

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141323.9 - phase: 0
(523 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
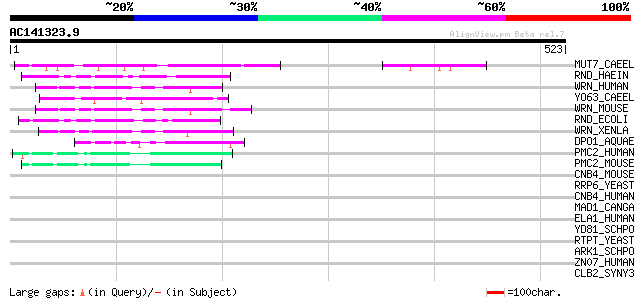
Score E
Sequences producing significant alignments: (bits) Value
MUT7_CAEEL (P34607) Probable exonuclease mut-7 (EC 3.1.-.-) 61 6e-09
RND_HAEIN (P44442) Ribonuclease D (EC 3.1.26.3) (RNase D) 55 3e-07
WRN_HUMAN (Q14191) Werner syndrome helicase 54 8e-07
YO63_CAEEL (P34603) Hypothetical protein ZK1098.3 in chromosome III 54 1e-06
WRN_MOUSE (O09053) Werner syndrome helicase homolog 53 2e-06
RND_ECOLI (P09155) Ribonuclease D (EC 3.1.26.3) (RNase D) 50 1e-05
WRN_XENLA (O93530) Werner syndrome helicase homolog (Focus formi... 47 9e-05
DPO1_AQUAE (O67779) DNA polymerase I (EC 2.7.7.7) (POL I) 47 9e-05
PMC2_HUMAN (Q01780) Polymyositis/scleroderma autoantigen 2 (Exos... 46 2e-04
PMC2_MOUSE (P56960) Polymyositis/scleroderma autoantigen 100 kDa... 44 8e-04
CNB4_MOUSE (Q8VEG4) Protein C14orf114 homolog 42 0.005
RRP6_YEAST (Q12149) Exosome complex exonuclease RRP6 (EC 3.1.13.... 39 0.033
CNB4_HUMAN (Q9NVH0) Protein C14orf114 38 0.056
MAD1_CANGA (Q6FS63) Spindle assembly checkpoint component MAD1 (... 36 0.28
ELA1_HUMAN (Q14241) Transcription elongation factor B polypeptid... 34 0.81
YD81_SCHPO (Q10146) Hypothetical protein C1F3.01 in chromosome I 34 1.1
RTPT_YEAST (P17558) 40S ribosomal protein PET123, mitochondrial ... 33 1.4
ARK1_SCHPO (O59790) Serine/threonine-protein kinase ark1 (EC 2.7... 33 2.4
ZN07_HUMAN (P17097) Zinc finger protein 7 (Zinc finger protein K... 32 3.1
CLB2_SYNY3 (P74361) Chaperone clpB 2 32 4.0
>MUT7_CAEEL (P34607) Probable exonuclease mut-7 (EC 3.1.-.-)
Length = 910
Score = 61.2 bits (147), Expect = 6e-09
Identities = 67/274 (24%), Positives = 115/274 (41%), Gaps = 43/274 (15%)
Query: 5 NQKPLEIHFVTTTDSPEFTHLTRSLTQTS------LVGLDAEWKP---VRTHQNSFPTVS 55
N++ +IH V T E +L + S VG D+EWKP H + +
Sbjct: 398 NERRTQIHMVKTES--EMNYLCSEIKSLSDEPAPVYVGFDSEWKPSNLTAVHDSKIAIIQ 455
Query: 56 LLQIACQLGDDEVVFLLDLISLPLSSI----WEPLREMLVSADILKL-GFRFKQDL---- 106
L C V+L+D + L +++ W+ L +K+ GF + DL
Sbjct: 456 LFFKNC-------VWLVDCVELEKANMADDWWQKFASRLFGDSPVKVVGFDMRNDLDAMA 508
Query: 107 --VYLSSTFCEQGCNPGFDK---VEPYLDITSVYNHLQFKKNGRIASKQNKSLSTICGEL 161
L S+ + FD E DI L K+ L+ + L
Sbjct: 509 TIPALKSSMKIEDTKNAFDLKRLAENVCDIDMEILELP---------KKTFKLADLTHYL 559
Query: 162 LGITLSKELQCSDWSQRPLTEEQMTYAAMDAHCLLGIFKVFQATVAKEGELVNKTNILSI 221
LG+ L K QCS+W RPL ++Q+ YAA+DA ++ FK + V ++ + + I +
Sbjct: 560 LGLELDKTEQCSNWQCRPLRKKQIVYAALDAVVVVETFKKILSIVEEKNKDADIEKI--V 617
Query: 222 RSANLGLKELFRKHDTSDKVHSTQFCEALAIVQA 255
R +N+ + + H + K+ + + E I+++
Sbjct: 618 RESNVMAPKKDKGHKSYRKLKTIPWLELYDILRS 651
Score = 48.9 bits (115), Expect = 3e-05
Identities = 32/109 (29%), Positives = 55/109 (50%), Gaps = 11/109 (10%)
Query: 352 KFLCDVMVEGLAKHLRCVGIDAAVP---SSKKPEPRELITKAQKEKRVLLTRDAK---LL 405
K + D M+ G K+LR VGID +P S + +E+ + R ++T +K L
Sbjct: 666 KVIVDTMLIGFGKNLRRVGIDVILPKDVSDFRKYLKEIERVGGEHLRHIITVPSKSYEAL 725
Query: 406 RHDYQIYKV-----KSLLKNEQLLEIIETFQLNINEDQLMSRCTKCNGR 449
+ DY Y + ++ +QL+E + F ++I + + RCT+CN R
Sbjct: 726 KMDYDNYTIAIPELNNMSPVDQLIEFFDLFNVDIRPEDVYPRCTECNSR 774
>RND_HAEIN (P44442) Ribonuclease D (EC 3.1.26.3) (RNase D)
Length = 399
Score = 55.5 bits (132), Expect = 3e-07
Identities = 50/198 (25%), Positives = 91/198 (45%), Gaps = 28/198 (14%)
Query: 12 HFVTTTDSPEFTHLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFL 71
HF TD+ + Q S V LD E+ V T+ FP + L+Q L D E V L
Sbjct: 29 HFRVVTDNTALLEVCNLAQQKSAVALDTEFMRVSTY---FPKLGLIQ----LYDGEHVSL 81
Query: 72 LDLISLPLSSIWEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKV-EPYLD 130
+D +++ + + P +L + +LK+ +DL+ F ++ FD++ P +D
Sbjct: 82 IDPLAI---TDFSPFVALLANPKVLKILHSCSEDLL----VFLQE-----FDQLPRPMID 129
Query: 131 ITSVYNHLQFKKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRPLTEEQMTYAAM 190
+ L + +A + + L + + K ++W +RPL++ Q+ YAA
Sbjct: 130 TQIMARFLGLGTSAGLAK--------LAQQYLNVEIDKGATRTNWIKRPLSDIQLQYAAG 181
Query: 191 DAHCLLGIFKVFQATVAK 208
D LL ++ + + +AK
Sbjct: 182 DVWYLLPLYHILEKELAK 199
>WRN_HUMAN (Q14191) Werner syndrome helicase
Length = 1432
Score = 54.3 bits (129), Expect = 8e-07
Identities = 47/178 (26%), Positives = 88/178 (49%), Gaps = 23/178 (12%)
Query: 25 LTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFLLDLISLPLSSIWE 84
++ SL+ +VG D EW P+ ++ V+L+Q+ + +L + S+ S +
Sbjct: 69 ISMSLSDGDVVGFDMEWPPLY-NRGKLGKVALIQLCVS---ESKCYLFHVSSM--SVFPQ 122
Query: 85 PLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPYLDITSVYNHLQFKKNG 144
L+ +L + + K G + D L F K++ ++++T V N
Sbjct: 123 GLKMLLENKAVKKAGVGIEGDQWKLLRDFDI--------KLKNFVELTDVANK------- 167
Query: 145 RIASKQNKSLSTICGELLGITLSKE--LQCSDWSQRPLTEEQMTYAAMDAHCLLGIFK 200
++ + SL+++ LLG L K+ ++CS+WS+ PLTE+Q YAA DA+ I++
Sbjct: 168 KLKCTETWSLNSLVKHLLGKQLLKDKSIRCSNWSKFPLTEDQKLYAATDAYAGFIIYR 225
>YO63_CAEEL (P34603) Hypothetical protein ZK1098.3 in chromosome III
Length = 784
Score = 53.9 bits (128), Expect = 1e-06
Identities = 43/186 (23%), Positives = 87/186 (46%), Gaps = 18/186 (9%)
Query: 29 LTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFLLDLISLP----LSSIWE 84
L + +G D+E+KP S ++++Q+ + + +L++ +++ +W
Sbjct: 441 LEEGMYIGYDSEFKPYHLIDVSTSRLAIIQLFFK----DKAWLINCVAIDNLASRDDVWI 496
Query: 85 PLREMLVSADILKL-GFRFKQDLVYLSSTFCEQGCNPGF--DKVEPYLDITSVYNHLQFK 141
L + L ++ + GF +QD+ + F N F + ++ + + S+ ++
Sbjct: 497 RLYKGLFESNKFSIVGFDIRQDI---EAMFTVPSINKNFKIENIQNVICVKSLAENVNAL 553
Query: 142 KNGRI-ASKQNKSLSTICGELLGITLSKELQCSDWSQRPLTEEQMTYAAMDAHCLLGIFK 200
+ S + LS + L+G+ + K QC +W RPL Q+ YA MDA + +F+
Sbjct: 554 SMDILNLSTKTSKLSVLADHLVGLKMDKSEQCGNWQCRPLRRNQIIYAVMDA---VAVFE 610
Query: 201 VFQATV 206
VFQ V
Sbjct: 611 VFQKIV 616
Score = 33.9 bits (76), Expect = 1.1
Identities = 42/167 (25%), Positives = 74/167 (44%), Gaps = 19/167 (11%)
Query: 287 LLKIVRKH--GDRILLKESDRAPKTSKKKRKKQLPINGIPKEKHLENFDEWQGTAPWDPL 344
++++VRKH LL ES +K R+ I+ IP + + + P PL
Sbjct: 615 IVEVVRKHELDAEKLLVESHMITVKKEKVRRDCKNISLIPWNEFYQIIHTHRN--PEKPL 672
Query: 345 VGGDGFPKFLCDVMVEGLAKHLRCVGIDAAVPSSKKPEPRELITKAQK----EKRVLL-- 398
K + D MV GL K+LR +G D +P E +E + K K E+R+++
Sbjct: 673 QKPSEL-KIVVDTMVLGLGKNLRLLGFDVYIPRD-VTELKEFLRKMDKMEESEQRLVISV 730
Query: 399 -TRDAKLLRHD------YQIYKVKSLLKNEQLLEIIETFQLNINEDQ 438
+R ++L+ D I + + + + + F ++I+ DQ
Sbjct: 731 PSRSYEMLKSDNPNAKFVLIPNIYEKVPIDLVCSFFDFFNIDISPDQ 777
>WRN_MOUSE (O09053) Werner syndrome helicase homolog
Length = 1401
Score = 53.1 bits (126), Expect = 2e-06
Identities = 52/206 (25%), Positives = 93/206 (44%), Gaps = 30/206 (14%)
Query: 25 LTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFLLDLISLPLSSIWE 84
++ L+ +VG D EW P+ V+++Q+ + +L + S+ S +
Sbjct: 63 ISMRLSDGDVVGFDMEWPPIYK-PGKRSRVAVIQLCVS---ENKCYLFHISSM--SVFPQ 116
Query: 85 PLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPYLDITSVYNHLQFKKNG 144
L+ +L + I K G + D L F K+E ++++T V N
Sbjct: 117 GLKMLLENKSIKKAGVGIEGDQWKLLRDFDV--------KLESFVELTDVANE------- 161
Query: 145 RIASKQNKSLSTICGELLGITLSKE--LQCSDWSQRPLTEEQMTYAAMDAHCLLGIFKVF 202
++ + SL+ + +LG L K+ ++CS+WS PLTE+Q YAA DA+ L I++
Sbjct: 162 KLKCAETWSLNGLVKHVLGKQLLKDKSIRCSNWSNFPLTEDQKLYAATDAYAGLIIYQ-- 219
Query: 203 QATVAKEGELVNKTNILSIRSANLGL 228
K G L + + ++ A L
Sbjct: 220 -----KLGNLGDTVQVFALNKAEENL 240
>RND_ECOLI (P09155) Ribonuclease D (EC 3.1.26.3) (RNase D)
Length = 375
Score = 50.1 bits (118), Expect = 1e-05
Identities = 52/190 (27%), Positives = 80/190 (41%), Gaps = 28/190 (14%)
Query: 9 LEIHFVTTTDSPEFTHLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEV 68
+ +TT D+ L ++ + LD E+ RT+ +P + L+Q L D E
Sbjct: 1 MNYQMITTDDA--LASLCEAVRAFPAIALDTEFVRTRTY---YPQLGLIQ----LFDGEH 51
Query: 69 VFLLDLISLPLSSIWEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPY 128
+ L+D + + + W PL+ +L I K +DL + F E +P
Sbjct: 52 LALIDPLGI---TDWSPLKAILRDPSITKFLHAGSEDLEVFLNVFGELP--------QPL 100
Query: 129 LDITSVYNHLQFKKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRPLTEEQMTYA 188
+D + GR S +++ E G+TL K +DW RPLTE Q YA
Sbjct: 101 IDTQILAAFC-----GRPMSW---GFASMVEEYSGVTLDKSESRTDWLARPLTERQCEYA 152
Query: 189 AMDAHCLLGI 198
A D LL I
Sbjct: 153 AADVWYLLPI 162
>WRN_XENLA (O93530) Werner syndrome helicase homolog (Focus forming
activity 1)
Length = 1436
Score = 47.4 bits (111), Expect = 9e-05
Identities = 44/186 (23%), Positives = 85/186 (45%), Gaps = 23/186 (12%)
Query: 28 SLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFLLDLISLPLSSIWEPLR 87
SL + ++G D EW PV T + V+L+Q+ ++ +L + P++ + L+
Sbjct: 66 SLLEEDVLGFDIEWPPVYT-KGKTGKVALIQVCVS---EKKCYLFHIS--PMAGFPKGLK 119
Query: 88 EMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPYLDITSVYNHLQFKKNGRIA 147
+L + K+G + D L S + K++ +++++ + N ++
Sbjct: 120 RLLEDESVRKVGVGIEGDQWKLMSDYEL--------KLKGFIELSEMANQ-------KLR 164
Query: 148 SKQNKSLSTICGELLGITL--SKELQCSDWSQRPLTEEQMTYAAMDAHCLLGIFKVFQAT 205
K+ + + + L L K +CS+W LTE+Q YAA DA+ L I+K +
Sbjct: 165 CKEKWTFNGLIKHLFKEQLYKRKSYRCSNWDIFLLTEDQKLYAATDAYAGLLIYKKLEGM 224
Query: 206 VAKEGE 211
A E +
Sbjct: 225 DAHESD 230
>DPO1_AQUAE (O67779) DNA polymerase I (EC 2.7.7.7) (POL I)
Length = 574
Score = 47.4 bits (111), Expect = 9e-05
Identities = 48/166 (28%), Positives = 73/166 (43%), Gaps = 31/166 (18%)
Query: 62 QLGDDEVVFLLDLISLPLSSIWEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPG 121
Q+GD+E +++DL + EPLR+++ I+ G K DL YL G P
Sbjct: 40 QIGDEENTYVIDLYEI---QDIEPLRKLINERGIV--GHNLKFDLKYLY----RYGIFPS 90
Query: 122 --FDKVEPYLDITSVYNHLQFKKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRP 179
FD T + ++L + SL+ I LLG ++ K Q SDW
Sbjct: 91 ATFD--------TMIASYL--------LGYERHSLNHIVSNLLGYSMDKSYQTSDWGASV 134
Query: 180 LTEEQMTYAAMDAHCLLGIFKVFQATV----AKEGELVNKTNILSI 221
L++ Q+ YAA D L +F + + A+ GE + KT I
Sbjct: 135 LSDAQLKYAANDVIVLRELFPKMRDMLNELDAERGEELLKTRTAKI 180
>PMC2_HUMAN (Q01780) Polymyositis/scleroderma autoantigen 2 (Exosome
component 10) (Autoantigen PM/Scl 2)
(Polymyositis/scleroderma autoantigen 100 kDa)
(PM/Scl-100) (P100 polymyositis-scleroderma overlap
syndrome associated autoantigen)
Length = 885
Score = 46.2 bits (108), Expect = 2e-04
Identities = 50/211 (23%), Positives = 84/211 (39%), Gaps = 31/211 (14%)
Query: 3 PPNQKPLE---IHFVTTTDSPEFTHLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQI 59
P +P+E HF+++ D E L L +D E ++++ L+QI
Sbjct: 277 PQLYRPIEETPCHFISSLD--ELVELNEKLLNCQEFAVDLEH---HSYRSFLGLTCLMQI 331
Query: 60 ACQLGDDEVVFLLDLISLPLSSIWEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCN 119
+ + D F++D +L L S L E L I+K+ D+ +L F
Sbjct: 332 STRTED----FIID--TLELRSDMYILNESLTDPAIVKVFHGADSDIEWLQKDFG----- 380
Query: 120 PGFDKVEPYLDITSVYNHLQFKKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRP 179
V N + R+ + SL + + +K+ Q +DW RP
Sbjct: 381 ------------LYVVNMFDTHQAARLLNLGRHSLDHLLKLYCNVDSNKQYQLADWRIRP 428
Query: 180 LTEEQMTYAAMDAHCLLGIFKVFQATVAKEG 210
L EE ++YA D H LL I+ + + + G
Sbjct: 429 LPEEMLSYARDDTHYLLYIYDKMRLEMWERG 459
>PMC2_MOUSE (P56960) Polymyositis/scleroderma autoantigen 100 kDa
(Exosome component 10) (Autoantigen PM/Scl 2) (P100
polymyositis-scleroderma overlap syndrome associated
autoantigen homolog)
Length = 887
Score = 44.3 bits (103), Expect = 8e-04
Identities = 47/188 (25%), Positives = 75/188 (39%), Gaps = 28/188 (14%)
Query: 12 HFVTTTDSPEFTHLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFL 71
H V++ D E L L +D E ++++ L+QI+ + D F+
Sbjct: 289 HLVSSLD--ELVELNEKLLGCQEFAVDLEH---HSYRSFLGLTCLMQISTRTED----FI 339
Query: 72 LDLISLPLSSIWEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPYLDI 131
+D +L L S L E L I+K+ D+ +L F
Sbjct: 340 VD--TLELRSDMYILNESLTDPAIVKVFHGADSDIEWLQKDFG----------------- 380
Query: 132 TSVYNHLQFKKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRPLTEEQMTYAAMD 191
V N + R+ + SL + G+ +K+ Q +DW RPL EE ++YA D
Sbjct: 381 LYVVNMFDTHQAARLLNLARHSLDHLLRLYCGVESNKQYQLADWRIRPLPEEMLSYARDD 440
Query: 192 AHCLLGIF 199
H LL I+
Sbjct: 441 THYLLYIY 448
>CNB4_MOUSE (Q8VEG4) Protein C14orf114 homolog
Length = 496
Score = 41.6 bits (96), Expect = 0.005
Identities = 35/118 (29%), Positives = 53/118 (44%), Gaps = 19/118 (16%)
Query: 86 LREMLVSADILKLGFRFKQDLVYLSSTF--CEQGCNPGFDKVEPYLDITSVYNHLQFKKN 143
L ++L ILK+G +D L + +GC LD+ +L K+
Sbjct: 27 LLDILADGAILKVGVGCSEDANKLLQDYGLIVRGC----------LDL----RYLAMKQG 72
Query: 144 GRIASKQNKSLSTICGELLGITLSKEL--QCSDWSQRPLTEEQMTYAAMDAHCLLGIF 199
I SL ++ +L L K L +CS+W LTE+Q+TYAA DA + +F
Sbjct: 73 NNILCN-GLSLKSLAETILNFPLDKSLLLRCSNWDAENLTEDQVTYAARDAQISVALF 129
>RRP6_YEAST (Q12149) Exosome complex exonuclease RRP6 (EC 3.1.13.-)
(Ribosomal RNA processing protein 6)
Length = 733
Score = 38.9 bits (89), Expect = 0.033
Identities = 42/194 (21%), Positives = 76/194 (38%), Gaps = 26/194 (13%)
Query: 19 SPEFTHLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFLLDLISLP 78
S E + L T + +D E R++ + V L+QI+ + D +L+D +L
Sbjct: 219 STELESMLEDLKNTKEIAVDLEHHDYRSY---YGIVCLMQISTRERD----YLVD--TLK 269
Query: 79 LSSIWEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPYLDITSVYNHL 138
L L E+ + I+K+ D+++L L + +++
Sbjct: 270 LRENLHILNEVFTNPSIVKVFHGAFMDIIWLQRDLG--------------LYVVGLFDTY 315
Query: 139 QFKKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRPLTEEQMTYAAMDAHCLLGI 198
K SL+ + SK+ Q +DW RPL++ YA D H LL I
Sbjct: 316 HASK---AIGLPRHSLAYLLENFANFKTSKKYQLADWRIRPLSKPMTAYARADTHFLLNI 372
Query: 199 FKVFQATVAKEGEL 212
+ + + + +L
Sbjct: 373 YDQLRNKLIESNKL 386
>CNB4_HUMAN (Q9NVH0) Protein C14orf114
Length = 496
Score = 38.1 bits (87), Expect = 0.056
Identities = 19/49 (38%), Positives = 28/49 (56%), Gaps = 2/49 (4%)
Query: 153 SLSTICGELLGITLSKEL--QCSDWSQRPLTEEQMTYAAMDAHCLLGIF 199
SL ++ +L L K L +CS+W LTE+Q+ YAA DA + +F
Sbjct: 81 SLKSLAETVLNFPLDKSLLLRCSNWDAETLTEDQVIYAARDAQISVALF 129
>MAD1_CANGA (Q6FS63) Spindle assembly checkpoint component MAD1
(Mitotic arrest deficient protein 1)
Length = 657
Score = 35.8 bits (81), Expect = 0.28
Identities = 28/92 (30%), Positives = 41/92 (44%), Gaps = 2/92 (2%)
Query: 392 KEKRVLLTRDAKLLRHDYQIYKVKSLLK--NEQLLEIIETFQLNINEDQLMSRCTKCNGR 449
K R+L RD+ LL+ + K LL+ N+ LL +IE N NE T N
Sbjct: 482 KNIRILQQRDSPLLKLQFVRRKENELLREENQNLLTMIENITENRNESFDKIPLTSYNSL 541
Query: 450 FIQKPLSTEEAIEAAKGFQKIPNCLFNKNLEF 481
Q +S + + K F ++ K+LEF
Sbjct: 542 KYQYEISMDTNKKQEKKFSRLKQIFNKKSLEF 573
>ELA1_HUMAN (Q14241) Transcription elongation factor B polypeptide 3
(RNA polymerase II transcription factor SIII subunit A1)
(SIII p110) (Elongin A) (EloA) (Elongin 110 kDa subunit)
(MSTP059)
Length = 772
Score = 34.3 bits (77), Expect = 0.81
Identities = 31/104 (29%), Positives = 44/104 (41%), Gaps = 13/104 (12%)
Query: 235 HDTSDKVHSTQFCEALAIVQATSCSDVVRVISSAGAVIQ----KSSCRDTKPMD----EF 286
H K HS F + L Q + G V Q KSS +D +P+D E
Sbjct: 207 HQKPGKGHSNAFQDRLGASQERHLGEP----HGKGVVSQNKEHKSSHKDKRPVDAKSDEK 262
Query: 287 LLKIVRKHGDRILLKESDRAPKTSKKKRKKQLPINGIPKEKHLE 330
+ R+ + L KE +R P + R+K P +G+ KEK E
Sbjct: 263 ASVVSREKSHKALSKEENRRPPSGDNAREKP-PSSGVKKEKDRE 305
>YD81_SCHPO (Q10146) Hypothetical protein C1F3.01 in chromosome I
Length = 777
Score = 33.9 bits (76), Expect = 1.1
Identities = 37/179 (20%), Positives = 74/179 (40%), Gaps = 26/179 (14%)
Query: 21 EFTHLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFLLDLISLPLS 80
+ + + + L + + +D E R+ + V L+QI+ + D +++D +L L
Sbjct: 226 QLSDMLKELQNSKEIAVDLEHHDYRSFRGF---VCLMQISNREKD----WIVD--TLELR 276
Query: 81 SIWEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPYLDITSVYNHLQF 140
E L + + +I+K+ D+++L F L + ++++
Sbjct: 277 EELEALNVVFTNPNIIKVFHGATMDIIWLQRDFG--------------LYVVNLFDTYYA 322
Query: 141 KKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRPLTEEQMTYAAMDAHCLLGIF 199
K + + L+ + + K Q +DW RPL E + YA D H LL I+
Sbjct: 323 TK---VLGFEGHGLAFLLQKYCDYDADKRYQMADWRIRPLPREMLKYAQSDTHYLLYIW 378
>RTPT_YEAST (P17558) 40S ribosomal protein PET123, mitochondrial
precursor
Length = 318
Score = 33.5 bits (75), Expect = 1.4
Identities = 32/133 (24%), Positives = 57/133 (42%), Gaps = 12/133 (9%)
Query: 201 VFQATVAKEGELVNKTN------ILSIRSANLGLKELFRKHDTSDKVHSTQFCEALAIVQ 254
+ ++ K+ +++NK +L I A G+K H S KV+ +A
Sbjct: 21 ILKSPTTKQTDIINKVKSPKPKGVLGIGYAK-GVKHPKGSHRLSPKVNFIDVDNLIAKTV 79
Query: 255 ATSCSDVVRVISSAGAVIQKSSCRDTKPMDEFLLKIVRKHGDRILLKESDRAPKTSKKKR 314
A S I S+ QK + + +FL++ RK R+L K +T + ++
Sbjct: 80 AEPQS-----IKSSNGSAQKVRLQKAELRRKFLIEAFRKEEARLLHKHEYLQKRTKELEK 134
Query: 315 KKQLPINGIPKEK 327
K+L + + KEK
Sbjct: 135 AKELELEKLNKEK 147
>ARK1_SCHPO (O59790) Serine/threonine-protein kinase ark1 (EC
2.7.1.37) (Aurora-related kinase 1)
Length = 355
Score = 32.7 bits (73), Expect = 2.4
Identities = 27/96 (28%), Positives = 45/96 (46%), Gaps = 12/96 (12%)
Query: 318 LPINGIPKEKHLENFDEWQ-GTAPWDPLVGGDGFPKFLCDVMVEGLAKHLRCVGIDAAVP 376
LP + ++H E D W G ++ LVG F + M A + R +D +P
Sbjct: 252 LPPEMVEGKEHTEKVDLWSLGVLTYEFLVGAPPF-----EDMSGHSATYKRIAKVDLKIP 306
Query: 377 SSKKPEPRELITKA---QKEKRVLLTRDAKLLRHDY 409
S P+ R+LI++ EKR+ L +++RH +
Sbjct: 307 SFVPPDARDLISRLLQHNPEKRMSL---EQVMRHPW 339
>ZN07_HUMAN (P17097) Zinc finger protein 7 (Zinc finger protein
KOX4) (Zinc finger protein HF.16)
Length = 686
Score = 32.3 bits (72), Expect = 3.1
Identities = 27/97 (27%), Positives = 43/97 (43%), Gaps = 1/97 (1%)
Query: 168 KELQCSDWSQRPLTEEQMTYAAMDAHCLLGIFKVFQATVAKEGELVNKTNILSIRSANLG 227
+E QC D QR L E M L G F VF+ + E + +L ++ A
Sbjct: 16 EEWQCLDPGQRALYREVMLENHSSVAGLAG-FLVFKPELISRLEQGEEPWVLDLQGAEGT 74
Query: 228 LKELFRKHDTSDKVHSTQFCEALAIVQATSCSDVVRV 264
K D++ + + Q CE + I+++ S VVR+
Sbjct: 75 EAPRTSKTDSTIRTENEQACEDMDILKSESYGTVVRI 111
>CLB2_SYNY3 (P74361) Chaperone clpB 2
Length = 872
Score = 32.0 bits (71), Expect = 4.0
Identities = 30/119 (25%), Positives = 55/119 (46%), Gaps = 19/119 (15%)
Query: 212 LVNKTNILSIRSANLGLKELFRKHDTSDKVHSTQFCEALAIVQATSCSD----------- 260
LV++ N+L S GLKE + H H + ++ A+V A S+
Sbjct: 339 LVDEPNVLDTISILRGLKERYEVH------HGVKIADS-ALVAAAMLSNRYISDRFLPDK 391
Query: 261 VVRVISSAGAVIQKSSCRDTKPMDEFLLKIVRKHGDRI-LLKESDRAPKTSKKKRKKQL 318
+ ++ A A ++ + +DE KI++ +R+ L +E+D A K +K +K+L
Sbjct: 392 AIDLVDEAAAKLKMEITSKPEELDEVDRKILQLEMERLSLQRENDSASKERLEKLEKEL 450
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 60,984,782
Number of Sequences: 164201
Number of extensions: 2567205
Number of successful extensions: 6167
Number of sequences better than 10.0: 23
Number of HSP's better than 10.0 without gapping: 14
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 6144
Number of HSP's gapped (non-prelim): 26
length of query: 523
length of database: 59,974,054
effective HSP length: 115
effective length of query: 408
effective length of database: 41,090,939
effective search space: 16765103112
effective search space used: 16765103112
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 68 (30.8 bits)
Medicago: description of AC141323.9


