

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140916.4 - phase: 0
(250 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
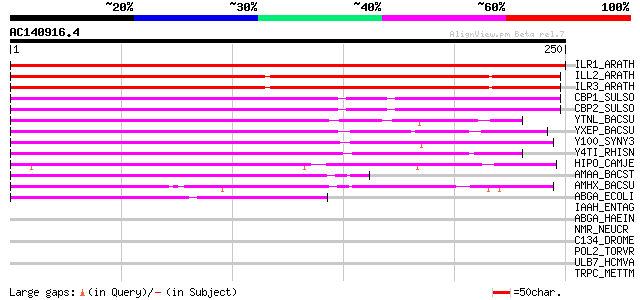
Score E
Sequences producing significant alignments: (bits) Value
ILR1_ARATH (P54968) IAA-amino acid hydrolase 1 (EC 3.5.1.-) 243 2e-64
ILL2_ARATH (P54970) IAA-amino acid hydrolase homolog 2 precursor 237 2e-62
ILR3_ARATH (P54969) IAA-amino acid hydrolase 3 precursor (EC 3.5... 236 4e-62
CBP1_SULSO (P80092) Thermostable carboxypeptidase 1 (EC 3.4.17.-) 152 7e-37
CBP2_SULSO (P58156) Thermostable carboxypeptidase 2 (EC 3.4.17.-) 151 1e-36
YTNL_BACSU (O34980) Hypothetical protein ytnL 135 7e-32
YXEP_BACSU (P54955) Hypothetical protein yxeP 126 4e-29
Y100_SYNY3 (P54984) Hypothetical protein sll0100 115 8e-26
Y4TI_RHISN (P55663) Hypothetical hydrolase Y4TI (EC 3.-.-.-) 100 6e-21
HIPO_CAMJE (P45493) Hippurate hydrolase (EC 3.5.1.32) (Benzoylgl... 97 5e-20
AMAA_BACST (P37112) N-acyl-L-amino acid amidohydrolase (EC 3.5.1... 78 2e-14
AMHX_BACSU (P54983) Amidohydrolase amhX (EC 3.5.1.-) (Aminoacylase) 58 2e-08
ABGA_ECOLI (P77357) Aminobenzoyl-glutamate utilization protein A 44 4e-04
IAAH_ENTAG (O50173) Indole-3-acetyl-aspartic acid hydrolase (EC ... 42 0.001
ABGA_HAEIN (P44765) Aminobenzoyl-glutamate utilization protein A... 37 0.045
NMR_NEUCR (P23762) Nitrogen metabolic regulation protein (NMR pr... 35 0.22
C134_DROME (Q9VG40) Probable cytochrome P450 313a4 (EC 1.14.-.-)... 32 1.4
POL2_TORVR (P25247) RNA2 polyprotein (P2) [Contains: X3 protein;... 31 2.5
ULB7_HCMVA (P16770) Hypothetical protein UL117 31 3.2
TRPC_METTM (P26939) Indole-3-glycerol phosphate synthase (EC 4.1... 30 4.2
>ILR1_ARATH (P54968) IAA-amino acid hydrolase 1 (EC 3.5.1.-)
Length = 442
Score = 243 bits (621), Expect = 2e-64
Identities = 120/250 (48%), Positives = 174/250 (69%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M++D L+D++ I +VHV P+G IGSRPG +LAG G F + G+ + AA P S D
Sbjct: 182 MLKDEILDDLDGILSVHVFPSIPSGGIGSRPGTVLAGAGLFTVTVHGQGSHAATPHFSKD 241
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PVLAAS+AV+++Q IVSRE +PL++ VV+V GG++ ++IP S GGTFR+ SN
Sbjct: 242 PVLAASSAVVALQQIVSRELDPLEAGVVTVGYIEGGHAQNVIPQSAKFGGTFRSLSNDGL 301
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
+ RI+++ QASVY C AEV+F EK+ +++P ND+ +YEH KKV+ ++G+ NF
Sbjct: 302 LFIQRRIKEISEAQASVYRCKAEVNFEEKKPSLHPVMNNDEGLYEHGKKVAEAMIGKNNF 361
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
P MG ED+SF++Q +A F +G++NETLG+ HSP+F +DE+ALP+GAA+HA
Sbjct: 362 HDFPVTMGGEDFSFFTQKTKAAIFVLGVKNETLGAGKPLHSPYFFVDEEALPVGAALHAA 421
Query: 241 IAERYLNEHG 250
+A YL+EHG
Sbjct: 422 MAVSYLDEHG 431
>ILL2_ARATH (P54970) IAA-amino acid hydrolase homolog 2 precursor
Length = 439
Score = 237 bits (604), Expect = 2e-62
Identities = 122/248 (49%), Positives = 176/248 (70%), Gaps = 3/248 (1%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M ++GAL++VEAIF +H+S P G SR G LAG G F AVI+GK AA P+++ D
Sbjct: 181 MREEGALKNVEAIFGIHLSARIPFGKAASRAGSFLAGAGVFEAVITGKGGHAAIPQHTID 240
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PV+AAS+ V+S+Q +VSRE++PLDS+VV+V+ NGGN+ ++IPDS+ IGGT RAF T F
Sbjct: 241 PVVAASSIVLSLQQLVSRETDPLDSKVVTVSKVNGGNAFNVIPDSITIGGTLRAF--TGF 298
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL +R+++VI +QA+V+ C A V+ PPTVN+ +Y+ KKV DLLGQ+ F
Sbjct: 299 TQLQQRVKEVITKQAAVHRCNASVNLTPNGREPMPPTVNNKDLYKQFKKVVRDLLGQEAF 358
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
P+MG+ED+S++++ IP F +G+++ET G + HSP + I+ED LP GAA+HA+
Sbjct: 359 VEAAPVMGSEDFSYFAETIPGHFSLLGMQDETNGYA-SSHSPLYRINEDVLPYGAAIHAS 417
Query: 241 IAERYLNE 248
+A +YL E
Sbjct: 418 MAVQYLKE 425
>ILR3_ARATH (P54969) IAA-amino acid hydrolase 3 precursor (EC
3.5.1.-)
Length = 438
Score = 236 bits (602), Expect = 4e-62
Identities = 122/248 (49%), Positives = 174/248 (69%), Gaps = 3/248 (1%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M ++GAL++VEAIF +H+S P G S G +AG G F AVI+GK AA P+++ D
Sbjct: 180 MREEGALKNVEAIFGIHLSPRTPFGKAASLAGSFMAGAGAFEAVITGKGGHAAIPQHTID 239
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
PV+AAS+ V+S+Q +VSRE++P DS+VV+VT NGGN+ ++IPDS+ IGGT RAF T F
Sbjct: 240 PVVAASSIVLSLQHLVSRETDPSDSKVVTVTKVNGGNAFNVIPDSITIGGTLRAF--TGF 297
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL ERI+++I +QA+V+ C A V+ PPTVN+ +Y+ KKV DLLGQ+ F
Sbjct: 298 TQLQERIKEIITKQAAVHRCNASVNLAPNGNQPMPPTVNNMDLYKKFKKVVRDLLGQEAF 357
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
P MG+ED+S++++ IP F +G+++ET G + HSPH+ I+ED LP GAA+HAT
Sbjct: 358 VEAVPEMGSEDFSYFAETIPGHFSLLGMQDETQGYA-SSHSPHYRINEDVLPYGAAIHAT 416
Query: 241 IAERYLNE 248
+A +YL +
Sbjct: 417 MAVQYLKD 424
>CBP1_SULSO (P80092) Thermostable carboxypeptidase 1 (EC 3.4.17.-)
Length = 393
Score = 152 bits (384), Expect = 7e-37
Identities = 73/248 (29%), Positives = 133/248 (53%), Gaps = 6/248 (2%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MI+ G + V+ +F +H+S +P+G+ +R GP++A F+ ++ GK + P + D
Sbjct: 152 MIEAGVMNGVDYVFGIHISSSYPSGVFATRKGPIMATPDAFKIIVHGKGGHGSAPHETID 211
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+ + +I GI +R+ +P+ ++S+T+ + G ++IPD + GT R+
Sbjct: 212 PIFISLQIANAIYGITARQIDPVQPFIISITTIHSGTKDNIIPDDAEMQGTIRSLDENVR 271
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
+ + + +++ +Y EV F E +YP TVN+ ++ + V K+ L
Sbjct: 272 SKAKDYMRRIVSSICGIYGATCEVKFME---DVYPTTVNNPEVTDEVMKI---LSSISTV 325
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
P++GAED+S + Q P +F++G RNE G + HS F +DED L +GA HA
Sbjct: 326 VETEPVLGAEDFSRFLQKAPGTYFFLGTRNEKKGCIYPNHSSKFCVDEDVLKLGALAHAL 385
Query: 241 IAERYLNE 248
+A ++ N+
Sbjct: 386 LAVKFSNK 393
>CBP2_SULSO (P58156) Thermostable carboxypeptidase 2 (EC 3.4.17.-)
Length = 393
Score = 151 bits (382), Expect = 1e-36
Identities = 75/248 (30%), Positives = 133/248 (53%), Gaps = 6/248 (2%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MI+ G + V+ +F +H+S +P+G+ +R GP++A F+ V+ GK + P + D
Sbjct: 152 MIEAGVMNGVDYVFGIHISSSYPSGVFATRKGPIMATPDAFKIVVHGKGGHGSAPHETID 211
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+ + +I GI +R+ +P+ V+S+T+ + G ++IPD + GT R+
Sbjct: 212 PIFISLQIANAIYGITARQIDPVQPFVISITTIHSGTKDNIIPDDAEMQGTIRSLDENVR 271
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
+ + + +++ +Y EV F E +YP TVN+ ++ + V K+ L
Sbjct: 272 SKAKDYMRRIVSSICGIYGATCEVKFME---DVYPITVNNPEVTDEVMKI---LSSISTV 325
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
P++GAED+S + Q P +F++G RNE G + HS F +DED L +GA HA
Sbjct: 326 VETEPVLGAEDFSRFLQKAPGMYFFLGTRNEKKGCIYPNHSSKFCVDEDVLKLGALAHAL 385
Query: 241 IAERYLNE 248
+A ++ N+
Sbjct: 386 LAIKFSNK 393
>YTNL_BACSU (O34980) Hypothetical protein ytnL
Length = 416
Score = 135 bits (341), Expect = 7e-32
Identities = 80/233 (34%), Positives = 128/233 (54%), Gaps = 15/233 (6%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
+I+DG L+ ++A+ +H + G +G + GPL+A F+ I GK A AA P N D
Sbjct: 172 VIEDGQLDGIDAVIGLHNKPDIAVGTVGLKTGPLMAAVDRFKVEIEGKGAHAALPHNGFD 231
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P++ AS ++++Q IVSR NPL S +++V NGG++ ++IPD+VVI GT R F +
Sbjct: 232 PIIGASQLIVALQTIVSRNVNPLQSAILTVGKINGGSTWNVIPDTVVIEGTVRTFDSEVR 291
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
Q+ +R V Q ++ +S A V K ++ PP ND+ + V+ D +
Sbjct: 292 NQVKQRFFAVTEQISAAFSLKANV----KWHSGPPPLCNDEAITGLVR----DAAHKAKL 343
Query: 181 RVV--PPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDAL 231
+V+ P ED+++Y + IP +F + G + H H P FTIDE A+
Sbjct: 344 QVIDPAPSTAGEDFAYYLEHIPGSFAFFGTDGD-----HDWHHPAFTIDETAI 391
>YXEP_BACSU (P54955) Hypothetical protein yxeP
Length = 380
Score = 126 bits (317), Expect = 4e-29
Identities = 78/242 (32%), Positives = 118/242 (48%), Gaps = 11/242 (4%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
+++ G L V AIF +H + P G IG + GPL+A F VI GK A+ P NS D
Sbjct: 142 VLEAGVLNGVSAIFGMHNKPDLPVGTIGVKEGPLMASVDRFEIVIKGKGGHASIPNNSID 201
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
P+ AA + +Q +VSR + L + VVS+T G S ++IPD + GT R F +
Sbjct: 202 PIAAAGQIISGLQSVVSRNISSLQNAVVSITRVQAGTSWNVIPDQAEMEGTVRTFQKEAR 261
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
+ E + +V A+ Y AE +F Y P+V +D + + + LG +
Sbjct: 262 QAVPEHMRRVAEGIAAGYGAQAEFKWFP-----YLPSVQNDGTFLNAASEAAARLGYQTV 316
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHAT 240
G ED++ Y + IP F ++G T H P FT+DE+AL + + A
Sbjct: 317 H-AEQSPGGEDFALYQEKIPGFFVWMG-----TNGTEEWHHPAFTLDEEALTVASQYFAE 370
Query: 241 IA 242
+A
Sbjct: 371 LA 372
>Y100_SYNY3 (P54984) Hypothetical protein sll0100
Length = 393
Score = 115 bits (289), Expect = 8e-26
Identities = 72/246 (29%), Positives = 114/246 (46%), Gaps = 5/246 (2%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MIQDGA++ V I VHV P +G R G L A I G+ A P + D
Sbjct: 146 MIQDGAMKGVSHILGVHVFPSIPAQQVGIRYGALTAAADDLEIFIQGESGHGARPHEAID 205
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
+ A+ + ++Q +SR NPL V+S+ +GG + ++I D V + GT R+ +
Sbjct: 206 AIWIAAQVITALQQAISRTQNPLRPMVLSLGQISGGRAPNVIADQVRMAGTVRSLHPETH 265
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
QL + IE ++ Y EV++ P ND Q+ + ++ + G+
Sbjct: 266 AQLPQWIEGIVANVCQTYGAKYEVNYRRG----VPSVQNDAQLNKLLENAVREAWGESAL 321
Query: 181 RVVP-PMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAVHA 239
+++P P +GAED++ Y + P A F +G H H P F DE A+ G +
Sbjct: 322 QIIPEPSLGAEDFALYLEHAPGAMFRLGTGFGDRQMNHPLHHPRFEADEAAILTGVVTLS 381
Query: 240 TIAERY 245
A +Y
Sbjct: 382 YAAWQY 387
>Y4TI_RHISN (P55663) Hypothetical hydrolase Y4TI (EC 3.-.-.-)
Length = 402
Score = 99.8 bits (247), Expect = 6e-21
Identities = 62/231 (26%), Positives = 110/231 (46%), Gaps = 6/231 (2%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
++++ LE + HV+ G G+R G + F+ +SG A + P N D
Sbjct: 157 VLEERLLEGFDNAVGFHVTPSIQVGKFGAREGAVSKSSDQFKVTVSGSAAHGSTPHNGID 216
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
+ A+A V +Q ++SRE D V+++ + +GG + ++I VV+ GT R +
Sbjct: 217 AITIAAAFVNEVQKVISREVPVDDRSVITIGTIHGGEATNIICPKVVMEGTIRTTNPELR 276
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNF 180
L +R+ ++ A+++ AEV E P +ND +M + D+ G
Sbjct: 277 PLLSQRVREIAEGVAALHRGKAEVVVTSGE----PAVINDPEMVRLFRDAVSDMAGSDAL 332
Query: 181 RVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDAL 231
+ G++D+ FYSQ IPS +F+ G + G+ H+P F + +D L
Sbjct: 333 TQGKAISGSDDFGFYSQCIPSIYFWFG--SGEPGNESGVHTPTFAVSDDVL 381
>HIPO_CAMJE (P45493) Hippurate hydrolase (EC 3.5.1.32)
(Benzoylglycine amidohydrolase) (Hippuricase)
Length = 383
Score = 96.7 bits (239), Expect = 5e-20
Identities = 64/251 (25%), Positives = 126/251 (49%), Gaps = 16/251 (6%)
Query: 1 MIQDGALE--DVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNS 58
MI+DG E D + +F H + G ++A + + G+ + P +
Sbjct: 143 MIEDGLFEKFDSDYVFGWHNMPFGSDKKFYLKKGAMMASSDSYSIEVIGRGGHGSAPEKA 202
Query: 59 ADPVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNT 118
DP+ AAS ++++Q IVSR +P +S VVS+ +FN G++ ++IPD I + RA N
Sbjct: 203 KDPIYAASLLIVALQSIVSRNVDPQNSAVVSIGAFNAGHAFNIIPDIATIKMSVRALDNE 262
Query: 119 SFYQLLERIEQVI--VQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLG 176
+ E+I ++ + QA+ +++ + + P T+N+D+ + +V+ +L G
Sbjct: 263 TRKLTEEKIYKICKGIAQAN------DIEIKINKNVVAPVTMNNDEAVDFASEVAKELFG 316
Query: 177 QKNFRV-VPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGA 235
+KN P+M +ED+ F+ ++ A+ ++ N+ H+ + ++ L A
Sbjct: 317 EKNCEFNHRPLMASEDFGFFCEMKKCAYAFLENENDIY-----LHNSSYVFNDKLLARAA 371
Query: 236 AVHATIAERYL 246
+ +A +A +YL
Sbjct: 372 SYYAKLALKYL 382
>AMAA_BACST (P37112) N-acyl-L-amino acid amidohydrolase (EC
3.5.1.14) (L-aminoacylase)
Length = 370
Score = 77.8 bits (190), Expect = 2e-14
Identities = 49/162 (30%), Positives = 79/162 (48%), Gaps = 4/162 (2%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M+Q G ++ V+ + H+ G IG GP++A F I GK A P + D
Sbjct: 149 MVQAGVMDGVDVVIGTHLWSPLERGKIGIVYGPMMAAPDRFFIRIIGKGGHGAMPHQTID 208
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
+ + V ++Q IVSR +PL+ V+SVT F G +H+++P V I GT R F T
Sbjct: 209 AIAIGAQVVTNLQHIVSRYVDPLEPLVLSVTQFVAGTAHNVLPGEVEIQGTVRTFDETLR 268
Query: 121 YQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQ 162
+ + +E+++ + E F +Y Y P +N D+
Sbjct: 269 RTVPQWMERIVKGITEAHGASYE---FRFDYG-YRPVINYDE 306
>AMHX_BACSU (P54983) Amidohydrolase amhX (EC 3.5.1.-) (Aminoacylase)
Length = 389
Score = 57.8 bits (138), Expect = 2e-08
Identities = 55/250 (22%), Positives = 102/250 (40%), Gaps = 18/250 (7%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
MI++G L+D++ ++ VHV T P L I G+ A A P +
Sbjct: 134 MIEEGVLDDIDYLYGVHVRPIQETQNGRCAPSILHGSSQHIEGTIIGEEAHGARPHLGKN 193
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFN-GGNSHDMIPDSVVIGGTFRAFSNTS 119
+ A+ V + G++ +P V +T GG S ++IP RA +N +
Sbjct: 194 SIEIAAFLVHKL-GLI--HIDPQIPHTVKMTKLQAGGESSNIIPGKASFSLDLRAQTNEA 250
Query: 120 FYQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKN 179
L+ E+ A+ + E+ KE+++ P + + + + +++G +
Sbjct: 251 MEALIAETERACEAAAAAFGAKIEL---HKEHSL-PAATQNKEAEAIMAEAITEIIGAER 306
Query: 180 FRVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLG---STHTG-HSPHFTIDEDALPIGA 235
G ED+ FY+ +P+ ++ LG G H PH T D +A+ G
Sbjct: 307 LDDPLVTTGGEDFHFYAVKVPN------LKTTMLGLGCGLQPGLHHPHMTFDRNAMFTGI 360
Query: 236 AVHATIAERY 245
+ A ++
Sbjct: 361 HILANACSKH 370
>ABGA_ECOLI (P77357) Aminobenzoyl-glutamate utilization protein A
Length = 436
Score = 43.9 bits (102), Expect = 4e-04
Identities = 31/143 (21%), Positives = 60/143 (41%), Gaps = 3/143 (2%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSAD 60
M+ G ++DV+ AVH+ P G + +A F + A P + +
Sbjct: 192 MVDAGVVDDVDYFTAVHIGTGVPAGTVVCGSDNFMATTKFDAHFTGTAAHAGAKPEDGHN 251
Query: 61 PVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSF 120
+LAA+ A +++ I + V+V G+ +++P S ++ R S+
Sbjct: 252 ALLAAAQATLALHAIAPHSEG---ASRVNVGVMQAGSGRNVVPASALLKVETRGASDVIN 308
Query: 121 YQLLERIEQVIVQQASVYSCFAE 143
+ +R +Q I A++Y E
Sbjct: 309 QYVFDRAQQAIQGAATMYGVGVE 331
>IAAH_ENTAG (O50173) Indole-3-acetyl-aspartic acid hydrolase (EC
3.5.1.-) (IAA-Asp hydrolase)
Length = 435
Score = 42.0 bits (97), Expect = 0.001
Identities = 33/145 (22%), Positives = 63/145 (42%), Gaps = 5/145 (3%)
Query: 1 MIQDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASA-ANPRNSA 59
M+ G ++DV+ A+H+ P G + +A F A+ +G A A P +
Sbjct: 191 MVAAGVVDDVDYFTAIHIGTGVPAGTVVCGSDNFMATTK-FDALFTGVAAHAGGKPEDGR 249
Query: 60 DPVLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTS 119
+ +LAA+ A I++ I + + V+V G +++P ++ R +
Sbjct: 250 NALLAAAQAAIALHAIAPHSAG---ASRVNVGVMQAGTGRNVVPSGALLKVETRGETEDI 306
Query: 120 FYQLLERIEQVIVQQASVYSCFAEV 144
+ ER +VI A++Y E+
Sbjct: 307 NRYVFERAREVIHGAAAMYGASVEL 331
>ABGA_HAEIN (P44765) Aminobenzoyl-glutamate utilization protein A
homolog
Length = 423
Score = 37.0 bits (84), Expect = 0.045
Identities = 48/250 (19%), Positives = 102/250 (40%), Gaps = 17/250 (6%)
Query: 3 QDGALEDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASA-ANPRNSADP 61
Q G ++D + + H+S TG + + P L+ GK A A A P +
Sbjct: 186 QSGIIDDADYFASSHISFCANTGTVIANPRNFLSTTK-IDIRYKGKPAHAGAAPHLGRNA 244
Query: 62 VLAASAAVISIQGIVSRESNPLDSQVVSVTSFNGGNSHDMIPDSVVIGGTFRAFSNTSFY 121
+LAA+ V + GI +R + ++V G ++IP S + R +
Sbjct: 245 LLAAAHTVTQLHGI-ARHGKGMTR--INVGVLKAGEGRNVIPSSAELQLEVRGENKAINE 301
Query: 122 QLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQMYEHVKKVSIDLLGQKNFR 181
+ E++ Q+ + ++ E + + + ND ++ + ++++S++ N
Sbjct: 302 YMTEQVMQIAKGISISFNVAYETEIVGEAVDMN----NDVELIKLIEEISLEQPQINNVN 357
Query: 182 VVPPMMGAEDYSFYSQVIP----SAFFYIGIRNETLGSTHTGHSPHFTIDEDALPIGAAV 237
+ED + + + A ++I + T G H F DE+ L G +
Sbjct: 358 SDYAFNASEDATILGRRVQEHGGKAIYFILGADRTAGH----HEAEFDFDENQLLTGVNI 413
Query: 238 HATIAERYLN 247
+ ++ ++ L+
Sbjct: 414 YTSLVQKLLS 423
>NMR_NEUCR (P23762) Nitrogen metabolic regulation protein (NMR
protein)
Length = 488
Score = 34.7 bits (78), Expect = 0.22
Identities = 35/137 (25%), Positives = 59/137 (42%), Gaps = 14/137 (10%)
Query: 49 RASAANPRNSADPVLAASAAVISIQGIVSRESNPLDS---QVVSVTSFNGGNSHDMIPDS 105
RA N ++ + V +QG + + P +S Q V VT NG S + D+
Sbjct: 92 RAQMRNLEGVVATEVSTNPNVTVLQGELYTKETPAESDKGQCVDVTK-NGPISGIGVNDA 150
Query: 106 VV---IGGTFRAFSNTSFYQLLERIEQVIVQQAS-------VYSCFAEVDFFEKEYTIYP 155
++ G AF NT+FY ERI + A VYS + + K++ P
Sbjct: 151 LISELFRGAQLAFINTTFYGDEERIGMALADAAKKAGVQHYVYSSMPDHHAYNKDWPSLP 210
Query: 156 PTVNDDQMYEHVKKVSI 172
+ ++ ++VK++ I
Sbjct: 211 LWASKHRVEDYVKEIGI 227
>C134_DROME (Q9VG40) Probable cytochrome P450 313a4 (EC 1.14.-.-)
(CYPCCCXIIIA4)
Length = 478
Score = 32.0 bits (71), Expect = 1.4
Identities = 22/85 (25%), Positives = 42/85 (48%), Gaps = 15/85 (17%)
Query: 105 SVVIGGTFRAFSNTSFYQLLERIEQVIVQQASVYSCFAEVD--FFEKEYTIYPPTVNDDQ 162
++++ G F +N +Y L+ + + F E FE+ TI+P T + D
Sbjct: 278 NIIVFGAFETTANAVYYTLM------------LLAMFPEYQERAFEEIKTIFPNTGDFDV 325
Query: 163 MYEHVKK-VSIDLLGQKNFRVVPPM 186
Y ++ V +DL+ ++ RV+PP+
Sbjct: 326 SYADTQQMVYLDLILNESMRVIPPV 350
>POL2_TORVR (P25247) RNA2 polyprotein (P2) [Contains: X3 protein; X4
protein; Movement protein (MP); Coat protein (CP)]
Length = 1882
Score = 31.2 bits (69), Expect = 2.5
Identities = 31/120 (25%), Positives = 51/120 (41%), Gaps = 8/120 (6%)
Query: 110 GTFRAFSNTSFYQLLERIEQVIVQQASVYSCFAEVDFFEKEYTIYPPTVNDDQM--YEHV 167
G+F A +F + +E IEQ Q V +C A + K YT Y + M Y +V
Sbjct: 1743 GSFSA--RLAFGKGVEEIEQTSTVQPLVGACEARIPVEFKTYTGYTTSGPPGSMEPYIYV 1800
Query: 168 KKVSIDLLGQKNFRVVPPMMGAEDYSFYSQVIPSAFFYIGIRNETLGSTHTGHSPHFTID 227
+ L+ + + V+ E +SFY + +G + TLG+ + P +D
Sbjct: 1801 RLTQAKLVDRLSVNVIL----QEGFSFYGPSVKHFKKEVGTPSATLGTNNPVGRPPENVD 1856
>ULB7_HCMVA (P16770) Hypothetical protein UL117
Length = 424
Score = 30.8 bits (68), Expect = 3.2
Identities = 27/103 (26%), Positives = 45/103 (43%), Gaps = 9/103 (8%)
Query: 8 EDVEAIFAVHVSHEHPTGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSADPVLAASA 67
E A V + P G++G G L AG G A ++ A+ A +A+PV+ A+A
Sbjct: 153 ESAPATAEVCLGDALPGGVMGG--GGLPAGVGSASAAVAAAAAAVAGVPVAANPVMPATA 210
Query: 68 AVIS--IQGIVSRES-----NPLDSQVVSVTSFNGGNSHDMIP 103
V + + + S P + + NGGN+ ++P
Sbjct: 211 TVTTPPMIDLTSHHRPLTLFTPASAAAAPAVATNGGNATYILP 253
>TRPC_METTM (P26939) Indole-3-glycerol phosphate synthase (EC
4.1.1.48) (IGPS)
Length = 274
Score = 30.4 bits (67), Expect = 4.2
Identities = 14/42 (33%), Positives = 23/42 (54%)
Query: 24 TGMIGSRPGPLLAGCGFFRAVISGKRASAANPRNSADPVLAA 65
+G++G +LAGCG +I A++PR D ++AA
Sbjct: 212 SGVMGPEDAEILAGCGADALLIGTSLMLASDPRELLDEIMAA 253
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.135 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,705,895
Number of Sequences: 164201
Number of extensions: 1191575
Number of successful extensions: 2526
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 17
Number of HSP's successfully gapped in prelim test: 9
Number of HSP's that attempted gapping in prelim test: 2491
Number of HSP's gapped (non-prelim): 26
length of query: 250
length of database: 59,974,054
effective HSP length: 108
effective length of query: 142
effective length of database: 42,240,346
effective search space: 5998129132
effective search space used: 5998129132
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC140916.4


