

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141111.6 - phase: 0 /pseudo
(379 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
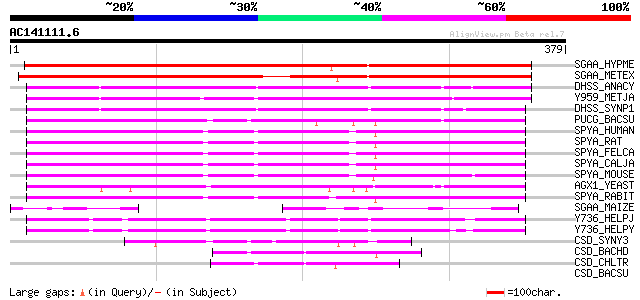
SGAA_HYPME (O08374) Serine--glyoxylate aminotransferase (EC 2.6.... 383 e-106
SGAA_METEX (P55819) Serine--glyoxylate aminotransferase (EC 2.6.... 374 e-103
DHSS_ANACY (P16421) Soluble hydrogenase 42 kDa subunit (EC 1.12.... 206 6e-53
Y959_METJA (Q58369) Putative aminotransferase MJ0959 (EC 2.6.1.-) 203 5e-52
DHSS_SYNP1 (P14776) Soluble hydrogenase, small subunit (EC 1.12.... 188 2e-47
PUCG_BACSU (O32148) Purine catabolism protein pucG (EC 2.-.-.-) 144 3e-34
SPYA_HUMAN (P21549) Serine--pyruvate aminotransferase (EC 2.6.1.... 130 4e-30
SPYA_RAT (P09139) Serine--pyruvate aminotransferase, mitochondri... 125 1e-28
SPYA_FELCA (P41689) Serine--pyruvate aminotransferase, mitochond... 125 2e-28
SPYA_CALJA (P31029) Serine--pyruvate aminotransferase, mitochond... 123 7e-28
SPYA_MOUSE (O35423) Serine--pyruvate aminotransferase, mitochond... 119 1e-26
AGX1_YEAST (P43567) Alanine--glyoxylate aminotransferase 1 (EC 2... 116 1e-25
SPYA_RABIT (P31030) Serine--pyruvate aminotransferase (EC 2.6.1.... 115 2e-25
SGAA_MAIZE (P84187) Serine--glyoxylate aminotransferase (EC 2.6.... 94 7e-19
Y736_HELPJ (Q9ZLA7) Putative aminotransferase JHP0673 (EC 2.6.1.-) 79 1e-14
Y736_HELPY (O25436) Putative aminotransferase HP0736 (EC 2.6.1.-) 78 4e-14
CSD_SYNY3 (Q55793) Probable cysteine desulfurase (EC 2.8.1.7) 47 6e-05
CSD_BACHD (Q9K7A0) Probable cysteine desulfurase (EC 2.8.1.7) 47 1e-04
CSD_CHLTR (O84693) Probable cysteine desulfurase (EC 2.8.1.7) 44 9e-04
CSD_BACSU (O32164) Probable cysteine desulfurase (EC 2.8.1.7) 42 0.003
>SGAA_HYPME (O08374) Serine--glyoxylate aminotransferase (EC
2.6.1.45) (SGAT)
Length = 404
Score = 383 bits (983), Expect = e-106
Identities = 191/348 (54%), Positives = 244/348 (69%), Gaps = 3/348 (0%)
Query: 11 HLFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGT 70
HLF+PGP NIPD V AM+ ED RSP P T L ED+KK FK G F+ P++GT
Sbjct: 5 HLFIPGPTNIPDAVRMAMNIPMEDMRSPEFPKFTLPLFEDLKKAFKMKDGRVFIFPSSGT 64
Query: 71 GAWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDILESKL 130
GAWESA+ NTL+ GD+++ GQFSLLWVD +RL V+V + EWG G ++ L
Sbjct: 65 GAWESAVENTLATGDKVLMSRFGQFSLLWVDMCERLGLKVEVCDEEWGTGVPVEKYADIL 124
Query: 131 ASDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEW 190
A D H IKA+ + HNETATGV++++A VR+ LDA +HPALL+VDGVSS+ +LD RM EW
Sbjct: 125 AKDKNHEIKAVFVTHNETATGVSSDVAGVRKALDAAKHPALLMVDGVSSVGSLDMRMGEW 184
Query: 191 GVDVAITGSQKALSLPTGIGFVVASPKA--IEASKSAKSLRVFFDWSDYLKFYKMGTYWP 248
GVD ++GSQK LPTG+G + S KA I SK+ + R FF + D +K G ++P
Sbjct: 185 GVDCCVSGSQKGFMLPTGLGILAVSQKALDINKSKNGRMNRCFFSFEDMIKTNDQG-FFP 243
Query: 249 YTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQEEEWFSDTV 308
YTP+ QLL GLR +LDL+F EGL+N+ ARH RL + R AV+AWGLK C +E +W+SDTV
Sbjct: 244 YTPATQLLRGLRTSLDLLFAEGLDNVFARHTRLASGVRAAVDAWGLKLCAKEPKWYSDTV 303
Query: 309 TAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGNLNE 356
+A++VP ID I + A+ RYN S GLGLNKVAGKVFRIGHLG L+E
Sbjct: 304 SAILVPEGIDSNAITKTAYYRYNTSFGLGLNKVAGKVFRIGHLGMLDE 351
>SGAA_METEX (P55819) Serine--glyoxylate aminotransferase (EC
2.6.1.45) (SGAT)
Length = 379
Score = 374 bits (959), Expect = e-103
Identities = 192/355 (54%), Positives = 243/355 (68%), Gaps = 24/355 (6%)
Query: 7 PGRNHLFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIP 66
PGRNHLFVPGP NIPD+V+RAM +ED+RS P+LTK L ED KK+F +T GT FL P
Sbjct: 2 PGRNHLFVPGPTNIPDRVMRAMMVQSEDHRSVDFPSLTKPLFEDTKKVFGSTEGTIFLFP 61
Query: 67 TTGTGAWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDIL 126
+GTG WESAL+NTL+ GD++++ GQFS LW+D QRL +V V E EWG GA + +
Sbjct: 62 ASGTGIWESALSNTLARGDKVLAARFGQFSHLWIDMAQRLGLDVVVQEEEWGTGAKPEKI 121
Query: 127 ESKLASDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFR 186
E L +D H IKA+ +VHNETATGVT+N+ VR+ +DA HPALL
Sbjct: 122 EEALRADKNHEIKAVMVVHNETATGVTSNIGAVRKAIDAAGHPALLF------------- 168
Query: 187 MDEWGVDVAITGSQKALSLPTGIGFVVASPKAIEAS-----KSAKSLRVFFDWSDYLKFY 241
VD AI GSQK L LP G+G + S KA++A+ ++ + RV+FDW D K
Sbjct: 169 -----VDCAIAGSQKGLMLPAGLGVICVSQKALKAAEGQSGRNDRLARVYFDWEDQKKQN 223
Query: 242 KMGTYWPYTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQEE 301
G Y+PYTP + LLYGLR AL +FEEGLEN+ RH LG ATR AV AWGLK C +
Sbjct: 224 PTG-YFPYTPPLPLLYGLREALACLFEEGLENVYHRHAVLGEATRQAVAAWGLKTCAKSP 282
Query: 302 EWFSDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGNLNE 356
EW SDTVTA++ P +D A+I++ A+ RYNL+LG GL++VAGKVFRIGH+G+LNE
Sbjct: 283 EWNSDTVTAILAPEGVDAAKIIKHAYVRYNLALGAGLSQVAGKVFRIGHVGDLNE 337
>DHSS_ANACY (P16421) Soluble hydrogenase 42 kDa subunit (EC
1.12.-.-) (Tritium exchange subunit)
Length = 383
Score = 206 bits (525), Expect = 6e-53
Identities = 120/345 (34%), Positives = 198/345 (56%), Gaps = 5/345 (1%)
Query: 12 LFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTG 71
L +PGP +P+ + A++++ +R+ + + +++K + +T + ++ +GTG
Sbjct: 7 LMIPGPTPVPEAALLALAKHPIGHRTSEFSNMMGEVTQNLKWLHQTESDV-LMLNVSGTG 65
Query: 72 AWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDILESKLA 131
A E+ + N LSPGDRI+ G+F WV+ Q NV+ + +EWG+ D D KL
Sbjct: 66 AVEAGMINFLSPGDRILVGSNGKFGERWVEVGQAFGLNVEAITAEWGQPLDPDKFAQKLQ 125
Query: 132 SDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWG 191
+D+ IKA+ I H+ET+TGV N+L + + + AL++VD V+S+ A + +D G
Sbjct: 126 ADTNKEIKAVIITHSETSTGVINDLVAINSHVKEHGQ-ALIIVDAVTSLGAYNVPVDALG 184
Query: 192 VDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMGTYWPYTP 251
+DV +GSQK +P G+GFV SPKA EA K+AK + + D Y K T P+TP
Sbjct: 185 LDVVASGSQKGYMIPPGLGFVSVSPKAWEAYKTAKLPKYYLDLGKYRKATAKNT-TPFTP 243
Query: 252 SIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQEEEWFSDTVTAV 311
+ L+ L L ++ +EGLE+I RH R ATR A++A L +E S +TAV
Sbjct: 244 PVNLMVALHTTLGMMKKEGLESIFTRHERQKNATRAAMKALNLP-LFAADECASPAITAV 302
Query: 312 VVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGNLNE 356
P ++ +I KR++++L G + ++ K+FR+GHLG +++
Sbjct: 303 ATPG-MEADKIRSLMKKRFDIALAGGQDHLSNKIFRVGHLGFVSD 346
>Y959_METJA (Q58369) Putative aminotransferase MJ0959 (EC 2.6.1.-)
Length = 385
Score = 203 bits (517), Expect = 5e-52
Identities = 121/346 (34%), Positives = 193/346 (54%), Gaps = 7/346 (2%)
Query: 12 LFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTG 71
L +PGP +P +V+ AM+ +R+ L + +E +KK+F T T FLI +GT
Sbjct: 10 LMIPGPTMVPPEVLNAMALPVIGHRTKDYSNLLEDTIEKLKKVFITENDT-FLITGSGTA 68
Query: 72 AWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDILESKLA 131
A + A++N + GD++++ V G F + + + K ++ EWG A+ + ++ L
Sbjct: 69 AMDMAISNIIKRGDKVLNIVTGNFGERFANIVKAYKGEAIRLDVEWGDMAEPEAVKEIL- 127
Query: 132 SDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWG 191
D IKA+ +VHNET+TG N + ++ +++ Y AL +VD VSS+ +D++
Sbjct: 128 -DKYDDIKAVTVVHNETSTGARNPIKEIGEVVKDYD--ALYIVDTVSSLGGDYVNVDKFH 184
Query: 192 VDVAITGSQKALSLPTGIGFVVASPKAIEA-SKSAKSLRVFFDWSDYLKFYKMGTYWPYT 250
+D+ +TGSQK L+ P G+ + S KA E K+ + + D Y K+Y+ PYT
Sbjct: 185 IDICVTGSQKCLAAPPGLAAITVSEKAWEVIKKNDDKVGFYLDLLAYKKYYEEKKQTPYT 244
Query: 251 PSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQEEEWFSDTVTA 310
PS+ L Y L ALDL+ EEG+EN + RH RL ATR +EA G++ +E S TVT+
Sbjct: 245 PSVNLTYALNVALDLVLEEGIENRVKRHERLAKATRAGLEAMGIELFAKERA-RSVTVTS 303
Query: 311 VVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGNLNE 356
P I+ ++ +YN+ + G +AGK+FRIGH+G E
Sbjct: 304 AKYPEGIEDSKFRGILSNKYNIVVAGGQKHLAGKIFRIGHMGICGE 349
>DHSS_SYNP1 (P14776) Soluble hydrogenase, small subunit (EC
1.12.-.-) (Tritium exchange subunit)
Length = 384
Score = 188 bits (477), Expect = 2e-47
Identities = 113/341 (33%), Positives = 185/341 (54%), Gaps = 5/341 (1%)
Query: 12 LFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTG 71
L +PGP +P+ V+ ++ ++ +RS + + +K + +T ++ +GTG
Sbjct: 7 LMIPGPTPVPESVLLSLGKHPIGHRSGEFSQIMAAMTAGIKWLHQTQNEV-LILAASGTG 65
Query: 72 AWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDILESKLA 131
A E+ + N LS GDR+V G+F W + + + + WG+ + D ++ L
Sbjct: 66 AMEAGIINFLSAGDRVVVGCNGKFGDRWGEVCDAYGLTTERISAPWGQPLNPDDFKAVLD 125
Query: 132 SDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWG 191
KA+ + H+ET+TGV N+L + + + A+ AL++VD V+S+ A+ +DEWG
Sbjct: 126 GHRQKPSKAVIVTHSETSTGVINDLEAINRHVKAHGQ-ALIIVDAVTSLGAVSVPIDEWG 184
Query: 192 VDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMGTYWPYTP 251
+DV +GSQK +P G+ FV SPKA EA K+A + + D Y K T P+TP
Sbjct: 185 LDVVGSGSQKGYMIPPGLAFVSVSPKAWEAYKTATLPKFYLDLGKYRKDAAKHT-TPFTP 243
Query: 252 SIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQEEEWFSDTVTAV 311
+ L + L+ AL+++ EGLE I RH RL ATR A++A L + S +TA
Sbjct: 244 PVNLFFALKTALEMMQAEGLEAIFQRHQRLMQATRAAMKALNLP-LYAADSCASPAITA- 301
Query: 312 VVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLG 352
V P ++ I KR++++L G + + G++FRIGHLG
Sbjct: 302 VAPQGVEAENIRSLMKKRFDIALAGGQDHLKGQIFRIGHLG 342
>PUCG_BACSU (O32148) Purine catabolism protein pucG (EC 2.-.-.-)
Length = 416
Score = 144 bits (364), Expect = 3e-34
Identities = 97/361 (26%), Positives = 170/361 (46%), Gaps = 26/361 (7%)
Query: 12 LFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTG 71
+ PGPV + +V+R MS PA + +E ++++F+T + I T
Sbjct: 14 IMTPGPVEVDPRVLRVMSTPVVGQFDPAFTGIMNETMEMLRELFQTKNRWAYPIDGTSRA 73
Query: 72 AWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDILESKLA 131
E+ L + + P D ++ + G+F L + +R NV ++E EWG D + + ++
Sbjct: 74 GIEAVLASVIEPEDDVLIPIYGRFGYLLTEIAERYGANVHMLECEWGTVFDPEDIIREIK 133
Query: 132 SDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWG 191
K + +VH ET+TG + L + + AL +VD V++I + ++DEW
Sbjct: 134 KVKP---KIVAMVHGETSTGRIHPLKAIGEA--CRTEDALFIVDAVATIGGCEVKVDEWK 188
Query: 192 VDVAITGSQKALSLPTG---------IGFVVASPKAIEASKSAKSLRVFFD-----WSDY 237
+D AI G+QK LS+P+G + V+A+ K +E + ++ R S+Y
Sbjct: 189 IDAAIGGTQKCLSVPSGMAPITYNERVADVIAARKKVERGIATQADRAALSGNRPITSNY 248
Query: 238 LKFYKMGTYWP------YTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEA 291
++ YW +T + +LY LR + L+ EEGLE RH A ++A
Sbjct: 249 FDLSQLEDYWSERRLNHHTEATTMLYALREGVRLVLEEGLETRFERHRHHEAALAAGIKA 308
Query: 292 WGLKNCTQEEEWFSDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHL 351
GL+ ++ VT V +P IDG + ++ + + +AGK++RIG +
Sbjct: 309 MGLR-LFGDDSCKMPVVTCVEIPGGIDGESVRDMLLAQFGIEIASSFGPLAGKIWRIGTM 367
Query: 352 G 352
G
Sbjct: 368 G 368
>SPYA_HUMAN (P21549) Serine--pyruvate aminotransferase (EC 2.6.1.51)
(SPT) (Alanine--glyoxylate aminotransferase) (EC
2.6.1.44) (AGT)
Length = 392
Score = 130 bits (328), Expect = 4e-30
Identities = 105/350 (30%), Positives = 158/350 (45%), Gaps = 18/350 (5%)
Query: 12 LFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTG 71
L PGP N+P +++ A S + + + E ++ +F+T +I +G
Sbjct: 25 LLGPGPSNLPPRIMAAGGLQMIGSMSKDMYQIMDEIKEGIQYVFQTRNPLTLVISGSGHC 84
Query: 72 AWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDILESKLA 131
A E+AL N L PGD + G + VD +R+ V + + G L +E LA
Sbjct: 85 ALEAALVNVLEPGDSFLVGANGIWGQRAVDIGERIGARVHPMTKDPGGHYTLQEVEEGLA 144
Query: 132 SDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWG 191
H + + H E++TGV L +L Y+ LLLVD V+S+ MD G
Sbjct: 145 Q---HKPVLLFLTHGESSTGVLQPLDGFGELCHRYK--CLLLVDSVASLGGTPLYMDRQG 199
Query: 192 VDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMGTYWP--- 248
+D+ +GSQKAL+ P G + S KA + S K+ F YL + +W
Sbjct: 200 IDILYSGSQKALNAPPGTSLISFSDKAKKKMYSRKTKPFSF----YLDIKWLANFWGCDD 255
Query: 249 ------YTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQEEE 302
+T + LY LR +L LI E+GLEN +H ++A GL+ ++
Sbjct: 256 QPRMYHHTIPVISLYSLRESLALIAEQGLENSWRQHREAAAYLHGRLQALGLQLFVKDPA 315
Query: 303 WFSDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLG 352
TVT V VP D +IV +++ + GL GKV RIG LG
Sbjct: 316 LRLPTVTTVAVPAGYDWRDIVSYVIDHFDIEIMGGLGPSTGKVLRIGLLG 365
>SPYA_RAT (P09139) Serine--pyruvate aminotransferase, mitochondrial
precursor (EC 2.6.1.51) (SPT) (Alanine--glyoxylate
aminotransferase) (EC 2.6.1.44) (AGT)
Length = 414
Score = 125 bits (315), Expect = 1e-28
Identities = 101/350 (28%), Positives = 156/350 (43%), Gaps = 18/350 (5%)
Query: 12 LFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTG 71
L PGP N+ +V+ A S + + + + + ++ +F+T ++ +G
Sbjct: 47 LLGPGPSNLAPRVLAAGSLRMIGHMQKEMFQIMDEIKQGIQYVFQTRNPLTLVVSGSGHC 106
Query: 72 AWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDILESKLA 131
A E+AL N L PGD + G + + + +R+ V + + G L +E LA
Sbjct: 107 AMETALFNLLEPGDSFLVGTNGIWGIRAAEIAERIGARVHQMIKKPGEHYTLQEVEEGLA 166
Query: 132 SDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWG 191
H + + H E++TGV L +L YQ LLLVD V+S+ + MD+ G
Sbjct: 167 Q---HKPVLLFLTHGESSTGVLQPLDGFGELCHRYQ--CLLLVDSVASLGGVPIYMDQQG 221
Query: 192 VDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMGTYWP--- 248
+D+ +GSQK L+ P GI + + KA S K+ V F Y + W
Sbjct: 222 IDILYSGSQKVLNAPPGISLISFNDKAKSKVYSRKTKPVSF----YTDITYLSKLWGCEG 277
Query: 249 ------YTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQEEE 302
+T + LY LR +L LI E+GLEN RH + GLK ++ E
Sbjct: 278 KTRVIHHTLPVISLYCLRESLALISEQGLENSWRRHREATAHLHKCLRELGLKFFVKDPE 337
Query: 303 WFSDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLG 352
T+T V VP + +IV +N+ + GL KV RIG LG
Sbjct: 338 IRLPTITTVTVPAGYNWRDIVSYVLDHFNIEISGGLGPSEDKVLRIGLLG 387
>SPYA_FELCA (P41689) Serine--pyruvate aminotransferase,
mitochondrial precursor (EC 2.6.1.51) (SPT)
(Alanine--glyoxylate aminotransferase) (EC 2.6.1.44)
(AGT)
Length = 414
Score = 125 bits (313), Expect = 2e-28
Identities = 102/350 (29%), Positives = 155/350 (44%), Gaps = 18/350 (5%)
Query: 12 LFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTG 71
L PGP N+ +V+ A + + + + + + ++ +F+T I +G
Sbjct: 47 LLGPGPSNLAPRVLVAGGKQMIGHMHKEMFQIMDDIKQGIQYVFQTKNPLTLAISGSGHC 106
Query: 72 AWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDILESKLA 131
A E+AL N L PGD + V G + D +R+ V + + G L LE LA
Sbjct: 107 ALEAALFNILEPGDPFLVGVNGIWGQRAADIGERIGARVHPMIKDPGNHYTLQELEEALA 166
Query: 132 SDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWG 191
H + + E+++GV L +L Y LLLVD V+S+C MD+ G
Sbjct: 167 Q---HKPVLLFLTQGESSSGVLQPLDGYGELCHRYN--CLLLVDSVASLCGTPIYMDQQG 221
Query: 192 VDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMGTYWP--- 248
+DV +GSQK L+ P G + S KA + K+ V F YL + W
Sbjct: 222 IDVLYSGSQKVLNSPPGTSLISFSDKAKNKIYTRKTKPVSF----YLDMKWLANIWGCDG 277
Query: 249 ------YTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQEEE 302
+T + LY LR +L LI E+GLEN +H + ++ GL+ ++
Sbjct: 278 KPRIYHHTTPVVSLYSLRESLALIAEQGLENSWRQHREVTAYLHGRLQGLGLQLFVKDPA 337
Query: 303 WFSDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLG 352
TVT V VP D +IV +++ + GL GKV RIG LG
Sbjct: 338 LRLPTVTTVAVPAGYDWRDIVNYVMDHFDIEITGGLGPSMGKVLRIGLLG 387
>SPYA_CALJA (P31029) Serine--pyruvate aminotransferase,
mitochondrial precursor (EC 2.6.1.51) (SPT)
(Alanine--glyoxylate aminotransferase) (EC 2.6.1.44)
(AGT)
Length = 414
Score = 123 bits (309), Expect = 7e-28
Identities = 101/350 (28%), Positives = 153/350 (42%), Gaps = 18/350 (5%)
Query: 12 LFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTG 71
L PGP N+P + + A + + + E ++ +F+T +I +G
Sbjct: 47 LLGPGPSNLPPRTMAAGGLQMLGHMHKETYQIMDEIKEGIQYVFQTRNPLTLVISGSGHC 106
Query: 72 AWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDILESKLA 131
A E+AL N L PGD + V G + D +RL V + + G L +E LA
Sbjct: 107 ALEAALINVLEPGDSFLVGVNGIWGQRAADIGERLGARVHPMTKDPGGHYTLQEVEEGLA 166
Query: 132 SDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWG 191
H + + H E+++GV L +L Y+ LLLVD V+S+ MD+ G
Sbjct: 167 Q---HKPVLLFLTHGESSSGVLQPLDGFGELCHRYK--CLLLVDSVASLGGAPLYMDQQG 221
Query: 192 VDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMGTYWP--- 248
+D+ +GSQK L+ P G + S A S K+ F YL + W
Sbjct: 222 IDILYSGSQKVLNAPPGTSLLSFSDTAKNKIYSRKTKPSSF----YLDVKYLANLWGCDG 277
Query: 249 ------YTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQEEE 302
+T + LY LR L L+ E+GLEN +H ++A GL+ ++
Sbjct: 278 QPRMYHHTTPVVSLYSLREGLALLSEQGLENSWRKHREAAAYLHGRLQALGLRLFVKDPA 337
Query: 303 WFSDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLG 352
TVT V VP D +IV +R+ + + GL GKV RIG +G
Sbjct: 338 LRLPTVTTVAVPTGYDWRDIVSYLIERFGIEITGGLGPSTGKVLRIGLMG 387
>SPYA_MOUSE (O35423) Serine--pyruvate aminotransferase,
mitochondrial precursor (EC 2.6.1.51) (SPT)
(Alanine--glyoxylate aminotransferase) (EC 2.6.1.44)
(AGT)
Length = 413
Score = 119 bits (299), Expect = 1e-26
Identities = 98/350 (28%), Positives = 156/350 (44%), Gaps = 19/350 (5%)
Query: 12 LFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTG 71
L PGP N+ +V+ A S + + + + + + ++ +F+T ++ +G
Sbjct: 47 LLGPGPSNLAPRVLAAGSLRMIGHMQKEMLQIMEEIKQGIQYVFQTRNPLTLVVSGSGHC 106
Query: 72 AWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDILESKLA 131
A E+AL N L PGD ++ G + + + R+ V + + G L +E LA
Sbjct: 107 AMETALFNLLEPGDSFLTGTNGIWGMRAAEIADRIGARVHQMIKKPGEHYTLQEVEEGLA 166
Query: 132 SDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWG 191
H + +VH E++TGV L +L YQ LLLVD V+S+ + MD+ G
Sbjct: 167 Q---HKPVLLFLVHGESSTGVVQPLDGFGELCHRYQ--CLLLVDSVASLGGVPIYMDQQG 221
Query: 192 VDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMGTYW---- 247
+D+ + SQK L+ P GI + + KA S K+ V F Y + W
Sbjct: 222 IDIMYSSSQKVLNAPPGISLISFNDKAKYKVYSRKTKPVSF----YTDITYLAKLWGCEG 277
Query: 248 -----PYTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQEEE 302
+T + LY LR +L LI E GLEN RH ++ GLK ++ E
Sbjct: 278 ETRVIHHTTPVTSLYCLRESLALIAERGLENCWRRHREATAHLHKHLQEMGLKFFVKDPE 337
Query: 303 WFSDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLG 352
T+T V Y + +IV +++ + GL +V RIG LG
Sbjct: 338 IRLPTITTVTAAGY-NWRDIVSYVLDHFSIEISGGLGPTEERVLRIGLLG 386
>AGX1_YEAST (P43567) Alanine--glyoxylate aminotransferase 1 (EC
2.6.1.44)
Length = 385
Score = 116 bits (290), Expect = 1e-25
Identities = 96/358 (26%), Positives = 169/358 (46%), Gaps = 23/358 (6%)
Query: 12 LFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGT---PFLIPTT 68
L +PGP+ + V +A+ + + SP ++ + +L++ + +FK+ + PF++ +
Sbjct: 8 LLIPGPIILSGAVQKALDVPSLGHTSPEFVSIFQRVLKNTRAVFKSAAASKSQPFVLAGS 67
Query: 69 GTGAWESALTNTL---SPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVES-EWGRGADLD 124
GT W+ +N + +P ++ G FS + D + VDVV + G L+
Sbjct: 68 GTLGWDIFASNFILSKAPNKNVLVVSTGTFSDRFADCLRSYGAQVDVVRPLKIGESVPLE 127
Query: 125 ILESKLASDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALD 184
++ KL+ +S A+ + H +T+T V ++L + Q + +VD V SI +
Sbjct: 128 LITEKLSQNS---YGAVTVTHVDTSTAVLSDLKAISQAIKQTSPETFFVVDAVCSIGCEE 184
Query: 185 FRMDEWGVDVAITGSQKALSLPTGIGFVVASPK----AIEASKSAKSLRVFFD---WSDY 237
F DEWGVD A+T SQKA+ P G+ + S + A+ SK+ F W+
Sbjct: 185 FEFDEWGVDFALTASQKAIGAPAGLSISLCSSRFMDYALNDSKNGHVHGYFSSLRRWTPI 244
Query: 238 LKFYK--MGTYWPYTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLK 295
++ Y+ G Y+ TP +QL+ L AL I EEGL H + + ++ GL+
Sbjct: 245 MENYEAGKGAYFA-TPPVQLINSLDVALKEILEEGLHKRWDLHREMSDWFKDSL-VNGLQ 302
Query: 296 NCTQEEEWFSDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAG-KVFRIGHLG 352
T + S+ + Y+ V K + + + G++K G K RIGH+G
Sbjct: 303 -LTSVSRYPSNMSAHGLTAVYVADPPDVIAFLKSHGVVIAGGIHKDIGPKYIRIGHMG 359
>SPYA_RABIT (P31030) Serine--pyruvate aminotransferase (EC 2.6.1.51)
(SPT) (Alanine--glyoxylate aminotransferase) (EC
2.6.1.44) (AGT)
Length = 392
Score = 115 bits (288), Expect = 2e-25
Identities = 97/350 (27%), Positives = 153/350 (43%), Gaps = 18/350 (5%)
Query: 12 LFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTG 71
L PGP N+P +V+ A + + + + + ++ F+T + +G
Sbjct: 25 LLGPGPSNLPPRVLAAGGLQMIGHMHEEMYQVMDEIKQGIQYAFQTRNALTLAVSGSGHC 84
Query: 72 AWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDILESKLA 131
A E+AL N L PGD + G + + +R+ V + + G L +E LA
Sbjct: 85 ALETALFNLLEPGDAFLVGANGIWGQRAAEVGERIGARVHPMIKDPGSHYTLQEVEECLA 144
Query: 132 SDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWG 191
H + + H E++TGV L +L Y+ LLLVD V+S+ MD+ G
Sbjct: 145 Q---HKPVLLFLTHGESSTGVLQPLDGFGELCHRYK--CLLLVDSVASLGGAPIYMDQQG 199
Query: 192 VDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMGTYWP--- 248
+DV +GSQKAL+ P G + S KA KS R +S Y+ + W
Sbjct: 200 IDVLYSGSQKALNAPPGTSLISFSDKA----KSKIYARKTKPFSFYMDVQLLANIWGCDG 255
Query: 249 ------YTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQEEE 302
+T + ++ LR +L L+ E+GLE RH + ++ GL+ ++
Sbjct: 256 KPRMYHHTTPVIGIFALRESLALLVEQGLEKSWQRHREVAQHLYRRLQELGLQLFVKDPA 315
Query: 303 WFSDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLG 352
TVT V+VP +IV + + + GL A KV RIG LG
Sbjct: 316 LRLPTVTTVIVPASYRWRDIVSYVMHHFGIEITGGLGPSADKVLRIGLLG 365
>SGAA_MAIZE (P84187) Serine--glyoxylate aminotransferase (EC
2.6.1.45) (SGAT) (Alanine--glyoxylate aminotransferase)
(EC 2.6.1.44) (AGT) (Fragments)
Length = 136
Score = 93.6 bits (231), Expect = 7e-19
Identities = 67/162 (41%), Positives = 72/162 (44%), Gaps = 80/162 (49%)
Query: 187 MDEWGVDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMGTY 246
MDEWGVDVA+TGSQKALS PTG+G V ASP RVFFDW DYL+ TY
Sbjct: 47 MDEWGVDVALTGSQKALSFPTGMGLVCASP------------RVFFDWKDYLR-----TY 89
Query: 247 WPYTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQEEEWFSD 306
W Y ++ L LAVEAWGL N
Sbjct: 90 WHYDQALDL------------------------------ELAVEAWGLSN---------- 109
Query: 307 TVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVA-GKVFR 347
RYNLSLGLGLNKVA GKVFR
Sbjct: 110 ----------------------RYNLSLGLGLNKVAGGKVFR 129
Score = 48.1 bits (113), Expect = 4e-05
Identities = 36/88 (40%), Positives = 41/88 (45%), Gaps = 45/88 (51%)
Query: 1 MDYVNGPGRNHLFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTG 60
+DYV GPGR RAM+ SPA+PALTK LLEDVKK
Sbjct: 1 LDYVYGPGR----------------RAMN-------SPAVPALTKVLLEDVKK------- 30
Query: 61 TPFLIPTTGTGAWESALTNTLSPGDRIV 88
ALTNTLSPGDR++
Sbjct: 31 ---------------ALTNTLSPGDRLL 43
>Y736_HELPJ (Q9ZLA7) Putative aminotransferase JHP0673 (EC 2.6.1.-)
Length = 369
Score = 79.3 bits (194), Expect = 1e-14
Identities = 82/343 (23%), Positives = 153/343 (43%), Gaps = 19/343 (5%)
Query: 12 LFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTG 71
LF PGPV I +++ + S+ +R+ + +++ E++KK+ T L+ ++GTG
Sbjct: 3 LFTPGPVAINEEMRSSFSQPMPHHRTKDFEKIFQSVRENLKKM--TGLEEVLLLSSSGTG 60
Query: 72 AWESALTNTLSPGDRIVSFV-IGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDILESKL 130
A E+++ +S + + FV G+F + + + EW A +D + + L
Sbjct: 61 AMEASV---ISLCQKELLFVNAGKFGERFGKIAKAHSIKAHELVYEWDTPAQVDGILNAL 117
Query: 131 ASDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEW 190
++ I A CI E++ G+ + + K+ Q + ++VD ++++ +
Sbjct: 118 KANP--NIDAFCIQACESSGGLRHPVEKIAQAIKETNPNVFVIVDAITALGVEPLEITH- 174
Query: 191 GVDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMGTYWPYT 250
VD I GSQKA LP + + S KAI+ + +++ +F+ LK K T YT
Sbjct: 175 -VDALIGGSQKAFMLPPAMSLIALSQKAIDRIEE-RNVGFYFNLKSELKNQKNNT-TSYT 231
Query: 251 PSIQLLYGLRAALDLIFE-EGLENIIARHNRLGTATRLAVEAWGLKNCTQEEEWFSDTVT 309
I GL+ +L+ G E + R+ A++ AV A GLK + T+
Sbjct: 232 APILHTLGLQRYFELVQNLGGFEALYQETKRVALASQKAVLALGLKIFPKSPSLSMTTIV 291
Query: 310 AVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLG 352
+ E+ ++Y + G + RI H+G
Sbjct: 292 SE------HAKELKNLLKEKYQVQFAGGQEPYKDTLIRINHMG 328
>Y736_HELPY (O25436) Putative aminotransferase HP0736 (EC 2.6.1.-)
Length = 369
Score = 77.8 bits (190), Expect = 4e-14
Identities = 87/344 (25%), Positives = 157/344 (45%), Gaps = 21/344 (6%)
Query: 12 LFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTGTPFLIPTTGTG 71
LF PGPV I +++ + S+ +R+ + +++ E++KK+ T L+ ++GTG
Sbjct: 3 LFTPGPVAINEEMRTSFSQPMPHHRTKDFEKIFQSVRENLKKM--TGLEEVLLLSSSGTG 60
Query: 72 AWESALTNTLSPGDRIVSFV-IGQFSLLWVDQQQRLKFNVDVVESEWGRGADLDILESKL 130
A E+++ +S + + FV G+F + + + EW A +D + S L
Sbjct: 61 AMEASV---ISLCQKELLFVNAGKFGERFGKIAKAHSIKAHELVYEWDTPAQVDEILSVL 117
Query: 131 ASDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEW 190
++ I A CI E++ G+ + + K+ Q + ++VD ++++ +
Sbjct: 118 KANP--NIDAFCIQACESSGGLRHPVEKIAQAIKETNPNVFVIVDAITALGVEPLEITH- 174
Query: 191 GVDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMGTYWPYT 250
VD I GSQKA LP + V S AIE + +++ +F+ LK + T YT
Sbjct: 175 -VDALIGGSQKAFMLPPAMSLVALSQNAIERIEE-RNVGFYFNLKSELKNQRNNT-TSYT 231
Query: 251 PSIQLLYGLRAALDLIFE-EGLENIIARHNRLGTATRLAVEAWGLKNCTQEEEWFSDTVT 309
I GL+ +L+ G E + + AT+ AV A GLK + S ++T
Sbjct: 232 APILHTLGLQRYFELVQNLGGFEALYRETKKAALATQKAVLALGLKIFPKSP---SLSMT 288
Query: 310 AVVVPPYIDGAEIVRRAWK-RYNLSLGLGLNKVAGKVFRIGHLG 352
+V + A+ +R K +Y + G + RI H+G
Sbjct: 289 TIV----NEHAKELRNLLKEKYQVQFAGGQEPYKDALIRINHMG 328
>CSD_SYNY3 (Q55793) Probable cysteine desulfurase (EC 2.8.1.7)
Length = 420
Score = 47.4 bits (111), Expect = 6e-05
Identities = 55/207 (26%), Positives = 89/207 (42%), Gaps = 24/207 (11%)
Query: 79 NTLSPGDRIVSFVIGQFSLL--WVDQQQRLKFNVDVVESEWGRGADLDILESKLASDSAH 136
N L GD I++ V+ S L W + + V+ + DL+ ++ L+ +
Sbjct: 113 NNLKAGDEIITTVMEHHSNLVPWQMVAAKTGAVLKFVQLDEQESFDLEHFKTLLSEKT-- 170
Query: 137 TIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWGVDVA- 195
K + +VH G N ++ QL A+Q A +LVD S A + +D +D
Sbjct: 171 --KLVTVVHISNTLGCVNPAEEIAQL--AHQAGAKVLVDACQS--APHYPLDVQLIDCDW 224
Query: 196 ITGSQKALSLPTGIGFVVASPKAIEASK-----SAKSLRVFFDW---SDYLKFYKMGTYW 247
+ S + PTGIGF+ + +EA VFFD + ++ GT
Sbjct: 225 LVASGHKMCAPTGIGFLYGKEEILEAMPPFFGGGEMIAEVFFDHFTTGELPHKFEAGT-- 282
Query: 248 PYTPSIQLLYGLRAALDLIFEEGLENI 274
P+I L AA+D + + G+ENI
Sbjct: 283 ---PAIAEAIALGAAVDYLTDLGMENI 306
>CSD_BACHD (Q9K7A0) Probable cysteine desulfurase (EC 2.8.1.7)
Length = 406
Score = 46.6 bits (109), Expect = 1e-04
Identities = 37/146 (25%), Positives = 63/146 (42%), Gaps = 6/146 (4%)
Query: 139 KAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWGVDVAITG 198
K + I+H G N + ++ ++ A++H A++LVDG S + + E G D
Sbjct: 164 KIVAIMHVSNVLGTINPVKEIAEI--AHRHGAIMLVDGAQSAPHMKIDVQELGCDFFAFS 221
Query: 199 SQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMGTYWPY---TPSIQL 255
K ++ PTGIG + +E + + D+ W + TP I
Sbjct: 222 GHK-MAGPTGIGVLYGKKAHLEKMEPVEFGGEMIDFVGLYDSTWKELPWKFEGGTPIIAG 280
Query: 256 LYGLRAALDLIFEEGLENIIARHNRL 281
GL AA+D + + GL+ I + L
Sbjct: 281 AIGLGAAIDFLEDIGLDEIEKHEHEL 306
>CSD_CHLTR (O84693) Probable cysteine desulfurase (EC 2.8.1.7)
Length = 401
Score = 43.5 bits (101), Expect = 9e-04
Identities = 32/132 (24%), Positives = 57/132 (42%), Gaps = 6/132 (4%)
Query: 138 IKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWGVDVAIT 197
++ + + H +G L ++ L+ Y+ AL VDG + + EWGVD
Sbjct: 158 VQLVSLAHVSNVSGAVLPLPEIAHLVHRYE--ALFAVDGAQGVGKGPLNLSEWGVDFYAF 215
Query: 198 GSQKALSLPTGIGFVVASPKAIEA---SKSAKSLRVFFDWSDYLKFYKMGTYWPYTPSIQ 254
K L PTGIG + + +E+ + + + +D+ + + TP I
Sbjct: 216 SGHK-LYAPTGIGVLYGKKELLESLPPVEGGGDMVIVYDFEELSYQEPPLRFEAGTPHIA 274
Query: 255 LLYGLRAALDLI 266
+ GL AA+D +
Sbjct: 275 GVLGLGAAIDYL 286
>CSD_BACSU (O32164) Probable cysteine desulfurase (EC 2.8.1.7)
Length = 406
Score = 42.0 bits (97), Expect = 0.003
Identities = 34/146 (23%), Positives = 62/146 (42%), Gaps = 6/146 (4%)
Query: 139 KAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSICALDFRMDEWGVDVAITG 198
K + + H G N + ++ ++ A+ + A+++VDG S + + + D
Sbjct: 164 KIVAVSHVSNVLGTVNPIKEMAKI--AHDNGAVIVVDGAQSTPHMKIDVQDLDCDFFALS 221
Query: 199 SQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKFYKMGTYWPY---TPSIQL 255
S K PTG+G + +E + A+ D+ + W + TP I
Sbjct: 222 SHKMCG-PTGVGVLYGKKALLENMEPAEFGGEMIDFVGLYESTWKELPWKFEAGTPIIAG 280
Query: 256 LYGLRAALDLIFEEGLENIIARHNRL 281
GL AA+D + E GL+ I ++L
Sbjct: 281 AIGLGAAIDFLEEIGLDEISRHEHKL 306
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 44,625,739
Number of Sequences: 164201
Number of extensions: 1824509
Number of successful extensions: 4178
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 44
Number of HSP's that attempted gapping in prelim test: 4103
Number of HSP's gapped (non-prelim): 68
length of query: 379
length of database: 59,974,054
effective HSP length: 112
effective length of query: 267
effective length of database: 41,583,542
effective search space: 11102805714
effective search space used: 11102805714
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC141111.6


