

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141863.5 - phase: 0
(139 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
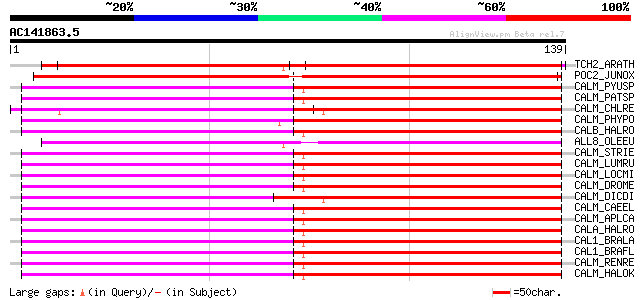
Score E
Sequences producing significant alignments: (bits) Value
TCH2_ARATH (P25070) Calmodulin-related protein 2, touch-induced 105 4e-23
POC2_JUNOX (O64943) Polcalcin Jun o 2 (Calcium-binding pollen al... 102 3e-22
CALM_PYUSP (P11121) Calmodulin (CaM) 93 2e-19
CALM_PATSP (P02595) Calmodulin (CaM) 93 2e-19
CALM_CHLRE (P04352) Calmodulin (CaM) 93 2e-19
CALM_PHYPO (O96102) Calmodulin (CaM) 92 3e-19
CALB_HALRO (O96081) Calmodulin B (CaM B) 92 3e-19
ALL8_OLEEU (Q9M7R0) Calcium-binding allergen Ole e 8 (PCA18/PCA23) 92 3e-19
CALM_STRIE (Q8STF0) Calmodulin (CaM) 91 6e-19
CALM_LUMRU (Q9GRJ1) Calmodulin (CaM) 91 6e-19
CALM_LOCMI (P62154) Calmodulin (CaM) 91 6e-19
CALM_DROME (P62152) Calmodulin (CaM) 91 6e-19
CALM_DICDI (P02599) Calmodulin (CaM) 91 6e-19
CALM_CAEEL (O16305) Calmodulin (CaM) 91 6e-19
CALM_APLCA (P62145) Calmodulin (CaM) 91 6e-19
CALA_HALRO (P62153) Calmodulin A (CaM A) 91 6e-19
CAL1_BRALA (P62148) Calmodulin 1 (CaM 1) 91 6e-19
CAL1_BRAFL (P62147) Calmodulin 1 (CaM 1) 91 6e-19
CALM_RENRE (P62184) Calmodulin (CaM) 91 8e-19
CALM_HALOK (Q95NI4) Calmodulin (CaM) 91 1e-18
>TCH2_ARATH (P25070) Calmodulin-related protein 2, touch-induced
Length = 161
Score = 105 bits (261), Expect = 4e-23
Identities = 52/135 (38%), Positives = 82/135 (60%), Gaps = 5/135 (3%)
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALME- 67
V + FD++GDGK+S EL++ +R + +E ++ D DG+G++ L+E +AL +
Sbjct: 21 VFQRFDKNGDGKISVDELKEVIRALSPTASPEETVTMMKQFDLDGNGFIDLDEFVALFQI 80
Query: 68 ----EGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDL 123
G + DL+EAFE+YD + G I+ K L ++K +GE S+ +CK MI D+
Sbjct: 81 GIGGGGNNRNDVSDLKEAFELYDLDGNGRISAKELHSVMKNLGEKCSVQDCKKMISKVDI 140
Query: 124 DGDGLLSFDEFITMM 138
DGDG ++FDEF MM
Sbjct: 141 DGDGCVNFDEFKKMM 155
Score = 51.6 bits (122), Expect = 5e-07
Identities = 21/65 (32%), Positives = 39/65 (59%)
Query: 75 LKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDEF 134
+ D+++ F+ +D G I+ LK +++ + + S +E M+K FDLDG+G + DEF
Sbjct: 15 MDDIKKVFQRFDKNGDGKISVDELKEVIRALSPTASPEETVTMMKQFDLDGNGFIDLDEF 74
Query: 135 ITMMQ 139
+ + Q
Sbjct: 75 VALFQ 79
Score = 45.8 bits (107), Expect = 3e-05
Identities = 19/58 (32%), Positives = 36/58 (61%)
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGG 70
+D DG+G++S EL ++ +GE+ +++ + I +D DGDG ++ +E +M GG
Sbjct: 102 YDLDGNGRISAKELHSVMKNLGEKCSVQDCKKMISKVDIDGDGCVNFDEFKKMMSNGG 159
Score = 28.1 bits (61), Expect = 6.1
Identities = 11/24 (45%), Positives = 17/24 (70%)
Query: 110 SIDECKAMIKHFDLDGDGLLSFDE 133
S+D+ K + + FD +GDG +S DE
Sbjct: 14 SMDDIKKVFQRFDKNGDGKISVDE 37
>POC2_JUNOX (O64943) Polcalcin Jun o 2 (Calcium-binding pollen
allergen Jun o 2)
Length = 165
Score = 102 bits (254), Expect = 3e-22
Identities = 54/132 (40%), Positives = 84/132 (62%), Gaps = 3/132 (2%)
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
E V + FD +GDGK+S +EL LR +G ++ E + +E D+DGDGY+SL+E + L
Sbjct: 28 EEVFKKFDANGDGKISGSELADILRSLGSDVGEAEVKAMMEEADADGDGYVSLQEFVDLN 87
Query: 67 EEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGD 126
+G +KDL+ AF+++D + G I+ L L+ +GE +I+E K +I + D +GD
Sbjct: 88 NKGA---SVKDLKNAFKVFDRDCNGSISAAELCHTLESVGEPCTIEESKNIIHNVDKNGD 144
Query: 127 GLLSFDEFITMM 138
GL+S +EF TMM
Sbjct: 145 GLISVEEFQTMM 156
Score = 48.1 bits (113), Expect = 6e-06
Identities = 23/66 (34%), Positives = 37/66 (55%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
EQ + +L E F+ +D+ G I+ L +L+ +G E KAM++ D DGDG +S
Sbjct: 21 EQSVHELEEVFKKFDANGDGKISGSELADILRSLGSDVGEAEVKAMMEEADADGDGYVSL 80
Query: 132 DEFITM 137
EF+ +
Sbjct: 81 QEFVDL 86
>CALM_PYUSP (P11121) Calmodulin (CaM)
Length = 148
Score = 93.2 bits (230), Expect = 2e-19
Identities = 45/136 (33%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DGDG + E +
Sbjct: 10 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGDGTIDFPEFL 69
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M ++ +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 70 TMMARKMKDTDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+TMM
Sbjct: 130 IDGDGQVNYEEFVTMM 145
Score = 58.2 bits (139), Expect = 5e-09
Identities = 25/67 (37%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DGDG + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGDGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
>CALM_PATSP (P02595) Calmodulin (CaM)
Length = 148
Score = 93.2 bits (230), Expect = 2e-19
Identities = 45/136 (33%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DGDG + E +
Sbjct: 10 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGDGTIDFPEFL 69
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M ++ +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 70 TMMARKMKDTDSEEEIREAFRVFDKDGDGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+TMM
Sbjct: 130 IDGDGQVNYEEFVTMM 145
Score = 58.2 bits (139), Expect = 5e-09
Identities = 25/67 (37%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DGDG + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGDGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
>CALM_CHLRE (P04352) Calmodulin (CaM)
Length = 162
Score = 93.2 bits (230), Expect = 2e-19
Identities = 48/136 (35%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 13 AEFKEAFALFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMISEVDADGNGTIDFPEFL 72
Query: 64 ALMEEGGEEQKLKD-LREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
LM +E +D LREAF+++D + GFI+ L+ ++ +GE S +E MI+ D
Sbjct: 73 MLMARKMKETDHEDELREAFKVFDKDGNGFISAAELRHVMTNLGEKLSEEEVDEMIREAD 132
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+ MM
Sbjct: 133 VDGDGQVNYEEFVRMM 148
Score = 53.9 bits (128), Expect = 1e-07
Identities = 30/80 (37%), Positives = 44/80 (54%), Gaps = 4/80 (5%)
Query: 1 MKNAGFEHVLR----YFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGY 56
MK E LR FD+DG+G +S AELR + +GE++ +E + I D DGDG
Sbjct: 79 MKETDHEDELREAFKVFDKDGNGFISAAELRHVMTNLGEKLSEEEVDEMIREADVDGDGQ 138
Query: 57 LSLEELIALMEEGGEEQKLK 76
++ EE + +M G + K K
Sbjct: 139 VNYEEFVRMMTSGATDDKDK 158
Score = 52.4 bits (124), Expect = 3e-07
Identities = 22/67 (32%), Positives = 42/67 (61%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 9 EEQIAEFKEAFALFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMISEVDADGNGTIDF 68
Query: 132 DEFITMM 138
EF+ +M
Sbjct: 69 PEFLMLM 75
>CALM_PHYPO (O96102) Calmodulin (CaM)
Length = 148
Score = 92.4 bits (228), Expect = 3e-19
Identities = 45/136 (33%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 10 AEFKEAFSLFDKDGDGNITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 69
Query: 64 ALM-EEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M + + +++REAF+++D + GFI+ L+ ++ +GE S +E MI+ D
Sbjct: 70 TMMARKMADTDTEEEIREAFKVFDKDGNGFISAAELRHVMTNLGEKLSDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG +++DEF+ MM
Sbjct: 130 VDGDGQVNYDEFVKMM 145
Score = 55.8 bits (133), Expect = 3e-08
Identities = 24/67 (35%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGNITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
>CALB_HALRO (O96081) Calmodulin B (CaM B)
Length = 148
Score = 92.4 bits (228), Expect = 3e-19
Identities = 45/136 (33%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 10 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 69
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M +E +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 70 TMMARKMKETDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+TMM
Sbjct: 130 IDGDGQVNYEEFVTMM 145
Score = 56.2 bits (134), Expect = 2e-08
Identities = 24/67 (35%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
>ALL8_OLEEU (Q9M7R0) Calcium-binding allergen Ole e 8 (PCA18/PCA23)
Length = 171
Score = 92.0 bits (227), Expect = 3e-19
Identities = 51/138 (36%), Positives = 79/138 (56%), Gaps = 12/138 (8%)
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALME- 67
V FD +GDGK+S EL L+ +G +E +E +D+D DG+++++E A ++
Sbjct: 24 VFNRFDANGDGKISGDELAGVLKALGSNTSKEEIGRIMEEIDTDKDGFINVQEFAAFVKA 83
Query: 68 -------EGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKH 120
GGE + L+EAFE+YD + G I+ L ++L ++GE + +C MIK
Sbjct: 84 ETDPYPSSGGENE----LKEAFELYDQDHNGLISSVELHKILTRLGERYAEHDCVEMIKS 139
Query: 121 FDLDGDGLLSFDEFITMM 138
D DGDG +SF+EF MM
Sbjct: 140 VDSDGDGYVSFEEFKKMM 157
Score = 35.4 bits (80), Expect = 0.038
Identities = 17/67 (25%), Positives = 35/67 (51%)
Query: 73 QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFD 132
Q+ +++ F +D+ G I+ L +LK +G + S +E +++ D D DG ++
Sbjct: 16 QEPNEVQGVFNRFDANGDGKISGDELAGVLKALGSNTSKEEIGRIMEEIDTDKDGFINVQ 75
Query: 133 EFITMMQ 139
EF ++
Sbjct: 76 EFAAFVK 82
Score = 35.0 bits (79), Expect = 0.050
Identities = 28/109 (25%), Positives = 47/109 (42%), Gaps = 10/109 (9%)
Query: 9 VLRYFDEDGDGKVSPAELRQRLRI-------MGEEILLKEAEMAIEAMDSDGDGYLSLEE 61
++ D D DG ++ E ++ G E LKEA E D D +G +S E
Sbjct: 60 IMEEIDTDKDGFINVQEFAAFVKAETDPYPSSGGENELKEA---FELYDQDHNGLISSVE 116
Query: 62 LIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKS 110
L ++ GE D E + DS+ G+++ + K+M+ + S
Sbjct: 117 LHKILTRLGERYAEHDCVEMIKSVDSDGDGYVSFEEFKKMMTNKSGNNS 165
>CALM_STRIE (Q8STF0) Calmodulin (CaM)
Length = 155
Score = 91.3 bits (225), Expect = 6e-19
Identities = 44/136 (32%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 17 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 76
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M ++ +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 77 TMMARKMKDTDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 136
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+TMM
Sbjct: 137 IDGDGQVNYEEFVTMM 152
Score = 56.2 bits (134), Expect = 2e-08
Identities = 24/67 (35%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 13 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 72
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 73 PEFLTMM 79
>CALM_LUMRU (Q9GRJ1) Calmodulin (CaM)
Length = 148
Score = 91.3 bits (225), Expect = 6e-19
Identities = 44/136 (32%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 10 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 69
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M ++ +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 70 TMMARKMKDTDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+TMM
Sbjct: 130 IDGDGQVNYEEFVTMM 145
Score = 56.2 bits (134), Expect = 2e-08
Identities = 24/67 (35%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
>CALM_LOCMI (P62154) Calmodulin (CaM)
Length = 148
Score = 91.3 bits (225), Expect = 6e-19
Identities = 44/136 (32%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 10 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 69
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M ++ +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 70 TMMARKMKDTDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+TMM
Sbjct: 130 IDGDGQVNYEEFVTMM 145
Score = 56.2 bits (134), Expect = 2e-08
Identities = 24/67 (35%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
>CALM_DROME (P62152) Calmodulin (CaM)
Length = 148
Score = 91.3 bits (225), Expect = 6e-19
Identities = 44/136 (32%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 10 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 69
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M ++ +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 70 TMMARKMKDTDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+TMM
Sbjct: 130 IDGDGQVNYEEFVTMM 145
Score = 56.2 bits (134), Expect = 2e-08
Identities = 24/67 (35%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
>CALM_DICDI (P02599) Calmodulin (CaM)
Length = 151
Score = 91.3 bits (225), Expect = 6e-19
Identities = 44/136 (32%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 12 AEFKEAFSLFDKDGDGSITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGNIDFPEFL 71
Query: 64 ALMEEGGEEQKLKD-LREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M ++ ++ +REAF+++D + G+I+ L+ ++ +GE + +E MI+ D
Sbjct: 72 TMMARKMQDTDTEEEIREAFKVFDKDGNGYISAAELRHVMTSLGEKLTNEEVDEMIREAD 131
Query: 123 LDGDGLLSFDEFITMM 138
LDGDG +++DEF+ MM
Sbjct: 132 LDGDGQVNYDEFVKMM 147
Score = 56.2 bits (134), Expect = 2e-08
Identities = 25/72 (34%), Positives = 45/72 (61%)
Query: 67 EEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGD 126
+E E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+
Sbjct: 3 QESLTEEQIAEFKEAFSLFDKDGDGSITTKELGTVMRSLGQNPTEAELQDMINEVDADGN 62
Query: 127 GLLSFDEFITMM 138
G + F EF+TMM
Sbjct: 63 GNIDFPEFLTMM 74
>CALM_CAEEL (O16305) Calmodulin (CaM)
Length = 148
Score = 91.3 bits (225), Expect = 6e-19
Identities = 44/136 (32%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 10 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 69
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M ++ +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 70 TMMARKMKDTDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+TMM
Sbjct: 130 IDGDGQVNYEEFVTMM 145
Score = 56.2 bits (134), Expect = 2e-08
Identities = 24/67 (35%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
>CALM_APLCA (P62145) Calmodulin (CaM)
Length = 148
Score = 91.3 bits (225), Expect = 6e-19
Identities = 44/136 (32%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 10 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 69
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M ++ +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 70 TMMARKMKDTDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+TMM
Sbjct: 130 IDGDGQVNYEEFVTMM 145
Score = 56.2 bits (134), Expect = 2e-08
Identities = 24/67 (35%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
>CALA_HALRO (P62153) Calmodulin A (CaM A)
Length = 148
Score = 91.3 bits (225), Expect = 6e-19
Identities = 44/136 (32%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 10 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 69
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M ++ +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 70 TMMARKMKDTDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+TMM
Sbjct: 130 IDGDGQVNYEEFVTMM 145
Score = 56.2 bits (134), Expect = 2e-08
Identities = 24/67 (35%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
>CAL1_BRALA (P62148) Calmodulin 1 (CaM 1)
Length = 148
Score = 91.3 bits (225), Expect = 6e-19
Identities = 44/136 (32%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 10 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 69
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M ++ +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 70 TMMARKMKDTDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+TMM
Sbjct: 130 IDGDGQVNYEEFVTMM 145
Score = 56.2 bits (134), Expect = 2e-08
Identities = 24/67 (35%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
>CAL1_BRAFL (P62147) Calmodulin 1 (CaM 1)
Length = 148
Score = 91.3 bits (225), Expect = 6e-19
Identities = 44/136 (32%), Positives = 80/136 (58%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 10 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 69
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M ++ +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 70 TMMARKMKDTDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+TMM
Sbjct: 130 IDGDGQVNYEEFVTMM 145
Score = 56.2 bits (134), Expect = 2e-08
Identities = 24/67 (35%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
>CALM_RENRE (P62184) Calmodulin (CaM)
Length = 148
Score = 90.9 bits (224), Expect = 8e-19
Identities = 44/136 (32%), Positives = 79/136 (57%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DGDG + E +
Sbjct: 10 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGDGTIDFPEFL 69
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M ++ +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 70 TMMARKMKDTDSEEEIREAFRVFDKDGDGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+ MM
Sbjct: 130 IDGDGQVNYEEFVKMM 145
Score = 58.2 bits (139), Expect = 5e-09
Identities = 25/67 (37%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DGDG + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGDGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
>CALM_HALOK (Q95NI4) Calmodulin (CaM)
Length = 148
Score = 90.5 bits (223), Expect = 1e-18
Identities = 44/136 (32%), Positives = 79/136 (57%), Gaps = 1/136 (0%)
Query: 4 AGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELI 63
A F+ FD+DGDG ++ EL +R +G+ E + I +D+DG+G + E +
Sbjct: 10 AEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDFPEFL 69
Query: 64 ALMEEGGEE-QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFD 122
+M +E +++REAF ++D + GFI+ L+ ++ +GE + +E MI+ D
Sbjct: 70 TMMARKMKETDSEEEIREAFRVFDKDGNGFISAAELRHVMTNLGEKLTDEEVDEMIREAD 129
Query: 123 LDGDGLLSFDEFITMM 138
+DGDG ++++EF+ MM
Sbjct: 130 IDGDGQVNYEEFVAMM 145
Score = 56.2 bits (134), Expect = 2e-08
Identities = 24/67 (35%), Positives = 43/67 (63%)
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ + +EAF ++D + G IT K L +++ +G++ + E + MI D DG+G + F
Sbjct: 6 EEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMINEVDADGNGTIDF 65
Query: 132 DEFITMM 138
EF+TMM
Sbjct: 66 PEFLTMM 72
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.138 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,947,701
Number of Sequences: 164201
Number of extensions: 696491
Number of successful extensions: 5294
Number of sequences better than 10.0: 651
Number of HSP's better than 10.0 without gapping: 460
Number of HSP's successfully gapped in prelim test: 194
Number of HSP's that attempted gapping in prelim test: 2675
Number of HSP's gapped (non-prelim): 1827
length of query: 139
length of database: 59,974,054
effective HSP length: 99
effective length of query: 40
effective length of database: 43,718,155
effective search space: 1748726200
effective search space used: 1748726200
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC141863.5


