

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141865.9 - phase: 0 /pseudo
(547 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
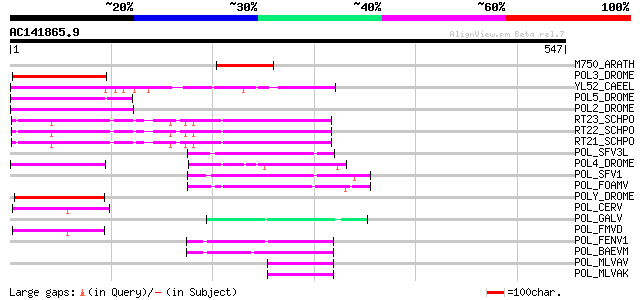
Score E
Sequences producing significant alignments: (bits) Value
M750_ARATH (P92516) Hypothetical mitochondrial protein AtMg00750... 85 4e-16
POL3_DROME (P04323) Retrovirus-related Pol polyprotein from tran... 82 4e-15
YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III 82 5e-15
POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from tran... 75 4e-13
POL2_DROME (P20825) Retrovirus-related Pol polyprotein from tran... 75 4e-13
RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protei... 69 2e-11
RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protei... 69 2e-11
RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protei... 69 2e-11
POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC 3.4.2... 68 7e-11
POL4_DROME (P10394) Retrovirus-related Pol polyprotein from tran... 67 2e-10
POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC 3.4.23... 65 3e-10
POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse transcript... 61 7e-09
POLY_DROME (P10401) Retrovirus-related Pol polyprotein from tran... 61 7e-09
POL_CERV (P05400) Enzymatic polyprotein [Contains: Aspartic prot... 59 3e-08
POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC 3.4.23... 56 2e-07
POL_FMVD (P09523) Enzymatic polyprotein [Contains: Aspartic prot... 55 4e-07
POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse transcript... 54 8e-07
POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC 3.4.2... 53 2e-06
POL_MLVAV (P03356) Pol polyprotein [Contains: Protease (EC 3.4.2... 50 1e-05
POL_MLVAK (P03357) Pol polyprotein [Contains: Reverse transcript... 50 1e-05
>M750_ARATH (P92516) Hypothetical mitochondrial protein AtMg00750
(ORF119)
Length = 119
Score = 85.1 bits (209), Expect = 4e-16
Identities = 36/56 (64%), Positives = 44/56 (78%)
Query: 205 ILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEMPLNNILEVEIFDVWVIDFI 260
+L +GF+WP+ FKD H F+S CD CQR GN TKRNEMP + ILEVE+FDVW I F+
Sbjct: 35 VLQAGFYWPTTFKDAHGFVSSCDACQRKGNFTKRNEMPQHFILEVEVFDVWGIYFM 90
>POL3_DROME (P04323) Retrovirus-related Pol polyprotein from
transposon 17.6 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1058
Score = 82.0 bits (201), Expect = 4e-15
Identities = 42/93 (45%), Positives = 58/93 (62%)
Query: 3 YASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAKPR 62
Y S+TL+ +NY+T EKELLA+V+A FR YL+G + +DH + +L KD +
Sbjct: 533 YISRTLNEHEINYSTIEKELLAIVWATKTFRHYLLGRHFEISSDHQPLSWLYRMKDPNSK 592
Query: 63 LIQLIFLLQEFDLEIKDKKGVENVVADHLSRLR 95
L + L EFD +IK KG EN VAD LSR++
Sbjct: 593 LTRWRVKLSEFDFDIKYIKGKENCVADALSRIK 625
>YL52_CAEEL (P34431) Hypothetical protein F44E2.2 in chromosome III
Length = 2186
Score = 81.6 bits (200), Expect = 5e-15
Identities = 91/355 (25%), Positives = 148/355 (41%), Gaps = 54/355 (15%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I +ASK L A Y T+ E LA+++A+ +F+ + G+ ITV+TDH + LL
Sbjct: 1271 IAFASKALSPAETRYHITDLEALAMMFALRRFKTIIYGTAITVFTDHKPLISLLKGSPLA 1330
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSR-------LRETNEDEL-----PLDDSF 108
RL + + EFD++I G N VAD LSR L E EL +
Sbjct: 1331 DRLWRWSIEILEFDVKIVYLAGKANAVADALSRGGCPPNELEEEQTKELTSIVNAIQTEL 1390
Query: 109 PD--DQLFLLAQTNA--PWYADFVNFLAAGV---------LPPELSYQQKKKFFNDLKHY 155
PD D L + + + + L G + E+S + K LK+
Sbjct: 1391 PDILDSSCWLERLKGEDEGWKEVIAALEGGKTKGTFKIVGIESEISLEYYKIVGGVLKNT 1450
Query: 156 YSDEPYLFKRG*DGIFRRCIPENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSL 215
+E R +PE + +L H GH +K ++++H F+WP +
Sbjct: 1451 EIEEQ----------SRSVVPEKIRTPLLKELHEGMLAGHFGIKK-MWRMVHRKFYWPQM 1499
Query: 216 FKDVHLFISKCDKC-------QRTGNITK-RNEMPLNNILEVEIFDVWVIDFIGPFPSSF 267
V + C KC + T ++T R PL I+ ++ DV + S
Sbjct: 1500 RVCVENCVRTCAKCLCANDHSKLTSSLTPYRMTFPL-EIVACDLMDVGL--------SVQ 1550
Query: 268 VNQYILVAVDYVSKWVEVIASLTNDAQVVIKLFKKNHFSRFG-VPRVVISDGGSQ 321
N+YIL +D +K+ + A+ V+K F + G +P +++D G +
Sbjct: 1551 GNRYILTIIDLFTKYGTAVPIPDKKAETVLKAFVERWAIGEGRIPLKLLTDQGKE 1605
>POL5_DROME (Q8I7P9) Retrovirus-related Pol polyprotein from
transposon opus [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1003
Score = 75.1 bits (183), Expect = 4e-13
Identities = 45/123 (36%), Positives = 74/123 (59%), Gaps = 3/123 (2%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGS-KITVYTDHSTIKYLLNKKDA 59
I Y S++L+ NYAT EKE+LA+++++D R YL G+ I VYTDH + + L ++
Sbjct: 461 IAYISRSLNKTEENYATIEKEMLAIIWSLDNLRAYLYGAGTIKVYTDHQPLTFALGNRNF 520
Query: 60 KPRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLD-DSFPDDQLFLLAQ 118
+L + ++E++ E+ K G NVVAD LSR+ ++L D D+ P+D + LA
Sbjct: 521 NAKLKRWKARIEEYNCELIYKPGKSNVVADALSRI-PPQLNQLSTDLDANPEDDMQSLAT 579
Query: 119 TNA 121
++
Sbjct: 580 AHS 582
>POL2_DROME (P20825) Retrovirus-related Pol polyprotein from
transposon 297 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1059
Score = 75.1 bits (183), Expect = 4e-13
Identities = 45/123 (36%), Positives = 68/123 (54%), Gaps = 1/123 (0%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I + S+TL+ +NY+ EKELLA+V+A FR YL+G + + +DH +++L N K+
Sbjct: 530 ISFISRTLNDHELNYSAIEKELLAIVWATKTFRHYLLGRQFLIASDHQPLRWLHNLKEPG 589
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLR-ETNEDELPLDDSFPDDQLFLLAQT 119
+L + L E+ +I KG EN VAD LSR++ E N S +D L+ T
Sbjct: 590 AKLERWRVRLSEYQFKIDYIKGKENSVADALSRIKIEENHHSEATQHSAEEDNSNLIHLT 649
Query: 120 NAP 122
P
Sbjct: 650 EKP 652
>RT23_SCHPO (Q9UR07) Retrotransposable element Tf2 155 kDa protein
type 3
Length = 1333
Score = 69.3 bits (168), Expect = 2e-11
Identities = 81/331 (24%), Positives = 145/331 (43%), Gaps = 32/331 (9%)
Query: 2 YYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGS--KITVYTDH-STIKYLLNKKD 58
YY++K + A +NY+ ++KE+LA++ ++ +R YL + + TDH + I + N+ +
Sbjct: 737 YYSAK-MSKAQLNYSVSDKEMLAIIKSLKHWRHYLESTIEPFKILTDHRNLIGRITNESE 795
Query: 59 AK-PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLA 117
+ RL + LQ+F+ EI + G N +AD LSR+ + E P+ D+ + +
Sbjct: 796 PENKRLARWQLFLQDFNFEINYRPGSANHIADALSRIVDETE---PIPKDSEDNSINFVN 852
Query: 118 QTNAPWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYS--DEPYLFKRG*DGIF---- 171
Q + DF N + Y K N L + +E K DG+
Sbjct: 853 QISIT--DDFKNQVVT-------EYTNDTKLLNLLNNEDKRVEENIQLK---DGLLINSK 900
Query: 172 -RRCIPENE--VSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDK 228
+ +P + +I+ H H + + IL F W + K + ++ C
Sbjct: 901 DQILLPNDTQLTRTIIKKYHEEGKLIHPGIELLTNIILRR-FTWKGIRKQIQEYVQNCHT 959
Query: 229 CQRTGNITKRNEMPLNNILEVE-IFDVWVIDFIGPFPSSFVNQYILVAVDYVSKW-VEVI 286
CQ + + PL I E ++ +DFI P S + V VD SK + V
Sbjct: 960 CQINKSRNHKPYGPLQPIPPSERPWESLSMDFITALPESSGYNALFVVVDRFSKMAILVP 1019
Query: 287 ASLTNDAQVVIKLFKKNHFSRFGVPRVVISD 317
+ + A+ ++F + + FG P+ +I+D
Sbjct: 1020 CTKSITAEQTARMFDQRVIAYFGNPKEIIAD 1050
>RT22_SCHPO (Q9C0R2) Retrotransposable element Tf2 155 kDa protein
type 2
Length = 1333
Score = 69.3 bits (168), Expect = 2e-11
Identities = 81/331 (24%), Positives = 145/331 (43%), Gaps = 32/331 (9%)
Query: 2 YYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGS--KITVYTDH-STIKYLLNKKD 58
YY++K + A +NY+ ++KE+LA++ ++ +R YL + + TDH + I + N+ +
Sbjct: 737 YYSAK-MSKAQLNYSVSDKEMLAIIKSLKHWRHYLESTIEPFKILTDHRNLIGRITNESE 795
Query: 59 AK-PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLA 117
+ RL + LQ+F+ EI + G N +AD LSR+ + E P+ D+ + +
Sbjct: 796 PENKRLARWQLFLQDFNFEINYRPGSANHIADALSRIVDETE---PIPKDSEDNSINFVN 852
Query: 118 QTNAPWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYS--DEPYLFKRG*DGIF---- 171
Q + DF N + Y K N L + +E K DG+
Sbjct: 853 QISIT--DDFKNQVVT-------EYTNDTKLLNLLNNEDKRVEENIQLK---DGLLINSK 900
Query: 172 -RRCIPENE--VSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDK 228
+ +P + +I+ H H + + IL F W + K + ++ C
Sbjct: 901 DQILLPNDTQLTRTIIKKYHEEGKLIHPGIELLTNIILRR-FTWKGIRKQIQEYVQNCHT 959
Query: 229 CQRTGNITKRNEMPLNNILEVE-IFDVWVIDFIGPFPSSFVNQYILVAVDYVSKW-VEVI 286
CQ + + PL I E ++ +DFI P S + V VD SK + V
Sbjct: 960 CQINKSRNHKPYGPLQPIPPSERPWESLSMDFITALPESSGYNALFVVVDRFSKMAILVP 1019
Query: 287 ASLTNDAQVVIKLFKKNHFSRFGVPRVVISD 317
+ + A+ ++F + + FG P+ +I+D
Sbjct: 1020 CTKSITAEQTARMFDQRVIAYFGNPKEIIAD 1050
>RT21_SCHPO (Q05654) Retrotransposable element Tf2 155 kDa protein
type 1
Length = 1333
Score = 69.3 bits (168), Expect = 2e-11
Identities = 81/331 (24%), Positives = 145/331 (43%), Gaps = 32/331 (9%)
Query: 2 YYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGS--KITVYTDH-STIKYLLNKKD 58
YY++K + A +NY+ ++KE+LA++ ++ +R YL + + TDH + I + N+ +
Sbjct: 737 YYSAK-MSKAQLNYSVSDKEMLAIIKSLKHWRHYLESTIEPFKILTDHRNLIGRITNESE 795
Query: 59 AK-PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLA 117
+ RL + LQ+F+ EI + G N +AD LSR+ + E P+ D+ + +
Sbjct: 796 PENKRLARWQLFLQDFNFEINYRPGSANHIADALSRIVDETE---PIPKDSEDNSINFVN 852
Query: 118 QTNAPWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYS--DEPYLFKRG*DGIF---- 171
Q + DF N + Y K N L + +E K DG+
Sbjct: 853 QISIT--DDFKNQVVT-------EYTNDTKLLNLLNNEDKRVEENIQLK---DGLLINSK 900
Query: 172 -RRCIPENE--VSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDK 228
+ +P + +I+ H H + + IL F W + K + ++ C
Sbjct: 901 DQILLPNDTQLTRTIIKKYHEEGKLIHPGIELLTNIILRR-FTWKGIRKQIQEYVQNCHT 959
Query: 229 CQRTGNITKRNEMPLNNILEVE-IFDVWVIDFIGPFPSSFVNQYILVAVDYVSKW-VEVI 286
CQ + + PL I E ++ +DFI P S + V VD SK + V
Sbjct: 960 CQINKSRNHKPYGPLQPIPPSERPWESLSMDFITALPESSGYNALFVVVDRFSKMAILVP 1019
Query: 287 ASLTNDAQVVIKLFKKNHFSRFGVPRVVISD 317
+ + A+ ++F + + FG P+ +I+D
Sbjct: 1020 CTKSITAEQTARMFDQRVIAYFGNPKEIIAD 1050
>POL_SFV3L (P27401) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1157
Score = 67.8 bits (164), Expect = 7e-11
Identities = 44/145 (30%), Positives = 71/145 (48%), Gaps = 6/145 (4%)
Query: 176 PENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNI 235
P+++ I+ H+ ++ G ST F + S +WWP+L KDV I +C +C T
Sbjct: 814 PKSDRPQIILQAHNIAHTGRDST----FLKVSSKYWWPNLRKDVVKVIRQCKQCLVTNAA 869
Query: 236 TKRNEMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQV 295
T L V+ FD + ID+IGP P S ++LV VD ++ +V + +
Sbjct: 870 TLAAPPILRPERPVKPFDKFFIDYIGPLPPSNGYLHVLVVVDSMTGFVWLYPTKAPSTSA 929
Query: 296 VIKLFKKNHFSRFGVPRVVISDGGS 320
+K N + VP+V+ SD G+
Sbjct: 930 TVKAL--NMLTSIAVPKVIHSDQGA 952
>POL4_DROME (P10394) Retrovirus-related Pol polyprotein from
transposon 412 [Contains: Protease (EC 3.4.23.-);
Reverse transcriptase (EC 2.7.7.49); Endonuclease]
Length = 1237
Score = 66.6 bits (161), Expect = 2e-10
Identities = 35/94 (37%), Positives = 55/94 (58%)
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
+ YAS+ N +TTE+EL A+ +AI FR Y+ G TV TDH + YL + +
Sbjct: 642 VAYASRAFTKGESNKSTTEQELAAIHWAIIHFRPYIYGKHFTVKTDHRPLTYLFSMVNPS 701
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRL 94
+L ++ L+E++ ++ KG +N VAD LSR+
Sbjct: 702 SKLTRIRLELEEYNFTVEYLKGKDNHVADALSRI 735
Score = 64.3 bits (155), Expect = 8e-10
Identities = 46/163 (28%), Positives = 80/163 (48%), Gaps = 10/163 (6%)
Query: 177 ENEVSSILTHCHSSSY-GGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNI 235
E E +IL+ H GGH K K+ ++W ++ K + ++ KC KCQ+
Sbjct: 890 EKEKEAILSTLHDDPIQGGHTGITKTLAKVKRH-YYWKNMSKYIKEYVRKCQKCQK-AKT 947
Query: 236 TKRNEMPLNNILEV--EIFDVWVIDFIGPFP-SSFVNQYILVAVDYVSKWVEVIASLTND 292
TK + P+ I E FD V+D IGP P S N+Y + + ++K++ I
Sbjct: 948 TKHTKTPM-TITETPEHAFDRVVVDTIGPLPKSENGNEYAVTLICDLTKYLVAIPIANKS 1006
Query: 293 AQVVIKLFKKNHFSRFGVPRVVISDGGSQ---NILKNFCKNLE 332
A+ V K ++ ++G + I+D G++ +I+ + CK L+
Sbjct: 1007 AKTVAKAIFESFILKYGPMKTFITDMGTEYKNSIITDLCKYLK 1049
>POL_SFV1 (P23074) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1161
Score = 65.5 bits (158), Expect = 3e-10
Identities = 49/186 (26%), Positives = 88/186 (46%), Gaps = 12/186 (6%)
Query: 176 PENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNI 235
P+ + I++ H+ ++ G + A+F + S +WWP+L KDV I +C +C T
Sbjct: 812 PKADREKIISTAHNIAHTG----RDATFLKVSSKYWWPNLRKDVVKSIRQCKQCLVTNAT 867
Query: 236 TKRNEMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQV 295
+ L + ++ FD + ID+IGP P S ++LV VD ++ +V + +
Sbjct: 868 NLTSPPILRPVKPLKPFDKFYIDYIGPLPPSNGYLHVLVVVDSMTGFVWLYPTKAPSTSA 927
Query: 296 VIKLFKKNHFSRFGVPRVVISDGGSQNILKNFCK-NLE*GTKLQ-----HPITHKLVDKW 349
+K N + +P+V+ SD G+ F E G +L+ HP + V++
Sbjct: 928 TVKAL--NMLTSIAIPKVLHSDQGAAFTSSTFADWAKEKGIQLEFSTPYHPQSSGKVERK 985
Query: 350 RSQIDK 355
S I +
Sbjct: 986 NSDIKR 991
>POL_FOAMV (P14350) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)]
Length = 886
Score = 61.2 bits (147), Expect = 7e-09
Identities = 48/187 (25%), Positives = 85/187 (44%), Gaps = 14/187 (7%)
Query: 176 PENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNI 235
P+++ I+ H+ ++ G +T KI + +WWP++ KDV + +C +C T
Sbjct: 604 PQSDRQKIVLQAHNLAHTGREATL---LKIANL-YWWPNMRKDVVKQLGRCQQCLITNAS 659
Query: 236 TKRNEMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQV 295
K + L + FD + ID+IGP P S Y+LV VD ++ + + +
Sbjct: 660 NKASGPILRPDRPQKPFDKFFIDYIGPLPPSQGYLYVLVVVDGMTGFTWLYPTKAPSTSA 719
Query: 296 VIKLFKKNHFSRFGVPRVVISDGGSQNILKNFCK-------NLE*GTKLQHPITHKLVDK 348
+K N + +P+V+ SD G+ F + +LE T HP + V++
Sbjct: 720 TVK--SLNVLTSIAIPKVIHSDQGAAFTSSTFAEWAKERGIHLEFSTP-YHPQSGSKVER 776
Query: 349 WRSQIDK 355
S I +
Sbjct: 777 KNSDIKR 783
>POLY_DROME (P10401) Retrovirus-related Pol polyprotein from
transposon gypsy [Contains: Reverse transcriptase (EC
2.7.7.49); Endonuclease]
Length = 1035
Score = 61.2 bits (147), Expect = 7e-09
Identities = 32/90 (35%), Positives = 55/90 (60%), Gaps = 1/90 (1%)
Query: 5 SKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSK-ITVYTDHSTIKYLLNKKDAKPRL 63
S+TL NYAT E+ELLA+V+A+ K + +L GS+ I ++TDH + + + ++ ++
Sbjct: 521 SRTLKQPEQNYATNERELLAIVWALGKLQNFLYGSREINIFTDHQPLTFAVADRNTNAKI 580
Query: 64 IQLIFLLQEFDLEIKDKKGVENVVADHLSR 93
+ + + + ++ K G EN VAD LSR
Sbjct: 581 KRWKSYIDQHNAKVFYKPGKENFVADALSR 610
>POL_CERV (P05400) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 659
Score = 58.9 bits (141), Expect = 3e-08
Identities = 37/100 (37%), Positives = 53/100 (53%), Gaps = 4/100 (4%)
Query: 3 YASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLN---KKDA 59
YAS + A NY + EKELLAV+ I KF YL S+ + TD+ + +N K D
Sbjct: 558 YASGSFKAAERNYHSNEKELLAVIRVIKKFSIYLTPSRFLIRTDNKNFTHFVNINLKGDR 617
Query: 60 KP-RLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETN 98
K RL++ L ++D +++ G +NV AD L TN
Sbjct: 618 KQGRLVRWQMWLSQYDFDVEHIAGTKNVFADFLQENTLTN 657
>POL_GALV (P21414) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1165
Score = 56.2 bits (134), Expect = 2e-07
Identities = 44/158 (27%), Positives = 64/158 (39%), Gaps = 6/158 (3%)
Query: 195 HASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEMPLNNILEVEIFDV 254
H +K + + P+L V S+C C T +T E +
Sbjct: 821 HLGPEKLLQLVNRTSLLIPNLQSAVREVTSQCQACAMTNAVTTYRETGKRQRGDRPGV-Y 879
Query: 255 WVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLFKKNHFSRFGVPRVV 314
W +DF P + N+Y+LV +D S WVE + T A +V K + RFG+P+V+
Sbjct: 880 WEVDFTEIKPGRYGNKYLLVFIDTFSGWVEAFPTKTETALIVCKKILEEILPRFGIPKVL 939
Query: 315 ISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWRSQ 352
SD G F + G Q I KL +R Q
Sbjct: 940 GSDNGPA-----FVAQVSQGLATQLGINWKLHCAYRPQ 972
>POL_FMVD (P09523) Enzymatic polyprotein [Contains: Aspartic
protease (EC 3.4.23.-); Endonuclease; Reverse
transcriptase (EC 2.7.7.49)]
Length = 666
Score = 55.5 bits (132), Expect = 4e-07
Identities = 35/95 (36%), Positives = 53/95 (54%), Gaps = 4/95 (4%)
Query: 3 YASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLN---KKDA 59
Y+S + A NY + +KELLAV I KF YL + TV TD+ Y L K D+
Sbjct: 568 YSSGSFKQAEKNYHSNDKELLAVKQVITKFSAYLTPVRFTVRTDNKNFTYFLRINLKGDS 627
Query: 60 KP-RLIQLIFLLQEFDLEIKDKKGVENVVADHLSR 93
K RL++ ++ +++ +GV+NV+AD L+R
Sbjct: 628 KQGRLVRWQNWFSKYQFDVEHLEGVKNVLADCLTR 662
>POL_FENV1 (P31792) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 1046
Score = 54.3 bits (129), Expect = 8e-07
Identities = 42/146 (28%), Positives = 62/146 (41%), Gaps = 6/146 (4%)
Query: 175 IPENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTG- 233
+P E +++ H+ + H S QK I + F P + S C CQ+
Sbjct: 689 LPRKEALAMIQQMHAWT---HLSNQKLKLLIEKTDFLIPKAGTLIEQVTSACKVCQQVNA 745
Query: 234 NITKRNEMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDA 293
T+ E ++ W IDF P +Y+LV VD S WVE + A
Sbjct: 746 GATRVPEGKRTRGNRPGVY--WEIDFTEVKPHYAGYKYLLVFVDTFSGWVEAYPTRQETA 803
Query: 294 QVVIKLFKKNHFSRFGVPRVVISDGG 319
+V K + F RFG+P+V+ SD G
Sbjct: 804 HMVAKKILEEIFPRFGLPKVIGSDNG 829
>POL_BAEVM (P10272) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1189
Score = 52.8 bits (125), Expect = 2e-06
Identities = 41/146 (28%), Positives = 61/146 (41%), Gaps = 6/146 (4%)
Query: 175 IPENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGN 234
+P+ E +++ H+ + H +K I + F P + S C CQ+
Sbjct: 832 LPQKEALAMIQQMHAWT---HLGNRKLKLLIEKTDFLIPRASTLIEQVTSACKVCQQVNA 888
Query: 235 ITKRNEMPLNNILEVEIFDV-WVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDA 293
R +P V W IDF P +Y+LV VD S WVE + A
Sbjct: 889 GATR--VPAGKRTRGNRPGVYWEIDFTEVKPHYAGYKYLLVFVDTFSGWVEAFPTRQETA 946
Query: 294 QVVIKLFKKNHFSRFGVPRVVISDGG 319
+V K + F RFG+P+V+ SD G
Sbjct: 947 HIVAKKILEEIFPRFGLPKVIGSDNG 972
>POL_MLVAV (P03356) Pol polyprotein [Contains: Protease (EC
3.4.23.-); Reverse transcriptase/ribonuclease H (EC
2.7.7.49) (EC 3.1.26.4) (RT); Integrase (IN)]
Length = 1196
Score = 50.4 bits (119), Expect = 1e-05
Identities = 27/65 (41%), Positives = 36/65 (54%)
Query: 255 WVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLFKKNHFSRFGVPRVV 314
W IDF P + +Y+LV VD S WVE + A+VV K + F RFG+P+V+
Sbjct: 913 WEIDFTEVKPGLYGYKYLLVFVDTFSGWVEAFPTKRETARVVSKKLLEEIFPRFGMPQVL 972
Query: 315 ISDGG 319
SD G
Sbjct: 973 GSDNG 977
>POL_MLVAK (P03357) Pol polyprotein [Contains: Reverse
transcriptase/ribonuclease H (EC 2.7.7.49) (EC 3.1.26.4)
(RT); Integrase (IN)] (Fragment)
Length = 843
Score = 50.4 bits (119), Expect = 1e-05
Identities = 27/65 (41%), Positives = 36/65 (54%)
Query: 255 WVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLFKKNHFSRFGVPRVV 314
W IDF P + +Y+LV VD S WVE + A+VV K + F RFG+P+V+
Sbjct: 561 WEIDFTEVKPGLYGYKYLLVFVDTFSGWVEAFPTKRETARVVSKKLLEEIFPRFGMPQVL 620
Query: 315 ISDGG 319
SD G
Sbjct: 621 GSDNG 625
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.336 0.148 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 57,725,421
Number of Sequences: 164201
Number of extensions: 2345537
Number of successful extensions: 8218
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 37
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 8142
Number of HSP's gapped (non-prelim): 62
length of query: 547
length of database: 59,974,054
effective HSP length: 115
effective length of query: 432
effective length of database: 41,090,939
effective search space: 17751285648
effective search space used: 17751285648
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC141865.9


