

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140913.9 - phase: 2 /pseudo
(273 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
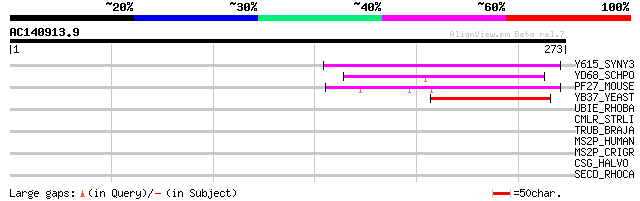
Y615_SYNY3 (P52876) Hypothetical UPF0016 protein sll0615 78 3e-14
YD68_SCHPO (Q10320) Hypothetical UPF0016 protein C17G8.08c in ch... 62 2e-09
PF27_MOUSE (P52875) Transmembrane protein PFT27 (TPA regulated l... 57 6e-08
YB37_YEAST (P38301) Hypothetical UPF0016 protein YBR187w 54 5e-07
UBIE_RHOBA (P59912) Menaquinone biosynthesis methyltransferase u... 35 0.20
CMLR_STRLI (P31141) Chloramphenicol resistance protein 32 2.2
TRUB_BRAJA (Q89WB1) tRNA pseudouridine synthase B (EC 4.2.1.70) ... 30 4.8
MS2P_HUMAN (O43462) Membrane-bound transcription factor site 2 p... 30 4.8
MS2P_CRIGR (O54862) Membrane-bound transcription factor site 2 p... 30 4.8
CSG_HALVO (P25062) Cell surface glycoprotein precursor (S-layer ... 30 6.3
SECD_RHOCA (O33517) Protein-export membrane protein secD 30 8.2
>Y615_SYNY3 (P52876) Hypothetical UPF0016 protein sll0615
Length = 206
Score = 77.8 bits (190), Expect = 3e-14
Identities = 42/117 (35%), Positives = 67/117 (56%)
Query: 155 AAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIVSTFLLV 214
A V L + FG L DA + + +E ++AE A++ + +V +F L
Sbjct: 70 AEVALFLIFGTKLLWDARRIKATANLEEMEDAEKAIASGEKKLKIVPRGWGIVVESFALT 129
Query: 215 FVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVTS 271
FVAEWGD++ +TIALAA+++ GV AG++ GH + +IAV+GG + +SEK +
Sbjct: 130 FVAEWGDRTQIATIALAASNNAWGVSAGAILGHTICAVIAVMGGKFVAGRISEKTVT 186
Score = 35.8 bits (81), Expect = 0.11
Identities = 21/54 (38%), Positives = 31/54 (56%), Gaps = 1/54 (1%)
Query: 212 LLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFL 265
LL+ V+E GDK+FF + LA V+ G + G T+++VL G + TFL
Sbjct: 10 LLITVSELGDKTFFIAMILAMRYPRRWVLVGVVGGLAAMTILSVLMGQIF-TFL 62
>YD68_SCHPO (Q10320) Hypothetical UPF0016 protein C17G8.08c in
chromosome I
Length = 287
Score = 61.6 bits (148), Expect = 2e-09
Identities = 30/101 (29%), Positives = 51/101 (49%), Gaps = 2/101 (1%)
Query: 165 VSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAA--ASTIVSTFLLVFVAEWGDK 222
+ LL+ ++ S + + +S G ++A + + F L FV+EWGD+
Sbjct: 159 IDQLLEEGAAPSHYTGHRSRSGHTLMSQLKSKGRNVMATLFSPLFIKAFALTFVSEWGDR 218
Query: 223 SFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGT 263
S +TIA+AA+ + GV G+ GH T +AV+ G + T
Sbjct: 219 SQIATIAMAASDNVYGVFMGANVGHACCTALAVISGKYIST 259
Score = 36.2 bits (82), Expect = 0.088
Identities = 21/65 (32%), Positives = 33/65 (50%)
Query: 206 TIVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFL 265
+++ + ++F E GDK+F LA +S L V AGS + + TL+ VL G
Sbjct: 49 SLIFSISMIFGCEIGDKTFIVAALLAFENSRLTVFAGSYSALFIMTLLGVLLGHAAPLLF 108
Query: 266 SEKVT 270
K+T
Sbjct: 109 PRKLT 113
>PF27_MOUSE (P52875) Transmembrane protein PFT27 (TPA regulated
locus protein)
Length = 323
Score = 56.6 bits (135), Expect = 6e-08
Identities = 40/138 (28%), Positives = 60/138 (42%), Gaps = 22/138 (15%)
Query: 156 AVCLLVYFGVSTLLDA--SSSDSQKSDDEQKEAELAVSDFSG------DGAGILAAAST- 206
+ L FG+ L + S D + + E+ +AEL D +G + ST
Sbjct: 163 STALFAIFGIRMLREGLKMSPDEGQEELEEVQAELKKKDEEFQRTKLLNGPDVETGTSTA 222
Query: 207 -------------IVSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLI 253
V L F+AEWGD+S +TI LAA P GV G GH + T +
Sbjct: 223 IPQKKWLHFISPIFVQALTLTFLAEWGDRSQLTTIVLAAREDPYGVAVGGTVGHCLCTGL 282
Query: 254 AVLGGSLLGTFLSEKVTS 271
AV+GG ++ +S + +
Sbjct: 283 AVIGGRMIAQKISVRTVT 300
Score = 34.3 bits (77), Expect = 0.33
Identities = 17/51 (33%), Positives = 30/51 (58%)
Query: 208 VSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGG 258
V+ ++ V+E GDK+FF +A + L V+AG++ + T ++VL G
Sbjct: 98 VAAISVIIVSELGDKTFFIAAIMAMRYNRLTVLAGAMLALALMTCLSVLFG 148
>YB37_YEAST (P38301) Hypothetical UPF0016 protein YBR187w
Length = 280
Score = 53.5 bits (127), Expect = 5e-07
Identities = 28/59 (47%), Positives = 39/59 (65%)
Query: 208 VSTFLLVFVAEWGDKSFFSTIALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLS 266
V FL+VF+ E GD+S S IA+A S VIAG++ GH + + +AV+GG LL T +S
Sbjct: 194 VQIFLMVFLGELGDRSQISIIAMATDSDYWYVIAGAVIGHAICSGLAVVGGKLLATRIS 252
>UBIE_RHOBA (P59912) Menaquinone biosynthesis methyltransferase ubiE
(EC 2.1.1.-)
Length = 291
Score = 35.0 bits (79), Expect = 0.20
Identities = 26/105 (24%), Positives = 45/105 (42%), Gaps = 4/105 (3%)
Query: 149 LPIDDIAAVCLLVYFGVSTLLDASSSDSQKSDDEQKEAELAVSDFSGDGAGILAAASTIV 208
LP DD A C+ V FG+ + D S+ + + ++ V +FS +L A
Sbjct: 145 LPFDDDAFQCVTVAFGLRNIADTDQGLSEMARVCKPGGQVLVLEFSQPTLPVLKQAYNFY 204
Query: 209 STFLLVFVAEW---GDKSFFSTIALAAASSPLG-VIAGSLAGHGV 249
+L + +W DKS + + + P G +AG + G+
Sbjct: 205 FRHVLPRIGQWMARNDKSAYEYLPESVGKFPCGDALAGRMRDVGL 249
>CMLR_STRLI (P31141) Chloramphenicol resistance protein
Length = 392
Score = 31.6 bits (70), Expect = 2.2
Identities = 19/42 (45%), Positives = 24/42 (56%)
Query: 229 ALAAASSPLGVIAGSLAGHGVATLIAVLGGSLLGTFLSEKVT 270
AL A +A L+G VAT+ V GGSLLGT+L + T
Sbjct: 118 ALVPADKQGRALAVLLSGTTVATVAGVPGGSLLGTWLGWRAT 159
>TRUB_BRAJA (Q89WB1) tRNA pseudouridine synthase B (EC 4.2.1.70)
(tRNA pseudouridine 55 synthase) (Psi55 synthase)
(Pseudouridylate synthase) (Uracil hydrolyase)
Length = 345
Score = 30.4 bits (67), Expect = 4.8
Identities = 20/54 (37%), Positives = 28/54 (51%), Gaps = 8/54 (14%)
Query: 119 VLFSLGHLAHLRRTFHYVDELLPFRFGETDL-PIDDIAAVCLLVYFGVSTLLDA 171
+L + GH+ LRRT L FGE D+ P+D + A+C G +L DA
Sbjct: 215 ILGTYGHICALRRT-------LVGPFGENDMIPLDQLEALCDRAASGEGSLADA 261
>MS2P_HUMAN (O43462) Membrane-bound transcription factor site 2
protease (EC 3.4.24.85) (S2P endopeptidase) (Site-2
protease) (Sterol-regulatory element-binding proteins
intramembrane protease)
Length = 519
Score = 30.4 bits (67), Expect = 4.8
Identities = 19/59 (32%), Positives = 27/59 (45%), Gaps = 4/59 (6%)
Query: 93 FC*YFSLNWETRLFSLQHC*QLEIQPVLFSLGHLAHLRRTFHYVDELLPFRFGETDLPI 151
FC SL TRL ++H Q++ + +GH HL T + F F DLP+
Sbjct: 385 FCIIPSLETHTRLIKVKHPPQID----MLYVGHPLHLHYTVSITSFIPRFNFLSIDLPV 439
>MS2P_CRIGR (O54862) Membrane-bound transcription factor site 2
protease (EC 3.4.24.85) (S2P endopeptidase) (Site-2
protease) (Sterol-regulatory element-binding proteins
intramembrane protease)
Length = 510
Score = 30.4 bits (67), Expect = 4.8
Identities = 19/59 (32%), Positives = 27/59 (45%), Gaps = 4/59 (6%)
Query: 93 FC*YFSLNWETRLFSLQHC*QLEIQPVLFSLGHLAHLRRTFHYVDELLPFRFGETDLPI 151
FC SL TRL ++H Q++ + +GH HL T + F F DLP+
Sbjct: 376 FCIVPSLETHTRLIKVKHPPQID----MLYVGHPLHLHYTVSITSFIPRFNFLSIDLPV 430
>CSG_HALVO (P25062) Cell surface glycoprotein precursor (S-layer
protein)
Length = 827
Score = 30.0 bits (66), Expect = 6.3
Identities = 14/31 (45%), Positives = 20/31 (64%)
Query: 176 SQKSDDEQKEAELAVSDFSGDGAGILAAAST 206
S SDD +E ++A+S SGDG+ IL+ T
Sbjct: 435 SVDSDDTFEEEDIALSGLSGDGSSILSLTGT 465
>SECD_RHOCA (O33517) Protein-export membrane protein secD
Length = 554
Score = 29.6 bits (65), Expect = 8.2
Identities = 19/67 (28%), Positives = 37/67 (54%), Gaps = 5/67 (7%)
Query: 198 AGILAAASTIVSTFLLVFVAEWGDKSFFSTIAL----AAASSPLGVIAGSLAGHGVATLI 253
AG++A+ V+ + +A +G FFS++AL A + +G I G++ G+A ++
Sbjct: 392 AGMVASVIGFVAV-VAYMIASYGLFGFFSSVALFINIAFIFAVMGAIGGTMTLPGIAGIV 450
Query: 254 AVLGGSL 260
+G S+
Sbjct: 451 LTIGTSV 457
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.138 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,749,973
Number of Sequences: 164201
Number of extensions: 1105567
Number of successful extensions: 4850
Number of sequences better than 10.0: 11
Number of HSP's better than 10.0 without gapping: 7
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 4839
Number of HSP's gapped (non-prelim): 14
length of query: 273
length of database: 59,974,054
effective HSP length: 109
effective length of query: 164
effective length of database: 42,076,145
effective search space: 6900487780
effective search space used: 6900487780
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 65 (29.6 bits)
Medicago: description of AC140913.9


