 88 KVLFD 92
+ FD
Sbjct: 229 LIFFD 233
>AT7B_MOUSE (Q64446) Copper-transporting ATPase 2 (EC 3.6.3.4)
(Copper pump 2) (Wilson disease-associated protein
homolog)
Length = 1462
Score = 30.0 bits (66), Expect = 5.8
Identities = 26/70 (37%), Positives = 37/70 (52%), Gaps = 12/70 (17%)
Query: 23 PGTSNE---SPLVTDDETKPTEVRTLTCPSCGHNIEFQDQ---GGINELPGLPAG---VK 73
PG S+E SP T + +++ +TC SC NIE Q G ++ L L +G VK
Sbjct: 474 PGHSSETPSSPGATASQKCFVQIKGMTCASCVSNIERSLQRHAGILSVLVALMSGKAEVK 533
Query: 74 FDPNDQEILQ 83
+DP EI+Q
Sbjct: 534 YDP---EIIQ 540
>ACHB_TORCA (P02712) Acetylcholine receptor protein, beta chain
precursor
Length = 493
Score = 30.0 bits (66), Expect = 5.8
Identities = 19/80 (23%), Positives = 33/80 (40%), Gaps = 1/80 (1%)
Query: 100 IDEFIPTLEGENGICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRR-KVQTDEEGS 158
I+ P L + + P+ P + +F +P+ + R VQ + S
Sbjct: 347 IETLPPFLWIQRPVTTPSPDSKPTIISRANDEYFIRKPAGDFVCPVDNARVAVQPERLFS 406
Query: 159 ETRWHKTGKTRPVIIDGVMK 178
E +WH G T+PV + +K
Sbjct: 407 EMKWHLNGLTQPVTLPQDLK 426
>P1L1_HUMAN (Q8TDX9) Polycystic kidney disease 1-like 1 protein
(Polycystin 1L1)
Length = 2849
Score = 29.6 bits (65), Expect = 7.6
Identities = 22/84 (26%), Positives = 35/84 (41%), Gaps = 13/84 (15%)
Query: 114 CYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVII 173
C EK+ ++ + + H PSKA GTR+R + + +TR +
Sbjct: 2239 CAGEVEKVLAARQQARHLRWAHPPSKAQLRGTRQRMR-------------RESRTRAALR 2285
Query: 174 DGVMKGFKKILVLYTNYGRQKKPE 197
D M +L+L YGR + E
Sbjct: 2286 DISMDILMLLLLLCVIYGRFSQDE 2309
>DYI1_MOUSE (O88485) Dynein intermediate chain 1, cytosolic (DH
IC-1) (Cytoplasmic dynein intermediate chain 1)
Length = 628
Score = 29.6 bits (65), Expect = 7.6
Identities = 17/65 (26%), Positives = 27/65 (41%)
Query: 28 ESPLVTDDETKPTEVRTLTCPSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEA 87
++PL T + E + P GH+ E ++Q E P + Q+IL E
Sbjct: 154 QTPLATHQSEEDEEDEEMVEPKIGHDSELENQEKKQETKEAPPRELTEEEKQQILHSEEF 213
Query: 88 KVLFD 92
+ FD
Sbjct: 214 LIFFD 218
>AT7B_HUMAN (P35670) Copper-transporting ATPase 2 (EC 3.6.3.4)
(Copper pump 2) (Wilson disease-associated protein)
Length = 1465
Score = 29.6 bits (65), Expect = 7.6
Identities = 19/52 (36%), Positives = 31/52 (59%), Gaps = 9/52 (17%)
Query: 41 EVRTLTCPSCGHNIE---FQDQGGINELPGLPAG---VKFDPNDQEILQHLE 86
+++ +TC SC NIE ++ G ++ L L AG +K+DP E++Q LE
Sbjct: 493 QIKGMTCASCVSNIERNLQKEAGVLSVLVALMAGKAEIKYDP---EVIQPLE 541
>SYL_UREPA (Q9PQC0) Leucyl-tRNA synthetase (EC 6.1.1.4)
(Leucine--tRNA ligase) (LeuRS)
Length = 806
Score = 29.3 bits (64), Expect = 9.9
Identities = 17/45 (37%), Positives = 25/45 (54%), Gaps = 7/45 (15%)
Query: 67 GLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGEN 111
G P V FD N+Q K++ D+P L P ++EF P+ GE+
Sbjct: 426 GEPFPVLFDENNQ-------IKIIEDLPVLLPNLNEFKPSKTGES 463
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,666,256
Number of Sequences: 164201
Number of extensions: 1558520
Number of successful extensions: 3790
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 5
Number of HSP's successfully gapped in prelim test: 19
Number of HSP's that attempted gapping in prelim test: 3773
Number of HSP's gapped (non-prelim): 28
length of query: 259
length of database: 59,974,054
effective HSP length: 108
effective length of query: 151
effective length of database: 42,240,346
effective search space: 6378292246
effective search space used: 6378292246
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 64 (29.3 bits)
Medicago: description of AC142143.13


