

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140670.14 + phase: 0
(359 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
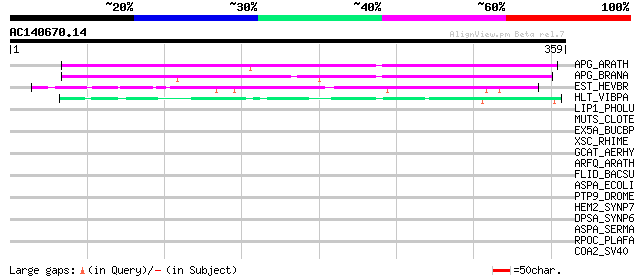
Score E
Sequences producing significant alignments: (bits) Value
APG_ARATH (P40602) Anter-specific proline-rich protein APG precu... 225 2e-58
APG_BRANA (P40603) Anter-specific proline-rich protein APG (Prot... 223 6e-58
EST_HEVBR (Q7Y1X1) Esterase precursor (EC 3.1.1.-) (Early nodule... 119 2e-26
HLT_VIBPA (Q99289) Thermolabile hemolysin precursor (TL) (Lecith... 51 5e-06
LIP1_PHOLU (P40601) Lipase 1 precursor (EC 3.1.1.3) (Triacylglyc... 35 0.38
MUTS_CLOTE (Q895H2) DNA mismatch repair protein mutS 34 0.65
EX5A_BUCBP (Q89AB2) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 32 1.9
XSC_RHIME (Q92UW6) Probable sulfoacetaldehyde acetyltransferase ... 32 2.5
GCAT_AERHY (P10480) Phosphatidylcholine-sterol acyltransferase p... 32 2.5
ARFQ_ARATH (Q84WU6) Auxin response factor 17 32 2.5
FLID_BACSU (P39738) Flagellar hook-associated protein 2 (HAP2) (... 32 3.2
ASPA_ECOLI (P04422) Aspartate ammonia-lyase (EC 4.3.1.1) (Aspart... 31 4.2
PTP9_DROME (P35832) Protein-tyrosine phosphatase 99A precursor (... 30 7.2
HEM2_SYNP7 (P43087) Delta-aminolevulinic acid dehydratase (EC 4.... 30 7.2
DPSA_SYNP6 (Q9R6T3) Nutrient-stress induced DNA binding protein ... 30 7.2
ASPA_SERMA (P33109) Aspartate ammonia-lyase (EC 4.3.1.1) (Aspart... 30 7.2
RPOC_PLAFA (P21422) DNA-directed RNA polymerase beta' chain (EC ... 30 9.4
COA2_SV40 (P03093) Coat protein VP2 [Contains: Coat protein VP3] 30 9.4
>APG_ARATH (P40602) Anter-specific proline-rich protein APG
precursor
Length = 534
Score = 225 bits (573), Expect = 2e-58
Identities = 113/331 (34%), Positives = 179/331 (53%), Gaps = 13/331 (3%)
Query: 34 VPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGI 93
+PA+ FGDS D GNNN + T +SN++PYG DF TGRFSNG +A+D++++ G+
Sbjct: 202 IPAVFFFGDSVFDTGNNNNLETKIKSNYRPYGMDFKFRVATGRFSNGMVASDYLAKYMGV 261
Query: 94 KPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAY 153
K +PAYLDP + TGVSFAS GY+ TS+ + IP+ QL Y+++Y +K+
Sbjct: 262 KEIVPAYLDPKIQPNDLLTGVSFASGGAGYNPTTSEAANAIPMLDQLTYFQDYIEKVNRL 321
Query: 154 L----------GEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGI 203
+ G +K + I+K + I+ G+ND + Y+ + + Y +A
Sbjct: 322 VRQEKSQYKLAGLEKTNQLISKGVAIVVGGSNDLIITYFGSGAQRLKNDIDSYTTIIADS 381
Query: 204 AQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLT 263
A +F+ +LY GA++I + G PP+GC+P +R C N + FN KL +
Sbjct: 382 AASFVLQLYGYGARRIGVIGTPPLGCVPSQRLKK---KKICNEELNYASQLFNSKLLLIL 438
Query: 264 TKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSC 323
+L K LP V+ + Y ++ +++ P YGF+ CC TG+ G C +++ C
Sbjct: 439 GQLSKTLPNSTFVYMDIYTIISQMLETPAAYGFEETKKPCCKTGLLSAGALCKKSTSKIC 498
Query: 324 MDASRYVFWDSFHPTEKTNGIVANYLVKNAL 354
+ S Y+FWD HPT++ + L+K L
Sbjct: 499 PNTSSYLFWDGVHPTQRAYKTINKVLIKEYL 529
>APG_BRANA (P40603) Anter-specific proline-rich protein APG (Protein
CEX) (Fragment)
Length = 449
Score = 223 bits (568), Expect = 6e-58
Identities = 118/327 (36%), Positives = 182/327 (55%), Gaps = 15/327 (4%)
Query: 34 VPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFGI 93
+PA+ FGDS D GNNN + T + N++PYG DF G TGRFSNGR+A+D+IS+ G+
Sbjct: 123 IPAVFFFGDSIFDTGNNNNLDTKLKCNYRPYGMDFPMGVATGRFSNGRVASDYISKYLGV 182
Query: 94 KPYIPAYLDPSFNI------SQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQ 147
K +PAY+D S TGVSFAS GY TS+ V + QL Y+++Y+
Sbjct: 183 KEIVPAYVDKKLQQNNELQQSDLLTGVSFASGGAGYLPQTSESWKVTTMLDQLTYFQDYK 242
Query: 148 KKLGAYLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNF---LAGIA 204
K++ +G+KK K+ ++K I+ G+ND + Y+ G +Q+ ++ +F +A A
Sbjct: 243 KRMKKLVGKKKTKKIVSKGAAIVVAGSNDLIYTYF---GNGAQHLKNDVDSFTTMMADSA 299
Query: 205 QNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTT 264
+F+ +LY GA++I + G PP+GC P +R C + N A FN KL +
Sbjct: 300 ASFVLQLYGYGARRIGVIGTPPIGCTPSQRVKK---KKICNEDLNYAAQLFNSKLVIILG 356
Query: 265 KLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCM 324
+L K LP +V+ + Y + +++ P YGF+ CC G+ + G C +L +
Sbjct: 357 QLSKTLPNSTIVYGDIYSIFSKMLESPEDYGFEEIKKPCCKIGLTKGGVFCKERTLKNMS 416
Query: 325 DASRYVFWDSFHPTEKTNGIVANYLVK 351
+AS Y+FWD HP+++ I LVK
Sbjct: 417 NASSYLFWDGLHPSQRAYEISNRKLVK 443
>EST_HEVBR (Q7Y1X1) Esterase precursor (EC 3.1.1.-) (Early
nodule-specific protein homolog) (Latex allergen Hev b
13)
Length = 391
Score = 119 bits (297), Expect = 2e-26
Identities = 103/356 (28%), Positives = 156/356 (42%), Gaps = 44/356 (12%)
Query: 15 ILCILLLQRLDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPT 74
+LC+L L ++ PAI FGDS+ D G + PYG F + T
Sbjct: 17 LLCMLSLAYAS----ETCDFPAIFNFGDSNSDTGGK---AAAFYPLNPPYGETFFH-RST 68
Query: 75 GRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLS-- 132
GR+S+GR+ DFI+E+F + PY+ YL S S F G FA+A + T+ + +
Sbjct: 69 GRYSDGRLIIDFIAESFNL-PYLSPYL--SSLGSNFKHGADFATAGSTIKLPTTIIPAHG 125
Query: 133 -VIPLWKQLEYYK--------EYQKKLGAYLGEKKAKET-ITKALYIISLGTNDFLENYY 182
P + ++Y + ++ ++ G E +E KALY +G ND E +
Sbjct: 126 GFSPFYLDVQYSQFRQFIPRSQFIRETGGIFAELVPEEYYFEKALYTFDIGQNDLTEGFL 185
Query: 183 TIPGRASQYT-PSEYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGG 241
+ T P +F A + K+YDLGA+ + P+GCL T
Sbjct: 186 NLTVEEVNATVPDLVNSFSANVK-----KIYDLGARTFWIHNTGPIGCLSFILTYFPWAE 240
Query: 242 ND---CVSNYNNIALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLGVVKKPGQYGFQV 298
D C YN +A FN KL ++ +L+KDLP V + Y V + +P ++GF+
Sbjct: 241 KDSAGCAKAYNEVAQHFNHKLKEIVAQLRKDLPLATFVHVDIYSVKYSLFSEPEKHGFEF 300
Query: 299 ASMACCATG---MFEMGYAC---------SRASLFSCMDASRYVFWDSFHPTEKTN 342
+ CC G F + C ++ + SC S V WD H TE N
Sbjct: 301 PLITCCGYGGKYNFSVTAPCGDTVTADDGTKIVVGSCACPSVRVNWDGAHYTEAAN 356
>HLT_VIBPA (Q99289) Thermolabile hemolysin precursor (TL)
(Lecithin-dependent haemolysin) (LDH) (Atypical
phospholipase) (Phospholipase A2) (Lysophospholipase)
Length = 418
Score = 50.8 bits (120), Expect = 5e-06
Identities = 71/330 (21%), Positives = 123/330 (36%), Gaps = 61/330 (18%)
Query: 33 KVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRIATDFISEAFG 92
++ ++ GDS D GN I ++ F FLG FSNG + T++I++A
Sbjct: 143 QINKVVALGDSLSDTGN---IFNASQWRFPNPNSWFLG-----HFSNGFVWTEYIAKAKN 194
Query: 93 IKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGA 152
+ Y + A G + +++ + +Q+ Y Y K
Sbjct: 195 LPLY---------------------NWAVGGAAGENQYIALTGVGEQVSSYLTYAKLAKN 233
Query: 153 YLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLY 212
Y K T L+ + G NDF+ +P + Y + + +L
Sbjct: 234 Y----KPANT----LFTLEFGLNDFMNYNRGVPEVKADYAEA-------------LIRLT 272
Query: 213 DLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFNGKLNKLTTKLKKDLPG 272
D GAK L LP P + + + + LE N + K G
Sbjct: 273 DAGAKNFMLMTLPDATKAPQFKYST----QEEIDKIRAKVLEMNEFIKAQAMYYKAQ--G 326
Query: 273 IRLVFSNPYDVLLGVVKKPGQYGFQVASMACC---ATGMFEMGYACSRASLFSCMDASRY 329
+ + + + + P ++GF AS C + + Y + S + A ++
Sbjct: 327 YNITLFDTHALFETLTSAPEEHGFVNASDPCLDINRSSSVDYMYTHALRSECAASGAEKF 386
Query: 330 VFWDSFHPTEKTNGIVANYLVK--NALAQF 357
VFWD HPT T+ VA +++ N LA++
Sbjct: 387 VFWDVTHPTTATHRYVAEKMLESSNNLAEY 416
>LIP1_PHOLU (P40601) Lipase 1 precursor (EC 3.1.1.3)
(Triacylglycerol lipase)
Length = 645
Score = 34.7 bits (78), Expect = 0.38
Identities = 24/65 (36%), Positives = 34/65 (51%), Gaps = 5/65 (7%)
Query: 287 VVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASR-YVFWDSFHPTEKTNGIV 345
V+ P +YGF CA G+ A S S + DAS+ ++F D FHPT + + IV
Sbjct: 284 VIANPLRYGFLNTIGYACAQGV----NAGSCRSKDTGFDASKPFLFADDFHPTPEAHHIV 339
Query: 346 ANYLV 350
+ Y V
Sbjct: 340 SQYTV 344
>MUTS_CLOTE (Q895H2) DNA mismatch repair protein mutS
Length = 881
Score = 33.9 bits (76), Expect = 0.65
Identities = 35/169 (20%), Positives = 65/169 (37%), Gaps = 23/169 (13%)
Query: 104 SFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAYLGEKKAKETI 163
S I + + F +TG ATS L + ++ + + L + + KE
Sbjct: 128 SLYIEDKSVAMCFCDISTGDFFATSHSLDKSIILDEISKFDPSELLLNKDIDKNILKEIK 187
Query: 164 TKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYDLGAKKISLGG 223
+ I+ D+ ++Y + G+ S ++ EY L + Y L +K+SLG
Sbjct: 188 NRFTISITNLEKDYFKDYNLLEGQFSDFSKEEYSLVLIKCGNGLLK--YILETQKMSLGH 245
Query: 224 LPPMGCLPLERTTNFAGGNDCVSNYN---NIALEFNGKLNKLTTKLKKD 269
+ DC NYN ++++ N + N T+ +D
Sbjct: 246 I------------------DCFKNYNIVDYLSMDINSRRNLEITETIRD 276
>EX5A_BUCBP (Q89AB2) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5)
Length = 618
Score = 32.3 bits (72), Expect = 1.9
Identities = 18/58 (31%), Positives = 31/58 (53%), Gaps = 3/58 (5%)
Query: 246 SNYNNIALEFNGKLNKLTTKLKKDLPGIRLV-FSNPYDVLLGVVKKPGQYGFQVASMA 302
+NYNNI E +K + LK+ LP ++++ N Y +L + K G +G ++A
Sbjct: 407 TNYNNIIFEITNSYSKYWSALKQSLPFLKIINIFNNYKIL--CILKNGLFGITELNLA 462
>XSC_RHIME (Q92UW6) Probable sulfoacetaldehyde acetyltransferase (EC
2.3.3.15)
Length = 591
Score = 32.0 bits (71), Expect = 2.5
Identities = 17/43 (39%), Positives = 24/43 (55%), Gaps = 2/43 (4%)
Query: 153 YLGEKKAKETITKALYIISLGTNDFLENYYTIPGRASQYTPSE 195
Y G K A E I+KA +++LGT L + T+PG Y P +
Sbjct: 251 YNGSKAAMELISKADVVLALGTR--LNPFSTLPGYGIDYWPKD 291
>GCAT_AERHY (P10480) Phosphatidylcholine-sterol acyltransferase
precursor (EC 2.3.1.43) (Glycerophospholipid-cholesterol
acyltransferase) (GCAT)
Length = 335
Score = 32.0 bits (71), Expect = 2.5
Identities = 75/348 (21%), Positives = 125/348 (35%), Gaps = 75/348 (21%)
Query: 16 LCILLLQRLDLSLVKSAKV-PAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPT 74
+C+L L L + S I++FGDS D G R +L P
Sbjct: 6 VCLLGLVALTVQAADSRPAFSRIVMFGDSLSDTGK-----------MYSKMRGYLPSSPP 54
Query: 75 ---GRFSNGRIATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVL 131
GRFSNG + + ++ F G++ A+ A G A +
Sbjct: 55 YYEGRFSNGPVWLEQLTNEF--------------------PGLTIANEAEGGPTAVA--- 91
Query: 132 SVIPLWKQLEYYKEYQ--KKLGAYLGEKKAKETITKA-LYIISLGTNDFLENYYTIPGRA 188
+ ++ + +YQ L + + K++ L I+ +G ND+L G
Sbjct: 92 -----YNKISWNPKYQVINNLDYEVTQFLQKDSFKPDDLVILWVGANDYLAY-----GWN 141
Query: 189 SQYTPSEYQNFLAGIAQNFIHKLYDLGAKKISLGGLPPMGCLPLERTTNFAGGNDCVSNY 248
++ ++ ++ A + GAK+I L LP +G P R+ VS Y
Sbjct: 142 TEQDAKRVRDAISDAANRMVLN----GAKEILLFNLPDLGQNPSARSQKVVEAASHVSAY 197
Query: 249 NNIALEFNGKLNKLTTKLKKDLPGIRLVFSNPYDVLLG-VVKKPGQYGFQVASMACCATG 307
+ N+L L + L +V D +++ P +G AC
Sbjct: 198 H----------NQLLLNLARQLAPTGMVKLFEIDKQFAEMLRDPQNFGLSDQRNACYGGS 247
Query: 308 MFEMGYACSRASLFSCMDASRYVFWDSFHPTEKTNGIVANYLVKNALA 355
+A AS S + A F+P E+ I N L+ A+A
Sbjct: 248 YVWKPFASRSASTDSQLSA--------FNPQERL-AIAGNPLLAQAVA 286
>ARFQ_ARATH (Q84WU6) Auxin response factor 17
Length = 585
Score = 32.0 bits (71), Expect = 2.5
Identities = 18/54 (33%), Positives = 26/54 (47%), Gaps = 3/54 (5%)
Query: 206 NFIHKLYDLG---AKKISLGGLPPMGCLPLERTTNFAGGNDCVSNYNNIALEFN 256
NF+ L DLG + ++ G P P TTN + GND V N ++ + N
Sbjct: 469 NFLSPLPDLGKVSTEMMNFGSPPSDNLSPNSNTTNLSSGNDLVGNRGPLSKKVN 522
>FLID_BACSU (P39738) Flagellar hook-associated protein 2 (HAP2)
(Filament cap protein) (Flagellar cap protein)
Length = 498
Score = 31.6 bits (70), Expect = 3.2
Identities = 28/94 (29%), Positives = 45/94 (47%), Gaps = 14/94 (14%)
Query: 97 IPAYLDPSFNISQFATGVSFASAATG-------YDNATSDVLSVIPLWKQLEYYKEYQKK 149
+ A+ D +N +++ ++F+S ATG D+AT+D +S QL + + K
Sbjct: 164 VSAFKDKIWNGTEYVETIAFSSKATGAGGSIQAADSATADFMS-----GQLGFSLDADNK 218
Query: 150 LGAYLGEKKAKETITKALYIISLGTNDFLENYYT 183
L AY AK TI + + TN+F N T
Sbjct: 219 LTAYKEGTNAKVTING--FEMEKLTNNFTVNGVT 250
>ASPA_ECOLI (P04422) Aspartate ammonia-lyase (EC 4.3.1.1)
(Aspartase)
Length = 478
Score = 31.2 bits (69), Expect = 4.2
Identities = 32/125 (25%), Positives = 52/125 (41%)
Query: 23 RLDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRI 82
R++ L+ + +VPA +G ++ A N +IS S+ + R + K +N +
Sbjct: 6 RIEEDLLGTREVPADAYYGVHTLRAIENFYISNNKISDIPEFVRGMVMVKKAAAMANKEL 65
Query: 83 ATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEY 142
T S A I L+ + QF V A T + T++VL+ I L
Sbjct: 66 QTIPKSVANAIIAACDEVLNNGKCMDQFPVDVYQGGAGTSVNMNTNEVLANIGLELMGHQ 125
Query: 143 YKEYQ 147
EYQ
Sbjct: 126 KGEYQ 130
>PTP9_DROME (P35832) Protein-tyrosine phosphatase 99A precursor (EC
3.1.3.48) (Receptor-linked protein-tyrosine phosphatase
99A)
Length = 1301
Score = 30.4 bits (67), Expect = 7.2
Identities = 21/82 (25%), Positives = 34/82 (40%), Gaps = 3/82 (3%)
Query: 269 DLPGIRLVFSNPYDVLLGVVKKPGQYGFQVASMACCATGMFEMGYACSRASLFSCMDASR 328
D+P L F+N + V + G + A+ A + Y S+ + +D S
Sbjct: 128 DVPATTLAFANAFPVPVAGEMGNGNGNYNDATPPYAAV---DDNYVPSKPQNLTILDVSA 184
Query: 329 YVFWDSFHPTEKTNGIVANYLV 350
S+HP + NG +A Y V
Sbjct: 185 NSITMSWHPPKNQNGAIAGYHV 206
>HEM2_SYNP7 (P43087) Delta-aminolevulinic acid dehydratase (EC
4.2.1.24) (Porphobilinogen synthase) (ALAD) (ALADH)
Length = 326
Score = 30.4 bits (67), Expect = 7.2
Identities = 21/72 (29%), Positives = 37/72 (51%), Gaps = 5/72 (6%)
Query: 158 KAKETITKALYIISLGTNDFLENYYTIPGR--ASQYT--PSEYQNFLAGIAQNFIHKLYD 213
++ ET+ + + +L + DF+ + +PG A + T P YQ + I + ++YD
Sbjct: 11 RSSETLRRMVRETTLTSADFIYPLFAVPGEGVAKEVTSMPGVYQLSIDKIVEE-AKEVYD 69
Query: 214 LGAKKISLGGLP 225
LG I L G+P
Sbjct: 70 LGIPSIILFGIP 81
>DPSA_SYNP6 (Q9R6T3) Nutrient-stress induced DNA binding protein
(Fragment)
Length = 175
Score = 30.4 bits (67), Expect = 7.2
Identities = 25/87 (28%), Positives = 41/87 (46%), Gaps = 4/87 (4%)
Query: 161 ETITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYDLGAKKIS 220
E + +AL + + ++++T+ G A Y+ E+ G ++ +H DLG +
Sbjct: 30 EGLNRALASFQVLYLQYQKHHFTVQG-AEFYSLHEFFEDSYGSVKDHVH---DLGERLNG 85
Query: 221 LGGLPPMGCLPLERTTNFAGGNDCVSN 247
LGG+P L L T FA D V N
Sbjct: 86 LGGVPVAHPLKLAELTCFAIEEDGVFN 112
>ASPA_SERMA (P33109) Aspartate ammonia-lyase (EC 4.3.1.1)
(Aspartase)
Length = 478
Score = 30.4 bits (67), Expect = 7.2
Identities = 32/125 (25%), Positives = 51/125 (40%)
Query: 23 RLDLSLVKSAKVPAIIVFGDSSVDAGNNNFISTVARSNFQPYGRDFLGGKPTGRFSNGRI 82
R++ L+ + +VPA +G ++ A N +IS S+ + R + K +N +
Sbjct: 6 RIEEDLLGTREVPADAYYGVHTLRAIENFYISNSKISDVPEFVRGMVMVKKAAAMANKEL 65
Query: 83 ATDFISEAFGIKPYIPAYLDPSFNISQFATGVSFASAATGYDNATSDVLSVIPLWKQLEY 142
T A I LD + QF V A T + T++VL+ I L
Sbjct: 66 KTIPRKIADVIIQACDEVLDKGKCMDQFPVDVFQGGAGTSLNMNTNEVLANIGLELMGHQ 125
Query: 143 YKEYQ 147
EYQ
Sbjct: 126 KGEYQ 130
>RPOC_PLAFA (P21422) DNA-directed RNA polymerase beta' chain (EC
2.7.7.6)
Length = 575
Score = 30.0 bits (66), Expect = 9.4
Identities = 23/78 (29%), Positives = 33/78 (41%), Gaps = 2/78 (2%)
Query: 163 ITKALYIISLGTNDFLENYYTIPGRASQYTPSEYQNFLAGIAQNFIHKLYDLGAKKIS-- 220
I K L+II N L+N + I RA Q+F + + + K Y LG +
Sbjct: 399 INKNLFIIQKFLNRLLQNQFIIINRAPTLHRMNLQSFKPLLTEGYSLKFYPLGCTSFNAD 458
Query: 221 LGGLPPMGCLPLERTTNF 238
G LPL +T+ F
Sbjct: 459 FDGDQMSIFLPLIKTSKF 476
>COA2_SV40 (P03093) Coat protein VP2 [Contains: Coat protein VP3]
Length = 352
Score = 30.0 bits (66), Expect = 9.4
Identities = 31/123 (25%), Positives = 47/123 (38%), Gaps = 18/123 (14%)
Query: 105 FNISQFATG-------VSFASAATGYDNATSDVLSVIPLWKQLEYYKEYQKKLGAYLGEK 157
F++++ A G V AS AT TS+ ++ I L Q Y A G
Sbjct: 24 FSVAEIAAGEAAAAIEVQLASVATVEGLTTSEAIAAIGLIPQA--YAVISGAPAAIAGFA 81
Query: 158 KAKETITKALYIISLGTNDFLE--------NYYTIPGRASQ-YTPSEYQNFLAGIAQNFI 208
+T+T + +G F + Y PG A Y P +Y + L Q F+
Sbjct: 82 ALLQTVTGVSAVAQVGYRFFSDWDHKVSTVGLYQQPGMAVDLYRPDDYYDILFPGVQTFV 141
Query: 209 HKL 211
H +
Sbjct: 142 HSV 144
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.138 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,951,402
Number of Sequences: 164201
Number of extensions: 1839110
Number of successful extensions: 4249
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 14
Number of HSP's that attempted gapping in prelim test: 4232
Number of HSP's gapped (non-prelim): 22
length of query: 359
length of database: 59,974,054
effective HSP length: 111
effective length of query: 248
effective length of database: 41,747,743
effective search space: 10353440264
effective search space used: 10353440264
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 66 (30.0 bits)
Medicago: description of AC140670.14


