

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140720.4 + phase: 0
(653 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
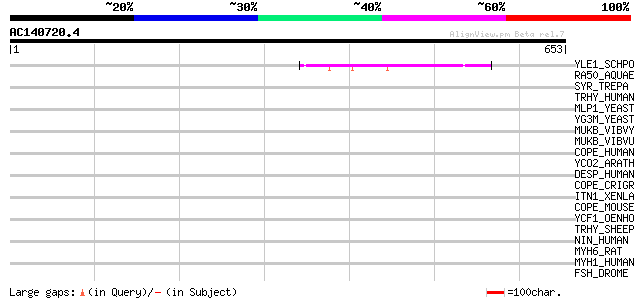
Score E
Sequences producing significant alignments: (bits) Value
YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I 52 5e-06
RA50_AQUAE (O67124) Probable DNA double-strand break repair rad5... 42 0.004
SYR_TREPA (O83803) Arginyl-tRNA synthetase (EC 6.1.1.19) (Argini... 42 0.005
TRHY_HUMAN (Q07283) Trichohyalin 39 0.033
MLP1_YEAST (Q02455) MLP1 protein (Myosin-like protein 1) 39 0.056
YG3M_YEAST (P48237) Hypothetical 101.4 kDa protein in RPL24B-RSR... 38 0.095
MUKB_VIBVY (Q7MJ64) Chromosome partition protein mukB (Structura... 36 0.28
MUKB_VIBVU (Q8DAP8) Chromosome partition protein mukB (Structura... 36 0.28
COPE_HUMAN (O14579) Coatomer epsilon subunit (Epsilon-coat prote... 36 0.28
YCO2_ARATH (Q9LME2) Hypothetical protein At1g22260 36 0.36
DESP_HUMAN (P15924) Desmoplakin (DP) (250/210 kDa paraneoplastic... 35 0.47
COPE_CRIGR (Q60445) Coatomer epsilon subunit (Epsilon-coat prote... 35 0.47
ITN1_XENLA (O42287) Intersectin 1 35 0.62
COPE_MOUSE (O89079) Coatomer epsilon subunit (Epsilon-coat prote... 35 0.62
YCF1_OENHO (Q9MTH5) Hypothetical 287.9 kDa protein ycf1 35 0.81
TRHY_SHEEP (P22793) Trichohyalin 35 0.81
NIN_HUMAN (Q8N4C6) Ninein (hNinein) 35 0.81
MYH6_RAT (P02563) Myosin heavy chain, cardiac muscle alpha isofo... 34 1.1
MYH1_HUMAN (P12882) Myosin heavy chain, skeletal muscle, adult 1... 34 1.1
FSH_DROME (P13709) Female sterile homeotic protein (Fragile-chor... 34 1.1
>YLE1_SCHPO (Q10451) Hypothetical protein C1093.01 in chromosome I
Length = 1261
Score = 52.0 bits (123), Expect = 5e-06
Identities = 56/242 (23%), Positives = 106/242 (43%), Gaps = 20/242 (8%)
Query: 342 LEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRK-------SSLKPNGATYGLAMEVMLQS 394
L P+V Y +++ +K++ V VF+ + SL+ L ++ M+ S
Sbjct: 805 LHPEV--YPTLISVLSKNKRFDAVQRVFEHSKHLYRKISTKSLEKANWFMALILDAMILS 862
Query: 395 GNYDLVHE---LF-EKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDME---RRGV 447
++ + LF + M+ G +P A T+ ++ + G D+A A+ E R V
Sbjct: 863 SSFARQFKSSNLFCDNMKMLGYIPRASTFAHLINNSTRRGDTDDATTALNIFEETKRHNV 922
Query: 448 MGTASVYYELACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCI 507
+ +Y + L R + +++K P VT+ +I ++ G
Sbjct: 923 KPSVFLYNAVLSKLGRARRTTECWKLFQEMKE-SGLLPTSVTYGTVINAACRIGDESLAE 981
Query: 508 CIFEYM--QDHCAPNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLML 565
+F M Q + P V NTM++ Q MF+ K LF ++ +D+ P ++TY L++
Sbjct: 982 KLFAEMENQPNYQPRVAPYNTMIQFEVQT-MFNREKALFYYNRLCATDIEPSSHTYKLLM 1040
Query: 566 EA 567
+A
Sbjct: 1041 DA 1042
Score = 37.0 bits (84), Expect = 0.16
Identities = 40/169 (23%), Positives = 73/169 (42%), Gaps = 24/169 (14%)
Query: 252 SKDHRKFQSRFVYTKLLAVLGKARRPKEALQIFNMMLGNIRVYPDMAAYHSI---AVTLG 308
+K H S F+Y +L+ LG+ARR E ++F M + + P Y ++ A +G
Sbjct: 917 TKRHNVKPSVFLYNAVLSKLGRARRTTECWKLFQEMKES-GLLPTSVTYGTVINAACRIG 975
Query: 309 QAGLLKELLNIVECMRQKPETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSK-QWKGVSW 367
L ++L +E P +P V YN ++ V + + +
Sbjct: 976 DESLAEKLFAEMEN----------------QPNYQPRVAPYNTMIQFEVQTMFNREKALF 1019
Query: 368 VFQQMRKSSLKPNGATYGLAMEV--MLQSGNYDLVHELFEKMQRNGEVP 414
+ ++ + ++P+ TY L M+ L+ N V + E M+R +VP
Sbjct: 1020 YYNRLCATDIEPSSHTYKLLMDAYGTLKPVNVGSVKAVLELMERT-DVP 1067
>RA50_AQUAE (O67124) Probable DNA double-strand break repair rad50
ATPase
Length = 978
Score = 42.4 bits (98), Expect = 0.004
Identities = 26/109 (23%), Positives = 55/109 (49%), Gaps = 9/109 (8%)
Query: 47 RKKQIQKHNRKLNRQNQLNPPLSKSQKQTLEEEQHFQELKHEYKQFTQNLEENQGGNKGL 106
R+K+++ +KL ++ LS+ E+E+ +++ K E++ ++ +E KG
Sbjct: 334 REKELEHRLKKLQEIKEILKELSQLSSSLKEKEREYEQAKQEFEDLSERVE------KGK 387
Query: 107 CLIGKPWEGVEKVDFLERIKVNYEHRGEKLKRESLIELKEMFRERKMDE 155
L+ + E +EK+ + + E+ K+K L+EL+ +E K E
Sbjct: 388 KLVAETEEKLEKI---KELFSEEEYTSLKMKERLLVELQRKLKELKEKE 433
Score = 35.8 bits (81), Expect = 0.36
Identities = 27/96 (28%), Positives = 49/96 (50%), Gaps = 6/96 (6%)
Query: 72 QKQTLEEEQHFQELKHEYKQFTQNLEENQGGNKGLCLIGKPWEGV-----EKVDFLERIK 126
+++ E E+ FQ LK + + + L+E +G + L I +E V EK L +K
Sbjct: 727 ERKIKEFEESFQSLKLKKSEIEEKLKEYEGIRE-LSDIKGEYESVKTQLEEKHKKLGEVK 785
Query: 127 VNYEHRGEKLKRESLIELKEMFRERKMDELKWVFED 162
EH GE+LKR+ ++ + E+K++ + + D
Sbjct: 786 RELEHLGERLKRKEELQKEISELEKKLEVYRVISND 821
>SYR_TREPA (O83803) Arginyl-tRNA synthetase (EC 6.1.1.19)
(Arginine--tRNA ligase) (ArgRS)
Length = 589
Score = 42.0 bits (97), Expect = 0.005
Identities = 47/188 (25%), Positives = 76/188 (40%), Gaps = 17/188 (9%)
Query: 79 EQHFQELKHEYKQFTQNLEENQGGNKGLCLIGKPWEGVEKVDFLERIKVNYEHRG---EK 135
+Q+ +E +H+ + Q E + L W L IK YE G +K
Sbjct: 215 QQYPEEAEHDVRDLLQRWESADPHVRALWRTMNEWA-------LRGIKQTYERTGISFDK 267
Query: 136 LKRESLIELKEMFRERKMDELKWVFEDDIEINEVWFDENNNGKRKKTSKRSEVQVVRFLV 195
L ES E RE L +E N +W D ++ G KK RS+ ++
Sbjct: 268 LYFES--ETYTKGREEVRRGLACGVFYQMEDNSIWVDLSSLGLDKKALLRSD-GTTMYIT 324
Query: 196 DRLCDKEIRAKDWKFSRLMKLSG--LSFTEGQLMMIIEMLGVKRCWKQALSVVQWVYNSK 253
+ RA+DW F +L+ + G ++ L ++ +LG W Q L V + +
Sbjct: 325 QDIGTAIFRAQDWPFDQLLYVVGNEQNYHFKVLFFVLRLLGYP--WAQQLHHVSYGMVNL 382
Query: 254 DHRKFQSR 261
H + +SR
Sbjct: 383 PHGRMKSR 390
>TRHY_HUMAN (Q07283) Trichohyalin
Length = 1898
Score = 39.3 bits (90), Expect = 0.033
Identities = 35/144 (24%), Positives = 63/144 (43%), Gaps = 17/144 (11%)
Query: 46 LRKKQIQKHNRKLNRQNQLN--PPLSKSQKQTLEEEQHFQELKHEYKQFTQNLEENQGGN 103
LR++Q + ++L R+ QL L + Q+ E+E+ E KHE ++ Q L+ Q
Sbjct: 414 LRREQQLRREQQLRREQQLRREQQLRREQQLRREQEEERHEQKHEQERREQRLKREQ--- 470
Query: 104 KGLCLIGKPWEGVEKVDFLERIKVNYEHRGEKLKRESLIELKEMFRERKMDELKWVFEDD 163
E+ D+L+R + H E+ K++ + +E RER + + +
Sbjct: 471 ------------EERRDWLKREEETERHEQERRKQQLKRDQEEERRERWLKLEEEERREQ 518
Query: 164 IEINEVWFDENNNGKRKKTSKRSE 187
E E +R++ KR E
Sbjct: 519 QERREQQLRREQEERREQRLKRQE 542
>MLP1_YEAST (Q02455) MLP1 protein (Myosin-like protein 1)
Length = 1875
Score = 38.5 bits (88), Expect = 0.056
Identities = 38/174 (21%), Positives = 79/174 (44%), Gaps = 25/174 (14%)
Query: 14 LRPLSQFQPDTDKIRRSL-----------------IQKGVTPTPKIIHTLRKKQIQKHNR 56
L + + Q D +K R L + + +T + I R+ Q Q
Sbjct: 1390 LEAIRKLQEDAEKASRELQAKLEESTTSYESTINGLNEEITTLKEEIEKQRQIQQQLQAT 1449
Query: 57 KLNRQNQLNPPLSKSQKQTLEEEQHFQELKHEYKQFTQNLEE-----NQGGNKGLCLIGK 111
N QN L+ + +S K++ EE++ + +K + ++ + + E NQ N + I K
Sbjct: 1450 SANEQNDLSN-IVESMKKSFEEDK-IKFIKEKTQEVNEKILEAQERLNQPSNINMEEIKK 1507
Query: 112 PWEGVEKVDFLERIKVNYEHRGEKLKRESLIELKEMFRERKMDELKWVFEDDIE 165
WE + + ++I+ E ++++ + ++ ++ ERK +EL+ FE+ +E
Sbjct: 1508 KWESEHEQEVSQKIREAEEALKKRIRLPTEEKINKII-ERKKEELEKEFEEKVE 1560
>YG3M_YEAST (P48237) Hypothetical 101.4 kDa protein in RPL24B-RSR1
intergenic region
Length = 864
Score = 37.7 bits (86), Expect = 0.095
Identities = 19/75 (25%), Positives = 39/75 (51%), Gaps = 5/75 (6%)
Query: 328 ETLKYMYRKNWDPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLA 387
+T+K++ K++ PD+ YN +L C + + +FQ+++ ++KP TY +
Sbjct: 344 DTMKFLSTKHF-----PDICTYNTMLRICEKERNFPKALDLFQEIQDHNIKPTTNTYIMM 398
Query: 388 MEVMLQSGNYDLVHE 402
V+ S + +V E
Sbjct: 399 ARVLASSSSNAVVSE 413
Score = 32.7 bits (73), Expect = 3.1
Identities = 17/66 (25%), Positives = 32/66 (47%)
Query: 519 PNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLEASSRGHQWEYFE 578
PN + T+++ YS+ +M A F+ +K + PD TYN ML + +
Sbjct: 318 PNKQNLTTVIQFYSRKEMTKQAWNTFDTMKFLSTKHFPDICTYNTMLRICEKERNFPKAL 377
Query: 579 HVYKEM 584
+++E+
Sbjct: 378 DLFQEI 383
Score = 31.2 bits (69), Expect = 8.9
Identities = 26/114 (22%), Positives = 49/114 (42%), Gaps = 18/114 (15%)
Query: 519 PNVGTVNTMLKVYSQNDMFSTAKVLFEEVKVAKSDLRPDAYTYNLMLE--ASSRGHQ--- 573
P++ T NTML++ + F A LF+E++ +++P TY +M ASS +
Sbjct: 355 PDICTYNTMLRICEKERNFPKALDLFQEIQ--DHNIKPTTNTYIMMARVLASSSSNAVVS 412
Query: 574 -----------WEYFEHVYKEMILSGYHLDQNKHLPLLVKASRAGKVDNILVIY 616
W+Y + + + D N L ++ A+ G ++ +Y
Sbjct: 413 EGKSDSLRLLGWKYLHELEDKNLYRHKKDDLNLFLAMMALAAFDGDIELSRALY 466
>MUKB_VIBVY (Q7MJ64) Chromosome partition protein mukB (Structural
maintenance of chromosome related protein)
Length = 1487
Score = 36.2 bits (82), Expect = 0.28
Identities = 29/146 (19%), Positives = 68/146 (45%), Gaps = 7/146 (4%)
Query: 24 TDKIRRSLIQKGVTPTPKIIHTLRKKQIQKHNRKLNRQNQLNPPLSKSQKQTLEEEQHFQ 83
T + R L + +T + + + + ++KL + L L S++ +E+ +
Sbjct: 251 TTQADRDLFKHLITESTNYVAADYMRHANERHKKLEQSLSLRSELFSSRETLIEQNRLLN 310
Query: 84 ELKHEYKQFTQN---LEEN-QGGNKGLCLIGKPWEGVEKVDFLERIKVNYEHRGEKLKRE 139
+++ E + ++ LE++ QG + L L+ EK+ ER + + E E+L+ +
Sbjct: 311 QVQQELEMLVESESALEQDYQGASDHLQLVQNALRQQEKI---ERYQEDLEELNERLEEQ 367
Query: 140 SLIELKEMFRERKMDELKWVFEDDIE 165
S++ + R +E V E++++
Sbjct: 368 SMVVEEAQERVLMAEEQSTVAENEVD 393
>MUKB_VIBVU (Q8DAP8) Chromosome partition protein mukB (Structural
maintenance of chromosome related protein)
Length = 1487
Score = 36.2 bits (82), Expect = 0.28
Identities = 29/146 (19%), Positives = 68/146 (45%), Gaps = 7/146 (4%)
Query: 24 TDKIRRSLIQKGVTPTPKIIHTLRKKQIQKHNRKLNRQNQLNPPLSKSQKQTLEEEQHFQ 83
T + R L + +T + + + + ++KL + L L S++ +E+ +
Sbjct: 251 TTQADRDLFKHLITESTNYVAADYMRHANERHKKLEQSLSLRSELFSSRETLIEQNRLLN 310
Query: 84 ELKHEYKQFTQN---LEEN-QGGNKGLCLIGKPWEGVEKVDFLERIKVNYEHRGEKLKRE 139
+++ E + ++ LE++ QG + L L+ EK+ ER + + E E+L+ +
Sbjct: 311 QVQQELEMLVESESALEQDYQGASDHLQLVQNALRQQEKI---ERYQEDLEELNERLEEQ 367
Query: 140 SLIELKEMFRERKMDELKWVFEDDIE 165
S++ + R +E V E++++
Sbjct: 368 SMVVEEAQERVLMAEEQSTVAENEVD 393
>COPE_HUMAN (O14579) Coatomer epsilon subunit (Epsilon-coat protein)
(Epsilon-COP)
Length = 307
Score = 36.2 bits (82), Expect = 0.28
Identities = 44/205 (21%), Positives = 88/205 (42%), Gaps = 13/205 (6%)
Query: 342 LEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVH 401
+E DV +Y A L +++ GV V +++ SS A A + +S +V
Sbjct: 49 VERDVFLYRAYL-----AQRKFGV--VLDEIKPSSAPELQAVRMFADYLAHESRRDSIVA 101
Query: 402 ELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMGTASVYYELACCL 461
EL +M R+ +V T+ +M + + + +A A+R + + + ++ ++ L
Sbjct: 102 ELDREMSRSVDVTNT-TFLLMAASIYLHDQNPDA--ALRALHQGDSLECTAMTVQI---L 155
Query: 462 CNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPNV 521
R A E+++++ L L T + + G + D IF+ M D C+P +
Sbjct: 156 LKLDRLDLARKELKRMQDLDEDATLTQLATAWVSLATGGEKLQDAYYIFQEMADKCSPTL 215
Query: 522 GTVNTMLKVYSQNDMFSTAKVLFEE 546
+N + + A+ L +E
Sbjct: 216 LLLNGQAACHMAQGRWEAAEGLLQE 240
>YCO2_ARATH (Q9LME2) Hypothetical protein At1g22260
Length = 871
Score = 35.8 bits (81), Expect = 0.36
Identities = 30/100 (30%), Positives = 48/100 (48%), Gaps = 6/100 (6%)
Query: 122 LERIKVNYEHRGEKLKRESLIELKEMFRERKMDELKWVFEDDIEINEVWFDENNN----- 176
L+ K YE +KL+ E ++ K+ R+R + +L+W D E + N N
Sbjct: 670 LKAFKCQYEDDCKKLQEELDLQRKKEERQRALVQLQWKVMSDNPPEEQEVNSNKNYSISK 729
Query: 177 GKRKKTSKRSEVQVVRFLVDRLCDKE-IRAKDWKFSRLMK 215
R SKRSE VR D + D ++AK+ S+++K
Sbjct: 730 DSRLGGSKRSEHIRVRSDNDNVQDSPFVKAKETPVSKILK 769
>DESP_HUMAN (P15924) Desmoplakin (DP) (250/210 kDa paraneoplastic
pemphigus antigen)
Length = 2871
Score = 35.4 bits (80), Expect = 0.47
Identities = 44/215 (20%), Positives = 83/215 (38%), Gaps = 38/215 (17%)
Query: 5 HMIQGHILPLRPLSQFQPDTDKIRRSLIQKGVTPTPKIIHTLRKKQIQKHNRKLNRQNQL 64
++ + H++ L + + D +RR + I+ + QI +NR L Q +
Sbjct: 1712 NLTKEHLMLEEELRNLRLEYDDLRRGRSEADSDKNATILELRSQLQIS-NNRTLELQGLI 1770
Query: 65 NP----------PLSKSQKQTLEEEQHFQELKHEYKQFTQNLEENQGGNKGLCLIGKPWE 114
N + K QKQ LE QE K++ Q Q E
Sbjct: 1771 NDLQRERENLRQEIEKFQKQALEASNRIQESKNQCTQVVQERESL--------------- 1815
Query: 115 GVEKVDFLERIKVNYEHRGEKLKR-ESLIELKEMFRER----------KMDELKWVFEDD 163
+ K+ LE+ K + ++L R +S +E + ++R +++ K +
Sbjct: 1816 -LVKIKVLEQDKARLQRLEDELNRAKSTLEAETRVKQRLECEKQQIQNDLNQWKTQYSRK 1874
Query: 164 IEINEVWFDENNNGKRKKTSKRSEVQVVRFLVDRL 198
E E +R+K S RSE++ ++ + R+
Sbjct: 1875 EEAIRKIESEREKSEREKNSLRSEIERLQAEIKRI 1909
>COPE_CRIGR (Q60445) Coatomer epsilon subunit (Epsilon-coat protein)
(Epsilon-COP) (LDLF)
Length = 307
Score = 35.4 bits (80), Expect = 0.47
Identities = 41/209 (19%), Positives = 86/209 (40%), Gaps = 15/209 (7%)
Query: 339 DPTLEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYD 398
D +E DV +Y A + +++ GV V +++ SS A A + ++
Sbjct: 46 DREVERDVFLYRAYI-----AQRKYGV--VLDEIKPSSAPELQAVRMFADYLATENRRDA 98
Query: 399 LVHELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMG-TASVYYEL 457
+V EL +M R+ +V + ++ + D A++ + + M T + +L
Sbjct: 99 IVVELDREMSRSVDVTNTTFLLMAASVYFHDQNPDAALRTLHQGDSLECMAMTIQILLKL 158
Query: 458 ACCLCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHC 517
R A E++K++ L T + ++ G + + IF+ + D C
Sbjct: 159 -------DRLDLARKELKKMQDQDEDATLTQLATAWVNLAVGGEKLQEAYYIFQELADKC 211
Query: 518 APNVGTVNTMLKVYSQNDMFSTAKVLFEE 546
+P + +N +S + TA+ + +E
Sbjct: 212 SPTLLLLNGQAACHSAQGRWETAEGVLQE 240
>ITN1_XENLA (O42287) Intersectin 1
Length = 1270
Score = 35.0 bits (79), Expect = 0.62
Identities = 45/206 (21%), Positives = 93/206 (44%), Gaps = 17/206 (8%)
Query: 19 QFQPDTDKIRRSLIQKGVTPTPKIIHTLRKKQIQKHNRKLN-RQNQLNPPLSK------S 71
Q + + ++ + L Q+ ++ +KK ++ LN +++QL L +
Sbjct: 444 QLEWERNRRQELLNQRNREQEDIVVLKAKKKTLEFELEALNDKKHQLEGKLQDIRCRLTT 503
Query: 72 QKQTLEEEQHFQELK-HEYKQFTQNLEENQGGNKGLCLIGKPW-EGVEKVDFLERIKVNY 129
Q+ +E +EL+ E Q L+E+Q L+GK E +D L++++ N
Sbjct: 504 QRHEIESTNKSRELRIAEITHLQQQLQESQQ------LLGKMIPEKQSLIDQLKQVQQNS 557
Query: 130 EHRGEKLKRESLIELKEMFRERKMDELKWV-FEDDIEINEVWFDENNNGKRKKTSKRSEV 188
HR L + +E KE+ R++ D+L V E ++ E+ N + ++ + +
Sbjct: 558 LHRDSLLTLKRALETKEIGRQQLRDQLDEVEKETRAKLQEIDVFNNQLKELRELYNKQQF 617
Query: 189 QVVR-FLVDRLCDKEIRAKDWKFSRL 213
Q + F +++ KE+ K + +L
Sbjct: 618 QKQQDFETEKIKQKELERKTSELDKL 643
>COPE_MOUSE (O89079) Coatomer epsilon subunit (Epsilon-coat protein)
(Epsilon-COP)
Length = 307
Score = 35.0 bits (79), Expect = 0.62
Identities = 41/206 (19%), Positives = 85/206 (40%), Gaps = 15/206 (7%)
Query: 342 LEPDVVIYNAVLNACVPSKQWKGVSWVFQQMRKSSLKPNGATYGLAMEVMLQSGNYDLVH 401
+E DV +Y A L +++ GV V +++ SS A A + ++ +V
Sbjct: 49 VERDVFLYRAYL-----AQRKYGV--VLDEIKPSSAPELQAVRMFAEYLASENQRDSIVL 101
Query: 402 ELFEKMQRNGEVPEALTYKVMVRTFWKEGKVDEAVKAVRDMERRGVMG-TASVYYELACC 460
EL +M R+ +V + ++ + D A++ + + M T + +L
Sbjct: 102 ELDREMSRSVDVTNTTFLLMAASIYFHDQNPDAALRTLHQGDGLECMAMTIQILLKL--- 158
Query: 461 LCNCGRWQDATLEVEKIKRLPHAKPLEVTFTGMIRSSMDGGHIDDCICIFEYMQDHCAPN 520
R A E++K++ L T + ++ G + + IF+ + D C+P
Sbjct: 159 ----DRLDLARKELKKMQDQDEDATLTQLATAWVNLAVGGEKLQEAYYIFQELADKCSPT 214
Query: 521 VGTVNTMLKVYSQNDMFSTAKVLFEE 546
+ +N +S + TA+ + +E
Sbjct: 215 LLLLNGQAACHSAQGRWETAEGVLQE 240
>YCF1_OENHO (Q9MTH5) Hypothetical 287.9 kDa protein ycf1
Length = 2434
Score = 34.7 bits (78), Expect = 0.81
Identities = 28/121 (23%), Positives = 60/121 (49%), Gaps = 17/121 (14%)
Query: 41 KIIHTLRKKQIQKHNRKLNRQNQLNPPLSKSQK-------------QTLEEEQHFQELKH 87
++I R+K+I+KH RK+ +Q + + ++K + LE+E+ +E K
Sbjct: 2043 QLIEKKRQKEIEKHKRKIEKQMRKKEKIENAKKKIENEKKKIETEEEKLEKEKRKKERKK 2102
Query: 88 E--YKQFTQNLEENQGGNKGLCLIGKPWEGVEKVDFLERIKVNYEHRGEKLKRESLIELK 145
E K+ +N+E+ + NK + K E ++K ++ E + + KR+ ++++
Sbjct: 2103 EKLKKKVAKNIEKLK--NKVAKNVAKNIEKLKKQRAKNIARLEEEDKKARKKRKRKVQVQ 2160
Query: 146 E 146
E
Sbjct: 2161 E 2161
>TRHY_SHEEP (P22793) Trichohyalin
Length = 1549
Score = 34.7 bits (78), Expect = 0.81
Identities = 40/179 (22%), Positives = 80/179 (44%), Gaps = 25/179 (13%)
Query: 48 KKQIQKHNRKLNRQNQLNPPLSKSQKQTLEEEQHFQELKHEYKQFTQNLEENQGGNK-GL 106
++Q+QK R L + L + + + E+E+ Q+L+ E ++ TQ +G ++
Sbjct: 238 EQQLQKQKRGLE-ERLLEQERREQELRRKEQERREQQLRQEQEEATQEEISERGESRTSR 296
Query: 107 CLIGKPWEGVEKVDFLERIKVNYEHRGEKLKRESLIELKEMFRERKMDELKWVFEDD--- 163
C W+ + D +R + HR E+ R EL E +E+++ E ++D
Sbjct: 297 C----QWQLESEADARQRKVYSRPHRQEQQSRRQEQELLERQQEQQISEEVQSLQEDQGR 352
Query: 164 --------IEINEVWFDENNNGKRKKT--SKRSEVQVVRFLVDRLCDKEIRAKDWKFSR 212
+ N W E + +R+ T +K ++ + VR ++++R K+ K R
Sbjct: 353 QRLKQEQRYDQNWRWQLEEESQRRRYTLYAKPAQREQVRE------EEQLRLKEEKLQR 405
>NIN_HUMAN (Q8N4C6) Ninein (hNinein)
Length = 2090
Score = 34.7 bits (78), Expect = 0.81
Identities = 37/123 (30%), Positives = 53/123 (43%), Gaps = 21/123 (17%)
Query: 46 LRKKQIQKHNRKLN------------RQNQLNPPL-SKSQKQTLEEEQHFQELKHEYKQF 92
LRK+ ++KH R+L R +Q+ S QK T E Q L+ Y+Q
Sbjct: 784 LRKELLEKHQRELQEGREKMETECNRRTSQIEAQFQSDCQKVTERCESALQSLEGRYRQE 843
Query: 93 TQNLEENQGGNKGLCLIGKPWEGVEKVDFLERIKVNYEHRGEKLKRESLIELKEMFRERK 152
++L+E Q K WE EK + + E E LKRE L + +ER+
Sbjct: 844 LKDLQEQQREEK------SQWE-FEKDELTQECAEAQELLKETLKREKTTSL-VLTQERE 895
Query: 153 MDE 155
M E
Sbjct: 896 MLE 898
>MYH6_RAT (P02563) Myosin heavy chain, cardiac muscle alpha isoform
(MyHC-alpha)
Length = 1938
Score = 34.3 bits (77), Expect = 1.1
Identities = 41/160 (25%), Positives = 72/160 (44%), Gaps = 15/160 (9%)
Query: 68 LSKSQKQTLEEEQHFQELKHEYKQFTQNLEENQGGNKGL-----CLIGKPWEGVEKVDFL 122
L SQK+ +LK+ Y++ ++LE + NK L L + EG + V L
Sbjct: 1468 LESSQKEARSLSTELFKLKNAYEESLEHLETFKRENKNLQEEISDLTEQLGEGGKNVHEL 1527
Query: 123 ERIKVNYEHRGEKLKRESLIELKEMFRERKMDELKWVFEDDIEINEVWFDENNNGKRK-- 180
E+I+ E EKL+ +S +E E E + + + +E N++ + K
Sbjct: 1528 EKIRKQLE--VEKLELQSALEEAEASLEHEEGK---ILRAQLEFNQIKAEIERKLAEKDE 1582
Query: 181 --KTSKRSEVQVVRFLVDRLCDKEIRAKDWKFSRLMKLSG 218
+ +KR+ ++VV L L D E R+++ K+ G
Sbjct: 1583 EMEQAKRNHLRVVDSLQTSL-DAETRSRNEALRVKKKMEG 1621
>MYH1_HUMAN (P12882) Myosin heavy chain, skeletal muscle, adult 1
(Myosin heavy chain IIx/d) (MyHC-IIx/d)
Length = 1939
Score = 34.3 bits (77), Expect = 1.1
Identities = 38/160 (23%), Positives = 72/160 (44%), Gaps = 15/160 (9%)
Query: 68 LSKSQKQTLEEEQHFQELKHEYKQFTQNLEENQGGNKGL-----CLIGKPWEGVEKVDFL 122
L SQK++ ++K+ Y++ LE + NK L L + EG +++ L
Sbjct: 1471 LEASQKESRSLSTELFKIKNAYEESLDQLETLKRENKNLQQEISDLTEQIAEGGKRIHEL 1530
Query: 123 ERIKVNYEHRGEKLKRESLIELKEMFRERKMDELKWVFEDDIEINEVWFDENNNGKRKKT 182
E+IK E EK + ++ +E E E + + + +E+N+V + + K
Sbjct: 1531 EKIKKQVEQ--EKSELQAALEEAEASLEHEEGK---ILRIQLELNQVKSEVDRKIAEKDE 1585
Query: 183 S----KRSEVQVVRFLVDRLCDKEIRAKDWKFSRLMKLSG 218
KR+ +++V + L D EIR+++ K+ G
Sbjct: 1586 EIDQMKRNHIRIVESMQSTL-DAEIRSRNDAIRLKKKMEG 1624
>FSH_DROME (P13709) Female sterile homeotic protein (Fragile-chorion
membrane protein)
Length = 2038
Score = 34.3 bits (77), Expect = 1.1
Identities = 30/116 (25%), Positives = 57/116 (48%), Gaps = 15/116 (12%)
Query: 2 EALHMIQGHILPLRP-LSQFQPDTDKIRRSLIQKGVTPTPKIIHTLRKKQIQKHNRKLNR 60
+ L+M L L P L Q Q ++++ Q+ T H +++ Q+H+++ +
Sbjct: 1482 DGLNMTAASFLDLEPSLQQQQMQQMQLQQQHHQQQQQQT----HQQQQQHQQQHHQQ--Q 1535
Query: 61 QNQLNPPLSKSQKQTLEEEQHFQELKHEYKQFTQNLEENQGGNKGLCLIGKPWEGV 116
Q QL + Q+Q +++QH Q+ +H+ + +Q NK L +I KP E +
Sbjct: 1536 QQQLTQQQLQQQQQQQQQQQHLQQQQHQQ-------QHHQAANK-LLIIPKPIESM 1583
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.136 0.408
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 76,209,770
Number of Sequences: 164201
Number of extensions: 3239177
Number of successful extensions: 9830
Number of sequences better than 10.0: 77
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 70
Number of HSP's that attempted gapping in prelim test: 9710
Number of HSP's gapped (non-prelim): 191
length of query: 653
length of database: 59,974,054
effective HSP length: 117
effective length of query: 536
effective length of database: 40,762,537
effective search space: 21848719832
effective search space used: 21848719832
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 69 (31.2 bits)
Medicago: description of AC140720.4


