

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141436.5 + phase: 0 /pseudo
(364 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
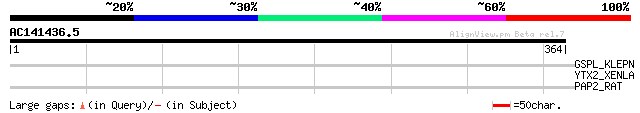
Score E
Sequences producing significant alignments: (bits) Value
GSPL_KLEPN (P15751) General secretion pathway protein L (Pullula... 31 4.3
YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein ... 30 7.3
PAP2_RAT (P35231) Pancreatitis-associated protein 2 precursor (L... 30 9.5
>GSPL_KLEPN (P15751) General secretion pathway protein L
(Pullulanase secretion protein pulL)
Length = 398
Score = 31.2 bits (69), Expect = 4.3
Identities = 14/38 (36%), Positives = 20/38 (51%), Gaps = 1/38 (2%)
Query: 11 DIKIWTHSPTGMYSVKSGYQWLTSNCLSFAGQTQQETW 48
D+ HSPTG +V+ G QWL C ++ G + W
Sbjct: 142 DVLALPHSPTGWSAVRCGEQWL-FRCDTWGGMAVETVW 178
>YTX2_XENLA (P14381) Transposon TX1 hypothetical 149 kDa protein (ORF
2)
Length = 1308
Score = 30.4 bits (67), Expect = 7.3
Identities = 16/63 (25%), Positives = 28/63 (44%), Gaps = 2/63 (3%)
Query: 44 QQETWQWIWKLHLPENLKHFIWLAMHGNLPTNAFRAGKHLTSDSSCKRCGASIESLLHTF 103
++ W+ + +P W +HG L T + A + S ++C CG ES+ H +
Sbjct: 1118 ERPQWRAFYSSLVPRPTGDLSWKVLHGALSTGEYLA-RFTDSPAACPFCGKG-ESVFHAY 1175
Query: 104 RDC 106
C
Sbjct: 1176 FTC 1178
>PAP2_RAT (P35231) Pancreatitis-associated protein 2 precursor
(Lithostathine 3) (Islet of langerhans regenerating
protein 3) (REG 3)
Length = 174
Score = 30.0 bits (66), Expect = 9.5
Identities = 13/33 (39%), Positives = 20/33 (60%), Gaps = 1/33 (3%)
Query: 8 SSQDIKIWTHSPT-GMYSVKSGYQWLTSNCLSF 39
++QDI IW H PT G G++W S+ L++
Sbjct: 97 NNQDIWIWLHDPTMGQQPNGGGWEWSNSDVLNY 129
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.337 0.147 0.506
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,668,820
Number of Sequences: 164201
Number of extensions: 1697579
Number of successful extensions: 5137
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 5137
Number of HSP's gapped (non-prelim): 3
length of query: 364
length of database: 59,974,054
effective HSP length: 112
effective length of query: 252
effective length of database: 41,583,542
effective search space: 10479052584
effective search space used: 10479052584
T: 11
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC141436.5


