

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141111.3 + phase: 0 /pseudo
(64 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
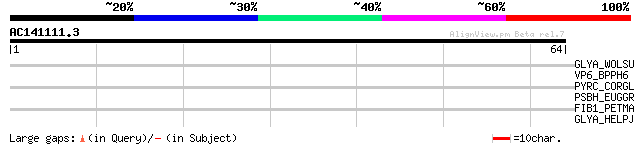
Score E
Sequences producing significant alignments: (bits) Value
GLYA_WOLSU (Q7MAR0) Serine hydroxymethyltransferase (EC 2.1.2.1)... 32 0.31
VP6_BPPH6 (P11128) P6 protein 28 5.8
PYRC_CORGL (Q8NQ39) Dihydroorotase (EC 3.5.2.3) (DHOase) 28 5.8
PSBH_EUGGR (P31555) Photosystem II reaction center H protein (Ph... 28 5.8
FIB1_PETMA (P02674) Fibrinogen alpha-1 chain precursor [Contains... 27 7.6
GLYA_HELPJ (Q9ZMP7) Serine hydroxymethyltransferase (EC 2.1.2.1)... 27 9.9
>GLYA_WOLSU (Q7MAR0) Serine hydroxymethyltransferase (EC 2.1.2.1)
(Serine methylase) (SHMT)
Length = 416
Score = 32.0 bits (71), Expect = 0.31
Identities = 18/55 (32%), Positives = 27/55 (48%), Gaps = 2/55 (3%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAG--NYEK 55
+ VYN ++ Y K MD + G LT G ++ +Y+S G G NY+K
Sbjct: 100 VAVYNALLKPYDKILGMDLSHGGHLTHGAKVSVTGQTYQSFFYGVELDGYINYDK 154
>VP6_BPPH6 (P11128) P6 protein
Length = 167
Score = 27.7 bits (60), Expect = 5.8
Identities = 13/58 (22%), Positives = 25/58 (42%), Gaps = 2/58 (3%)
Query: 1 MCIVVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYEKARW 58
+ I+ + + G+ ++ G G+F ++ I P TS+ S W G+ W
Sbjct: 35 VAIIYFAPYLAGFFTSAGFTGIGGIFSSIATTITPTLTSFLS--TAWSGVGSLASTAW 90
>PYRC_CORGL (Q8NQ39) Dihydroorotase (EC 3.5.2.3) (DHOase)
Length = 447
Score = 27.7 bits (60), Expect = 5.8
Identities = 11/36 (30%), Positives = 21/36 (57%)
Query: 3 IVVYNTMITGYGKASNMDGAEGVFLTLGGRIEPDET 38
+++ N ++ G G+ +N+ +GV +GG E D T
Sbjct: 22 LLISNVLVYGEGEPTNVFVKDGVIAAIGGTHEADRT 57
>PSBH_EUGGR (P31555) Photosystem II reaction center H protein
(Photosystem II 10 kDa phosphoprotein) (PSII-H)
Length = 70
Score = 27.7 bits (60), Expect = 5.8
Identities = 14/37 (37%), Positives = 21/37 (55%), Gaps = 3/37 (8%)
Query: 15 KASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAG 51
K SN ++G TLG ++P + Y ++ GWG AG
Sbjct: 8 KTSN---SKGKTTTLGTILKPLNSKYGKVLPGWGTAG 41
>FIB1_PETMA (P02674) Fibrinogen alpha-1 chain precursor [Contains:
Fibrinopeptide A] (Fragment)
Length = 966
Score = 27.3 bits (59), Expect = 7.6
Identities = 16/44 (36%), Positives = 21/44 (47%), Gaps = 2/44 (4%)
Query: 11 TGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAGNYE 54
TG A+N A+G + GGR EP+ S + WG G E
Sbjct: 402 TGGSTATNTGSAQGGSWSTGGRTEPNTGSGQG--GSWGTGGRTE 443
>GLYA_HELPJ (Q9ZMP7) Serine hydroxymethyltransferase (EC 2.1.2.1)
(Serine methylase) (SHMT)
Length = 416
Score = 26.9 bits (58), Expect = 9.9
Identities = 18/54 (33%), Positives = 27/54 (49%), Gaps = 2/54 (3%)
Query: 5 VYNTMITGYGKASNMDGAEGVFLTLGGRIEPDETSYRSMIEGWGRAG--NYEKA 56
VY+ ++ Y K MD + G LT G ++ Y+S G G G +YE+A
Sbjct: 102 VYHALLKLYDKILGMDLSCGGHLTHGAKVSLTGKHYQSFSYGVGLDGYIDYEEA 155
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.137 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,983,857
Number of Sequences: 164201
Number of extensions: 252649
Number of successful extensions: 597
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 3
Number of HSP's that attempted gapping in prelim test: 591
Number of HSP's gapped (non-prelim): 9
length of query: 64
length of database: 59,974,054
effective HSP length: 40
effective length of query: 24
effective length of database: 53,406,014
effective search space: 1281744336
effective search space used: 1281744336
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC141111.3


