

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140721.10 + phase: 0 /pseudo
(329 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
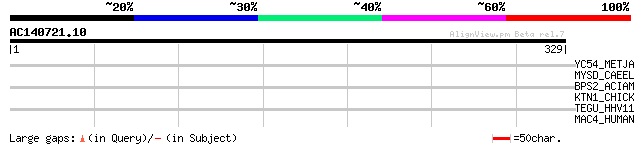
Score E
Sequences producing significant alignments: (bits) Value
YC54_METJA (Q58651) Hypothetical protein MJ1254 33 0.98
MYSD_CAEEL (P02567) Myosin heavy chain D (MHC D) 33 0.98
BPS2_ACIAM (P32985) Protein bps2 32 2.9
KTN1_CHICK (Q90631) Kinectin 30 6.4
TEGU_HHV11 (P10220) Large tegument protein (Virion protein UL36) 30 8.3
MAC4_HUMAN (Q96PK2) Microtubule-actin crosslinking factor 1, iso... 30 8.3
>YC54_METJA (Q58651) Hypothetical protein MJ1254
Length = 469
Score = 33.1 bits (74), Expect = 0.98
Identities = 40/185 (21%), Positives = 75/185 (39%), Gaps = 31/185 (16%)
Query: 170 VAELESKLTEQSKILLNLQQKVDDL------ASKHAYNPDE-------EYTEVESQLRAH 216
+ L++KL E+ + + NL K+ DL A+K+ N DE +E ESQ++
Sbjct: 289 LTSLKNKLKEKEEEIKNLAIKIKDLEDKLSKANKNLLNKDEIISVLNERISEYESQIQKL 348
Query: 217 LES-------------FLETARTFNMIYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKM 263
L+ ++ET + N ++R + + + G + L E K+
Sbjct: 349 LDENIIYKEKIESLNKYIETLKKENDKLKDKVRELSDIAKKYMEEKRGVLWSFLTEDEKI 408
Query: 264 LLKFLGNLQNLRDSHAALAFGSSETSGGPSSVSRIISECESEMTVINRDLGILSASIARE 323
++ + ++ G S+ VSRIISE E + +G ++ E
Sbjct: 409 IIDLIKKHGHITQKEIVEITGMSKPK-----VSRIISELEDRKIIRKEKIGRINKLTLTE 463
Query: 324 HGEKM 328
+K+
Sbjct: 464 ESKKL 468
>MYSD_CAEEL (P02567) Myosin heavy chain D (MHC D)
Length = 1938
Score = 33.1 bits (74), Expect = 0.98
Identities = 39/143 (27%), Positives = 67/143 (46%), Gaps = 11/143 (7%)
Query: 170 VAELESKLTEQSKILLNLQQKVDDLASK-HAYNPDEEYTEVES-QLRAHLESF---LETA 224
+A LE K KI+ + ++KVDD+ + A D T E +LR+ +++ +ET
Sbjct: 1446 IASLEKKQRAFDKIVDDWKRKVDDIQKEIDATTRDSRNTSTEVFKLRSSMDNLSEQIETL 1505
Query: 225 RTFNMIYTKEIRPWTHMMEVPQLHGFGPAANRLLEAYKMLLKFLGNLQN-LRDSHAALAF 283
R N I+++EIR Q+ G + ++ + L + LQ+ L ++ AAL
Sbjct: 1506 RRENKIFSQEIRDINE-----QITQGGRTYQEVHKSVRRLEQEKDELQHALDEAEAALEA 1560
Query: 284 GSSETSGGPSSVSRIISECESEM 306
S+ V +I SE E +
Sbjct: 1561 EESKVLRLQIEVQQIRSEIEKRI 1583
>BPS2_ACIAM (P32985) Protein bps2
Length = 582
Score = 31.6 bits (70), Expect = 2.9
Identities = 21/61 (34%), Positives = 32/61 (52%), Gaps = 3/61 (4%)
Query: 170 VAELESKLTEQSKILLNLQQKVDDLASKH--AYNPDEEYTEVESQLRAHLESFLETARTF 227
+ EL+SK E L NLQ ++DDL KH A + ++E ++ A LE E+ +
Sbjct: 130 INELKSKKEEMETKLHNLQAEIDDLNRKHKEAIELQTKIKQIEDEI-ARLEKEKESDKVL 188
Query: 228 N 228
N
Sbjct: 189 N 189
>KTN1_CHICK (Q90631) Kinectin
Length = 1364
Score = 30.4 bits (67), Expect = 6.4
Identities = 42/177 (23%), Positives = 69/177 (38%), Gaps = 38/177 (21%)
Query: 170 VAELESKLTEQSKILLNLQQKVDDLASKHAYNPDEEYTEVES--QLRAHLESFLETARTF 227
V E + +L + + +L+ +V+ L EE EVE+ + R HLES LE A
Sbjct: 1174 VEESQKELKQMRSSVASLEHEVERLK--------EEIKEVETLKKEREHLESELEKAEIE 1225
Query: 228 NMIYTKEIRPWTHMM-----EVPQLHGFGPAANRLLEAYKMLL---------------KF 267
Y E+R ++ ++ + N L KM L K
Sbjct: 1226 RSTYVSEVRELKDLLTELQKKLDDSYSEAVRQNEELNLLKMKLSETLSKLKVDQNERQKV 1285
Query: 268 LGNLQNLRDSHAALAFGSSETSGGPSSVSRIISECESEMT--------VINRDLGIL 316
G+L ++S A+L + G + + ESE+T +N+D+G L
Sbjct: 1286 AGDLPKAQESLASLEREIGKVVGDANVIENSDVRTESELTDKRLNVAVNLNQDVGHL 1342
>TEGU_HHV11 (P10220) Large tegument protein (Virion protein UL36)
Length = 3164
Score = 30.0 bits (66), Expect = 8.3
Identities = 32/128 (25%), Positives = 52/128 (40%), Gaps = 24/128 (18%)
Query: 189 QKVDDLASKHAYNPDEEYTEVESQLRAHLESF---LETARTFNMIYTKEIRPWTHMMEVP 245
+++ LA H YNP + E L + ++ LETA FN YT E + + +
Sbjct: 1382 RRLQALAGTHGYNPRDFRKRAEQALGTNAKAVTLALETALAFNP-YTPENQRHPMLPPLA 1440
Query: 246 QLH------GFGPAANRLLEAYKM-------LLKFLGNL-------QNLRDSHAALAFGS 285
+H FG AA+ + +++ LL+ G L D H A+ S
Sbjct: 1441 AIHRIDWSAAFGAAADTYADMFRVDTEPLARLLRLAGGLLERAQANDGFIDYHEAVLHLS 1500
Query: 286 SETSGGPS 293
+ G P+
Sbjct: 1501 EDLGGVPA 1508
>MAC4_HUMAN (Q96PK2) Microtubule-actin crosslinking factor 1, isoform
4
Length = 5938
Score = 30.0 bits (66), Expect = 8.3
Identities = 31/112 (27%), Positives = 47/112 (41%), Gaps = 7/112 (6%)
Query: 204 EEYTEVESQLRAHLESFLETARTFNMIYTKEIRPWTHMMEVPQLHG-FGPAA--NRLLEA 260
E TEV + LE F + ++RPW +ME + G GP + +L A
Sbjct: 2994 ERATEVTVARQRQLEESASHLACFQAAES-QLRPW--LMEKELMMGVLGPLSIDPNMLNA 3050
Query: 261 YKMLLKF-LGNLQNLRDSHAALAFGSSETSGGPSSVSRIISECESEMTVINR 311
K ++F L + R H L + GP VS S+ + E+ IN+
Sbjct: 3051 QKQQVQFMLKEFEARRQQHEQLNEAAQGILTGPGDVSLSTSQVQKELQSINQ 3102
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.332 0.143 0.430
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,249,880
Number of Sequences: 164201
Number of extensions: 1239248
Number of successful extensions: 4129
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 0
Number of HSP's successfully gapped in prelim test: 6
Number of HSP's that attempted gapping in prelim test: 4125
Number of HSP's gapped (non-prelim): 8
length of query: 329
length of database: 59,974,054
effective HSP length: 111
effective length of query: 218
effective length of database: 41,747,743
effective search space: 9101007974
effective search space used: 9101007974
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (22.0 bits)
S2: 66 (30.0 bits)
Medicago: description of AC140721.10


