

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140849.2 + phase: 0
(553 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
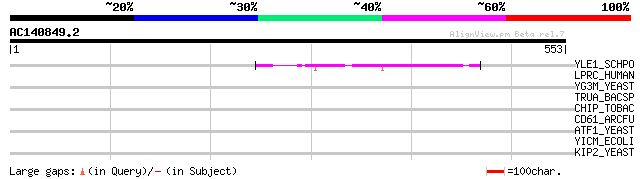
Score E
Sequences producing significant alignments: (bits) Value
YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I 45 6e-04
LPRC_HUMAN (P42704) 130 kDa leucine-rich protein (LRP 130) (GP13... 37 0.13
YG3M_YEAST (P48237) Hypothetical 101.4 kDa protein in RPL24B-RSR... 36 0.23
TRUA_BACSP (Q45557) tRNA pseudouridine synthase A (EC 4.2.1.70) ... 35 0.67
CHIP_TOBAC (P17513) Acidic endochitinase P precursor (EC 3.2.1.1... 32 4.3
CD61_ARCFU (O29995) Cell division control protein 6 homolog 1 (C... 32 5.6
ATF1_YEAST (P40353) Alcohol O-acetyltransferase 1 (EC 2.3.1.84) ... 32 5.6
YICM_ECOLI (P31438) Hypothetical protein yicM 31 7.4
KIP2_YEAST (P28743) Kinesin-like protein KIP2 31 9.6
>YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I
Length = 1261
Score = 44.7 bits (104), Expect = 6e-04
Identities = 51/241 (21%), Positives = 103/241 (42%), Gaps = 52/241 (21%)
Query: 246 LVSMYSRCKCIDSAYRVFCGIFNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKK-- 303
L+S+ S+ K D+ RVF E+ K L +RK++ KS +
Sbjct: 813 LISVLSKNKRFDAVQRVF----------------------EHSKHL--YRKISTKSLEKA 848
Query: 304 -------LDSVLIATVLASITQMANVL-PGCEIHGYVLRHGLESDVKVSSALIDMYSKCG 355
LD++++++ A + +N+ ++ GY+ R + LI+ ++ G
Sbjct: 849 NWFMALILDAMILSSSFARQFKSSNLFCDNMKMLGYIPR------ASTFAHLINNSTRRG 902
Query: 356 FLHLGTCVFRIMLER-------NIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVPDEG 408
T I E ++ YN+++ G ++ + +F EM + GL+P
Sbjct: 903 DTDDATTALNIFEETKRHNVKPSVFLYNAVLSKLGRARRTTECWKLFQEMKESGLLPTSV 962
Query: 409 TFSALLSACCHAGLVKDGRELFWRMKDEFNIKARPEHYVYMVKLLGGVGELEEAYNLTQS 468
T+ +++A C G +LF M+++ N + R Y M++ E++ +N ++
Sbjct: 963 TYGTVINAACRIGDESLAEKLFAEMENQPNYQPRVAPYNTMIQF-----EVQTMFNREKA 1017
Query: 469 L 469
L
Sbjct: 1018 L 1018
Score = 34.7 bits (78), Expect = 0.67
Identities = 19/61 (31%), Positives = 32/61 (52%), Gaps = 4/61 (6%)
Query: 57 AHHVFDKTSTR----SVFLWNSMIRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAIR 112
A ++F++T SVFL+N+++ +ARR + LF+ M + P + TY I
Sbjct: 910 ALNIFEETKRHNVKPSVFLYNAVLSKLGRARRTTECWKLFQEMKESGLLPTSVTYGTVIN 969
Query: 113 A 113
A
Sbjct: 970 A 970
>LPRC_HUMAN (P42704) 130 kDa leucine-rich protein (LRP 130) (GP130)
(Leucine-rich PPR-motif containing protein)
Length = 1273
Score = 37.0 bits (84), Expect = 0.13
Identities = 33/140 (23%), Positives = 60/140 (42%), Gaps = 5/140 (3%)
Query: 336 GLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNI----ISYNSMILAYGLHGCASQA 391
G DV +AL+ +Y + + T M E NI ++Y +I +Y G A
Sbjct: 36 GAVYDVSHYNALLKVYLQNEYKFSPTDFLAKMEEANIQPNRVTYQRLIASYCNVGDIEGA 95
Query: 392 FTMFDEMLQKGLVPDEGTFSALLSACCHAGLVKDGRELFWRMKDEFNIKARPEHYVYMVK 451
+ M K L E FSAL++ AG +++ + M+D I+ P+ Y+ ++
Sbjct: 96 SKILGFMKTKDLPVTEAVFSALVTGHARAGDMENAENILTVMRDA-GIEPGPDTYLALLN 154
Query: 452 LLGGVGELEEAYNLTQSLPK 471
G+++ + + K
Sbjct: 155 AYAEKGDIDHVKQTLEKVEK 174
Score = 31.2 bits (69), Expect = 7.4
Identities = 27/137 (19%), Positives = 58/137 (41%), Gaps = 8/137 (5%)
Query: 56 YAHHVFDKT----STRSVFLWNSMIRAFAKARRFSNAISLFRTMLVDDIRPDNYTYACAI 111
+AH ++D + V +N++++ + + + M +I+P+ TY I
Sbjct: 24 FAHRIWDTLQKLGAVYDVSHYNALLKVYLQNEYKFSPTDFLAKMEEANIQPNRVTYQRLI 83
Query: 112 RACADSFDFGMLRVVHGSAVSVGLGLDPICCSALVSAYSKLGVVHEARRVF----DGIVE 167
+ + D + G + L + SALV+ +++ G + A + D +E
Sbjct: 84 ASYCNVGDIEGASKILGFMKTKDLPVTEAVFSALVTGHARAGDMENAENILTVMRDAGIE 143
Query: 168 PDLVLWNSLISAYGGSG 184
P + +L++AY G
Sbjct: 144 PGPDTYLALLNAYAEKG 160
>YG3M_YEAST (P48237) Hypothetical 101.4 kDa protein in RPL24B-RSR1
intergenic region
Length = 864
Score = 36.2 bits (82), Expect = 0.23
Identities = 23/82 (28%), Positives = 36/82 (43%), Gaps = 6/82 (7%)
Query: 42 TQIIRLYAFNNHINYAHHVFDKTSTRS------VFLWNSMIRAFAKARRFSNAISLFRTM 95
T +I+ Y+ A + FD S + +N+M+R K R F A+ LF+ +
Sbjct: 324 TTVIQFYSRKEMTKQAWNTFDTMKFLSTKHFPDICTYNTMLRICEKERNFPKALDLFQEI 383
Query: 96 LVDDIRPDNYTYACAIRACADS 117
+I+P TY R A S
Sbjct: 384 QDHNIKPTTNTYIMMARVLASS 405
>TRUA_BACSP (Q45557) tRNA pseudouridine synthase A (EC 4.2.1.70)
(Pseudouridylate synthase I) (Pseudouridine synthase I)
(Uracil hydrolyase)
Length = 244
Score = 34.7 bits (78), Expect = 0.67
Identities = 28/108 (25%), Positives = 45/108 (40%), Gaps = 8/108 (7%)
Query: 31 KTHLSK--DPFYATQIIRLYAFNNHINYAHHVFDKTSTRSVFLWNSMIRAFAKARRFSNA 88
KT+L K + Y +R Y+ HI H+ + +F+ AF+ A+
Sbjct: 109 KTYLYKIWNEDYTHPFMRKYSL--HIEKKLHIDNMVKASQLFVGEHDFTAFSNAKSKKKT 166
Query: 89 ISLFRTMLVDDIRPDNYTYACAIRACADSFDFGMLRVVHGSAVSVGLG 136
T + I + IR C D F + M+R + G+ + VGLG
Sbjct: 167 ----NTRTIHSITIQDNQGFIDIRVCGDGFLYNMVRKMVGTLIEVGLG 210
>CHIP_TOBAC (P17513) Acidic endochitinase P precursor (EC 3.2.1.14)
(Pathogenesis-related protein P) (PR-P)
Length = 253
Score = 32.0 bits (71), Expect = 4.3
Identities = 30/117 (25%), Positives = 48/117 (40%), Gaps = 11/117 (9%)
Query: 268 NPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIATVLASIT-QMANVLPGC 326
NPDLV A IS ++ A+ F+ V+I + S Q AN PGC
Sbjct: 147 NPDLVATDATIS-------FKTAIWFWMTPQDNKPSSHDVIIGSWTPSAADQSANRAPGC 199
Query: 327 EIHGYVLRHGLESDVKVSSALIDMYSKCGFLHLGTCVFRIMLERNIISYNSMILAYG 383
+ ++ G+E V ++A+ D + G+ + + N+ YN A G
Sbjct: 200 GVITNIINGGIECGVGPNAAVED---RIGYYRRYCGMLNVAPGDNLDCYNQRNFAQG 253
>CD61_ARCFU (O29995) Cell division control protein 6 homolog 1 (CDC6
homolog 1)
Length = 409
Score = 31.6 bits (70), Expect = 5.6
Identities = 19/77 (24%), Positives = 35/77 (44%), Gaps = 8/77 (10%)
Query: 190 IQMFSSMRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQKSGLDSDCHVGSLLVSM 249
+ + S L G P + G G +++L + ++L S+K+ L+ H
Sbjct: 39 LALLLSPMLRGGTPSNIFIYGKTGTGKTATVLFVARQLEEASRKAKLNVAVH-------- 90
Query: 250 YSRCKCIDSAYRVFCGI 266
Y C+ +D+AYRV +
Sbjct: 91 YINCEIVDTAYRVLASL 107
>ATF1_YEAST (P40353) Alcohol O-acetyltransferase 1 (EC 2.3.1.84)
(AATase 1)
Length = 525
Score = 31.6 bits (70), Expect = 5.6
Identities = 21/59 (35%), Positives = 30/59 (50%), Gaps = 11/59 (18%)
Query: 423 VKDGRE---LFWRMKDEFN-IKARPEHYVYMVKLLGGVGELEEAYNLTQSLPKPVDKAI 477
+ DGR F ++DE N IK P+ Y+ K EE Y L + LP+P++K I
Sbjct: 193 MSDGRSSIHFFHDLRDELNNIKTPPKKLDYIFKY-------EEDYQLLRKLPEPIEKVI 244
>YICM_ECOLI (P31438) Hypothetical protein yicM
Length = 412
Score = 31.2 bits (69), Expect = 7.4
Identities = 43/158 (27%), Positives = 68/158 (42%), Gaps = 26/158 (16%)
Query: 196 MRLAGKKPDGFTLAGLLGGIADSSLLSIGQELHGLSQK---SGLDSDCHVGSLLVSMYSR 252
M LAG DG TL L GIA S+ + S K +G V +L+++++
Sbjct: 260 MNLAGFGVDGLTLVLLSFGIASFIGTSLSSFILKRSVKLALAGAPLILAVSALVLTLWGS 319
Query: 253 CKCIDSAYRVFCGI-FNPDLVTWSALISGYSQCGEYQKALLFFRKLNMKSKKLDSVLIAT 311
K + + + G+ F V WS I+ R L +++K S+ +A
Sbjct: 320 DKIVATGVAIIWGLTFALVPVGWSTWIT---------------RSLADQAEKAGSIQVAV 364
Query: 312 VLASITQMANVLPGCEIHGYVLRH-GLESDVKVSSALI 348
+ Q+AN G I GY L + GL S + +S L+
Sbjct: 365 I-----QLANTC-GAAIGGYALDNIGLTSPLMLSGTLM 396
>KIP2_YEAST (P28743) Kinesin-like protein KIP2
Length = 706
Score = 30.8 bits (68), Expect = 9.6
Identities = 16/54 (29%), Positives = 30/54 (54%), Gaps = 1/54 (1%)
Query: 352 SKCGFLHLGTCVFRIMLERNIISYNSMILAYGLHGCASQAFTMFDEMLQKGLVP 405
S C L + + M+++ ++ +N+ I AYG+ G + + FTM + GL+P
Sbjct: 171 SHCTNLEVYERTSKPMIDKLLMGFNATIFAYGMTG-SGKTFTMSGNEQELGLIP 223
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.324 0.138 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 63,392,352
Number of Sequences: 164201
Number of extensions: 2608265
Number of successful extensions: 7114
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 2
Number of HSP's successfully gapped in prelim test: 7
Number of HSP's that attempted gapping in prelim test: 7103
Number of HSP's gapped (non-prelim): 16
length of query: 553
length of database: 59,974,054
effective HSP length: 115
effective length of query: 438
effective length of database: 41,090,939
effective search space: 17997831282
effective search space used: 17997831282
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 68 (30.8 bits)
Medicago: description of AC140849.2


