

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140549.8 - phase: 0
(122 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
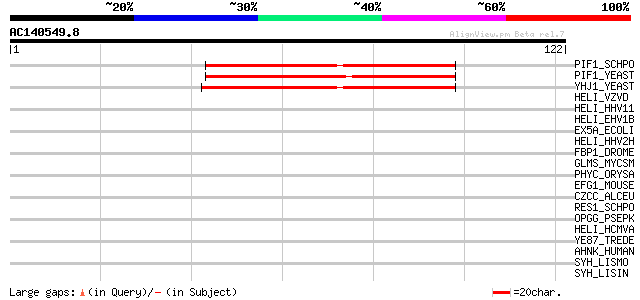
Score E
Sequences producing significant alignments: (bits) Value
PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1, m... 49 3e-06
PIF1_YEAST (P07271) DNA repair and recombination protein PIF1, m... 47 1e-05
YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergeni... 41 4e-04
HELI_VZVD (P09303) Probable helicase 33 0.11
HELI_HHV11 (P10189) Probable helicase 33 0.11
HELI_EHV1B (P28934) Probable helicase 33 0.15
EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC 3.1.1... 32 0.25
HELI_HHV2H (P28277) Probable helicase 31 0.43
FBP1_DROME (Q04691) Fat-body protein-1 precursor (P1 protein) 29 1.6
GLMS_MYCSM (O68956) Glucosamine--fructose-6-phosphate aminotrans... 29 2.1
PHYC_ORYSA (Q9ZWI9) Phytochrome C 28 2.8
EFG1_MOUSE (Q8K0D5) Elongation factor G 1, mitochondrial precurs... 28 2.8
CZCC_ALCEU (P13509) Cobalt-zinc-cadmium resistance protein czcC ... 28 2.8
RES1_SCHPO (P33520) Cell division cycle related-protein res1/sct1 28 3.6
OPGG_PSEPK (Q88D03) Glucans biosynthesis protein G precursor 28 3.6
HELI_HCMVA (P16736) Probable helicase 28 3.6
YE87_TREDE (P62042) Hypothetical UPF0082 protein TDE1487 27 6.2
AHNK_HUMAN (Q09666) Neuroblast differentiation associated protei... 27 6.2
SYH_LISMO (Q8Y708) Histidyl-tRNA synthetase (EC 6.1.1.21) (Histi... 27 8.1
SYH_LISIN (Q92BJ3) Histidyl-tRNA synthetase (EC 6.1.1.21) (Histi... 27 8.1
>PIF1_SCHPO (Q9UUA2) DNA repair and recombination protein pif1,
mitochondrial precursor
Length = 805
Score = 48.5 bits (114), Expect = 3e-06
Identities = 24/55 (43%), Positives = 40/55 (72%), Gaps = 1/55 (1%)
Query: 44 RRQFPISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKIL 98
R Q P+ +++A+ I+K+QGQ+L+ V V L +F GQ YVA+S+ T++ GL++L
Sbjct: 708 RSQIPLILAYAISIHKAQGQTLDRVKVDL-GRVFEKGQAYVALSRATTQEGLQVL 761
>PIF1_YEAST (P07271) DNA repair and recombination protein PIF1,
mitochondrial precursor
Length = 857
Score = 46.6 bits (109), Expect = 1e-05
Identities = 25/55 (45%), Positives = 38/55 (68%), Gaps = 1/55 (1%)
Query: 44 RRQFPISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKIL 98
R Q P+ +++++ I+KSQGQ+L V V L +F GQ YVA+S+ SR GL++L
Sbjct: 691 RVQLPLMLAWSLSIHKSQGQTLPKVKVDLRR-VFEKGQAYVALSRAVSREGLQVL 744
>YHJ1_YEAST (P38766) Hypothetical helicase in SLT2-PUT2 intergenic
region
Length = 723
Score = 41.2 bits (95), Expect = 4e-04
Identities = 21/56 (37%), Positives = 38/56 (67%), Gaps = 1/56 (1%)
Query: 43 KRRQFPISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKIL 98
+R Q P+ + +A+ I+K+QGQ+++ + V L IF GQ+YVA+S+ + L++L
Sbjct: 645 ERTQIPLMLCWALSIHKAQGQTIQRLKVDL-RRIFEAGQVYVALSRAVTMDTLQVL 699
>HELI_VZVD (P09303) Probable helicase
Length = 881
Score = 33.1 bits (74), Expect = 0.11
Identities = 19/54 (35%), Positives = 29/54 (53%)
Query: 47 FPISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKILIN 100
+ +S AM I +SQG SLE V + + +YVA+S+ S LK+ +N
Sbjct: 799 YGLSSKLAMTIARSQGLSLEKVAICFTADKLRLNSVYVAMSRTVSSRFLKMNLN 852
>HELI_HHV11 (P10189) Probable helicase
Length = 882
Score = 33.1 bits (74), Expect = 0.11
Identities = 22/65 (33%), Positives = 32/65 (48%), Gaps = 4/65 (6%)
Query: 47 FPISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKILIND----D 102
+ IS AM I +SQG SL+ V + YVA+S+ TS L++ +N
Sbjct: 800 YGISSKLAMTITRSQGLSLDKVAICFTPGNLRLNSAYVAMSRTTSSEFLRMNLNPLRERH 859
Query: 103 DDDDI 107
D DD+
Sbjct: 860 DRDDV 864
>HELI_EHV1B (P28934) Probable helicase
Length = 881
Score = 32.7 bits (73), Expect = 0.15
Identities = 18/54 (33%), Positives = 29/54 (53%)
Query: 47 FPISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKILIN 100
+ +S AM I +SQG SL+ V + P +YVA+S+ S L++ +N
Sbjct: 799 YGLSSKLAMTIARSQGLSLDKVAICFPRNNLRINSVYVAMSRTVSSRFLRMNLN 852
>EX5A_ECOLI (P04993) Exodeoxyribonuclease V alpha chain (EC
3.1.11.5) (Exodeoxyribonuclease V 67 kDa polypeptide)
Length = 608
Score = 32.0 bits (71), Expect = 0.25
Identities = 14/44 (31%), Positives = 26/44 (58%), Gaps = 3/44 (6%)
Query: 52 SFAMIINKSQGQSLEHVGVYLPS---PIFSHGQLYVAISQVTSR 92
++AM ++KSQG +H + LPS P+ + +Y A+++ R
Sbjct: 533 TWAMTVHKSQGSEFDHAALILPSQRTPVVTRELVYTAVTRARRR 576
>HELI_HHV2H (P28277) Probable helicase
Length = 881
Score = 31.2 bits (69), Expect = 0.43
Identities = 19/54 (35%), Positives = 27/54 (49%)
Query: 47 FPISVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKILIN 100
+ IS AM I +SQG SL+ V + YVA+S+ TS L + +N
Sbjct: 799 YGISSKLAMTITRSQGLSLDKVAICFTPGNLRLNSAYVAMSRTTSSEFLHMNLN 852
>FBP1_DROME (Q04691) Fat-body protein-1 precursor (P1 protein)
Length = 1030
Score = 29.3 bits (64), Expect = 1.6
Identities = 13/22 (59%), Positives = 16/22 (72%)
Query: 100 NDDDDDDIDVASNVVYREVFRN 121
NDDDDDD D NVV+R+ R+
Sbjct: 580 NDDDDDDNDDDVNVVHRQGLRS 601
>GLMS_MYCSM (O68956) Glucosamine--fructose-6-phosphate
aminotransferase [isomerizing] (EC 2.6.1.16)
(Hexosephosphate aminotransferase)
(D-fructose-6-phosphate amidotransferase) (GFAT)
(L-glutamine-D-fructose-6-phosphate amidotransferase)
(Glucosamine-6-ph
Length = 627
Score = 28.9 bits (63), Expect = 2.1
Identities = 13/41 (31%), Positives = 24/41 (57%), Gaps = 3/41 (7%)
Query: 70 VYLPSP---IFSHGQLYVAISQVTSRGGLKILINDDDDDDI 107
V +PSP H +L I ++ +RG + ++I ++DDD +
Sbjct: 533 VVMPSPKNAAMLHAKLLSNIREIQARGAVTVVIAEEDDDTV 573
>PHYC_ORYSA (Q9ZWI9) Phytochrome C
Length = 1137
Score = 28.5 bits (62), Expect = 2.8
Identities = 18/58 (31%), Positives = 29/58 (49%), Gaps = 1/58 (1%)
Query: 55 MIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKILINDDDDDDIDVASN 112
+I + S Q + G L +P H Q ++ V S + + IN+D+DDD D S+
Sbjct: 299 IIQDDSLTQPISICGSTLRAPHGCHAQYMASMGSVASLV-MSVTINEDEDDDGDTGSD 355
>EFG1_MOUSE (Q8K0D5) Elongation factor G 1, mitochondrial precursor
(mEF-G 1) (Elongation factor G1)
Length = 751
Score = 28.5 bits (62), Expect = 2.8
Identities = 19/48 (39%), Positives = 25/48 (51%), Gaps = 2/48 (4%)
Query: 58 NKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKILINDDDDD 105
NK L+ V YLP+P S Q Y ++Q S+ KIL+N DD
Sbjct: 304 NKGVQPLLDAVLEYLPNP--SEVQNYAILNQNDSKEKTKILMNPQRDD 349
>CZCC_ALCEU (P13509) Cobalt-zinc-cadmium resistance protein czcC
precursor (Cation efflux system protein czcC)
Length = 417
Score = 28.5 bits (62), Expect = 2.8
Identities = 17/49 (34%), Positives = 28/49 (56%), Gaps = 3/49 (6%)
Query: 43 KRRQFP-ISVSFAMIINKSQGQSLEHVGVYLPSPIF--SHGQLYVAISQ 88
+ RQ+P ++VS + +++ +GV +P PIF + G LY AI Q
Sbjct: 270 RSRQYPDLTVSLGAKRDTEANRNMAVIGVAIPLPIFDRNQGNLYSAIRQ 318
>RES1_SCHPO (P33520) Cell division cycle related-protein res1/sct1
Length = 637
Score = 28.1 bits (61), Expect = 3.6
Identities = 12/30 (40%), Positives = 17/30 (56%)
Query: 25 FIPRLSLTPSDNRIPFKFKRRQFPISVSFA 54
F+P + TP NR P + RR P+ SF+
Sbjct: 108 FMPMSTFTPQSNRKPTEAYRRNSPVKKSFS 137
>OPGG_PSEPK (Q88D03) Glucans biosynthesis protein G precursor
Length = 559
Score = 28.1 bits (61), Expect = 3.6
Identities = 13/45 (28%), Positives = 20/45 (43%)
Query: 73 PSPIFSHGQLYVAISQVTSRGGLKILINDDDDDDIDVASNVVYRE 117
P P H +Y + S G K+ + +D +DV S V R+
Sbjct: 191 PKPADKHLVIYALLDSPRSTGAYKLTLRPGNDTVVDVQSRVFLRD 235
>HELI_HCMVA (P16736) Probable helicase
Length = 956
Score = 28.1 bits (61), Expect = 3.6
Identities = 22/51 (43%), Positives = 29/51 (56%), Gaps = 6/51 (11%)
Query: 54 AMIINKSQGQSLEHVGV----YLPSPIFSHGQLYVAISQVTSRGGLKILIN 100
AM I KSQG SLE V V + + SH +YVA+S+VT L + +N
Sbjct: 879 AMTIAKSQGLSLEKVAVDFGDHPKNLKMSH--IYVAMSRVTDPEHLMMNVN 927
>YE87_TREDE (P62042) Hypothetical UPF0082 protein TDE1487
Length = 247
Score = 27.3 bits (59), Expect = 6.2
Identities = 19/68 (27%), Positives = 34/68 (49%), Gaps = 5/68 (7%)
Query: 53 FAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKILINDDDDDDID-VAS 111
FA ++ Q + E V + + YVA+ T++ LK++ ++DDD+ V+S
Sbjct: 180 FASVLEALQEKGFESVS----AAVSMVPDTYVALDADTTQKALKMIDKLEEDDDVQTVSS 235
Query: 112 NVVYREVF 119
N+ E F
Sbjct: 236 NIEIPEGF 243
>AHNK_HUMAN (Q09666) Neuroblast differentiation associated protein
AHNAK (Desmoyokin) (Fragments)
Length = 2960
Score = 27.3 bits (59), Expect = 6.2
Identities = 14/44 (31%), Positives = 25/44 (56%), Gaps = 1/44 (2%)
Query: 4 EGGTYQLEGRVISGSNIDEKVFIPRLSLTPSDNRIPFKFKRRQF 47
EGG L+G + ++D + P +++ D +IP KFK+ +F
Sbjct: 2008 EGGEVDLKGPKVEAPSLDVHMDSPDINIEGPDVKIP-KFKKPKF 2050
>SYH_LISMO (Q8Y708) Histidyl-tRNA synthetase (EC 6.1.1.21)
(Histidine--tRNA ligase) (HisRS)
Length = 425
Score = 26.9 bits (58), Expect = 8.1
Identities = 18/73 (24%), Positives = 37/73 (50%), Gaps = 2/73 (2%)
Query: 50 SVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKILINDDDDDDIDV 109
SV A + +K G+ + + +Y P+F + + + ++ G++ L +DD D++V
Sbjct: 85 SVVRAFVEHKLYGEVSQPIKMYYNEPMFRYERPQGGRQRQFTQMGIEALGSDDPSIDVEV 144
Query: 110 ASNVVYREVFRNV 122
S + E FR +
Sbjct: 145 IS--LAMEFFRKI 155
>SYH_LISIN (Q92BJ3) Histidyl-tRNA synthetase (EC 6.1.1.21)
(Histidine--tRNA ligase) (HisRS)
Length = 425
Score = 26.9 bits (58), Expect = 8.1
Identities = 18/73 (24%), Positives = 37/73 (50%), Gaps = 2/73 (2%)
Query: 50 SVSFAMIINKSQGQSLEHVGVYLPSPIFSHGQLYVAISQVTSRGGLKILINDDDDDDIDV 109
SV A + +K G+ + + +Y P+F + + + ++ G++ L +DD D++V
Sbjct: 85 SVVRAFVEHKLYGEVSQPIKMYYNEPMFRYERPQGGRQRQFTQMGIEALGSDDPSIDVEV 144
Query: 110 ASNVVYREVFRNV 122
S + E FR +
Sbjct: 145 IS--LAMEFFRKI 155
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.320 0.140 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,081,940
Number of Sequences: 164201
Number of extensions: 531682
Number of successful extensions: 1417
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 11
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 1402
Number of HSP's gapped (non-prelim): 22
length of query: 122
length of database: 59,974,054
effective HSP length: 98
effective length of query: 24
effective length of database: 43,882,356
effective search space: 1053176544
effective search space used: 1053176544
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC140549.8


