

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140549.19 - phase: 0
(373 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
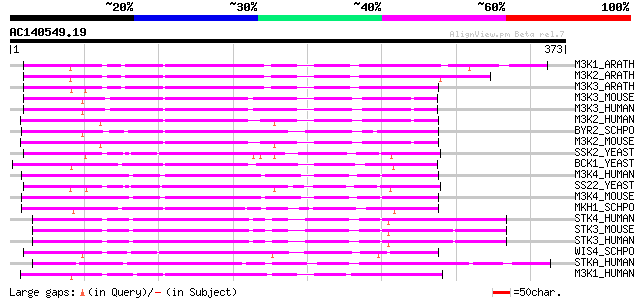
Score E
Sequences producing significant alignments: (bits) Value
M3K1_ARATH (O22040) Mitogen-activated protein kinase kinase kina... 197 4e-50
M3K2_ARATH (Q9FZ36) Mitogen-activated protein kinase kinase kina... 197 5e-50
M3K3_ARATH (O22042) Mitogen-activated protein kinase kinase kina... 196 8e-50
M3K3_MOUSE (Q61084) Mitogen-activated protein kinase kinase kina... 182 1e-45
M3K3_HUMAN (Q99759) Mitogen-activated protein kinase kinase kina... 182 2e-45
M3K2_HUMAN (Q9Y2U5) Mitogen-activated protein kinase kinase kina... 172 9e-43
BYR2_SCHPO (P28829) Protein kinase byr2 (EC 2.7.1.-) (Protein ki... 171 4e-42
M3K2_MOUSE (Q61083) Mitogen-activated protein kinase kinase kina... 170 6e-42
SSK2_YEAST (P53599) MAP kinase kinase kinase SSK2 (EC 2.7.1.-) (... 169 1e-41
BCK1_YEAST (Q01389) Serine/threonine-protein kinase BCK1/SLK1/SS... 166 7e-41
M3K4_HUMAN (Q9Y6R4) Mitogen-activated protein kinase kinase kina... 165 2e-40
SS22_YEAST (P25390) Serine/threonine-protein kinase SSK22 (EC 2.... 164 4e-40
M3K4_MOUSE (O08648) Mitogen-activated protein kinase kinase kina... 163 7e-40
MKH1_SCHPO (Q10407) MAP kinase kinase kinase mkh1 (EC 2.7.1.-) 162 1e-39
STK4_HUMAN (Q13043) Serine/threonine-protein kinase 4 (EC 2.7.1.... 160 5e-39
STK3_MOUSE (Q9JI10) Serine/threonine-protein kinase 3 (EC 2.7.1.... 158 2e-38
STK3_HUMAN (Q13188) Serine/threonine-protein kinase 3 (EC 2.7.1.... 157 4e-38
WIS4_SCHPO (O14299) MAP kinase kinase kinase wis4 (EC 2.7.1.-) (... 157 5e-38
STKA_HUMAN (O94804) Serine/threonine-protein kinase 10 (EC 2.7.1... 156 7e-38
M3K1_HUMAN (Q13233) Mitogen-activated protein kinase kinase kina... 155 1e-37
>M3K1_ARATH (O22040) Mitogen-activated protein kinase kinase kinase
1 (EC 2.7.1.-) (Arabidospsis NPK1-related protein kinase
1)
Length = 666
Score = 197 bits (501), Expect = 4e-50
Identities = 131/371 (35%), Positives = 191/371 (51%), Gaps = 55/371 (14%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVK----------KAHSEAGRDALENEVNILNT 59
W KG+++G G+FG+V++ MN +G L VK K ++A LE EV +L
Sbjct: 69 WRKGQLIGRGAFGTVYMGMNLDSGELLAVKQVLIAANFASKEKTQAHIQELEEEVKLLKN 128
Query: 60 LNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIV 119
L+ P IV+ LGT +D+ L++ +E++ GGS++ + KFG E VVR YTRQ++
Sbjct: 129 LSH---PNIVRYLGTV--REDDTLNILLEFVPGGSISSLLEKFG-PFPESVVRTYTRQLL 182
Query: 120 HGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTP 179
GL +LH H I+H D+K N+L+ + G +KLADFG +K+V + M + S GTP
Sbjct: 183 LGLEYLHNHAIMHRDIKGANILVDNKGCIKLADFGASKQVAELATMTGAKSM----KGTP 238
Query: 180 LWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMA 239
WMAPEV+L +S +ADIWS+GCTVIEM TG+ PW +A
Sbjct: 239 YWMAPEVILQTGHS-----------FSADIWSVGCTVIEMVTGKAPWSQQ-----YKEVA 282
Query: 240 AMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHY 299
A+F I P P S + DFL +CL P R TA ELL HPF++ HK
Sbjct: 283 AIFFIGTTKSHPPIPDTLSSDAKDFLLKCLQEVPNLRPTASELLKHPFVMG----KHKES 338
Query: 300 SASSPASV---------LEVHQFEDTYDDDDDDDDSDDNDELISPAGGNHFFITKELLCP 350
+++ SV L+++ + T D DD N G ++ + +
Sbjct: 339 ASTDLGSVLNNLSTPLPLQINNTKSTPDSTCDDVGDMCN------FGSLNYSLVDPVKSI 392
Query: 351 QGTTMWQLEDS 361
Q +WQ D+
Sbjct: 393 QNKNLWQQNDN 403
>M3K2_ARATH (Q9FZ36) Mitogen-activated protein kinase kinase kinase
2 (EC 2.7.1.-) (Arabidospsis NPK1-related protein kinase
2)
Length = 651
Score = 197 bits (500), Expect = 5e-50
Identities = 123/332 (37%), Positives = 180/332 (54%), Gaps = 44/332 (13%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVK----------KAHSEAGRDALENEVNILNT 59
W KG+++G G+FG+V++ MN +G L VK K ++A LE EV +L
Sbjct: 68 WRKGQLIGRGAFGTVYMGMNLDSGELLAVKQVLITSNCASKEKTQAHIQELEEEVKLLKN 127
Query: 60 LNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIV 119
L+ P IV+ LGT +D L++ +E++ GGS++ + KFG + E VVR YT Q++
Sbjct: 128 LSH---PNIVRYLGTV--REDETLNILLEFVPGGSISSLLEKFG-AFPESVVRTYTNQLL 181
Query: 120 HGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTP 179
GL +LH H I+H D+K N+L+ + G +KLADFG +K+V + ++ + S GTP
Sbjct: 182 LGLEYLHNHAIMHRDIKGANILVDNQGCIKLADFGASKQVAELATISGAKSM----KGTP 237
Query: 180 LWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMA 239
WMAPEV+L +S +ADIWS+GCTVIEM TG+ PW +A
Sbjct: 238 YWMAPEVILQTGHS-----------FSADIWSVGCTVIEMVTGKAPWSQQ-----YKEIA 281
Query: 240 AMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL-------VSTT 292
A+F I P P + S + DFL +CL ++P R TA ELL HPF+ S
Sbjct: 282 AIFHIGTTKSHPPIPDNISSDANDFLLKCLQQEPNLRPTASELLKHPFVTGKQKESASKD 341
Query: 293 LTHHKHYSASS-PASVLEVHQFEDTYDDDDDD 323
LT S S P+ + + ++ + DD D
Sbjct: 342 LTSFMDNSCSPLPSELTNITSYQTSTSDDVGD 373
>M3K3_ARATH (O22042) Mitogen-activated protein kinase kinase kinase
3 (EC 2.7.1.-) (Arabidospsis NPK1-related protein kinase
3)
Length = 651
Score = 196 bits (498), Expect = 8e-50
Identities = 115/289 (39%), Positives = 169/289 (57%), Gaps = 36/289 (12%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVKK---AHSEAGRDA-------LENEVNILNT 59
W KG+++GCG+FG V++ MN +G L +K+ A S A ++ LE EV +L
Sbjct: 68 WRKGELIGCGAFGRVYMGMNLDSGELLAIKQVLIAPSSASKEKTQGHIRELEEEVQLLKN 127
Query: 60 LNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIV 119
L+ P IV+ LGT + + L++ ME++ GGS++ + KFG S E V+ +YT+Q++
Sbjct: 128 LSH---PNIVRYLGTV--RESDSLNILMEFVPGGSISSLLEKFG-SFPEPVIIMYTKQLL 181
Query: 120 HGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTP 179
GL +LH +GI+H D+K N+L+ + G ++LADFG +K+V + +N + S GTP
Sbjct: 182 LGLEYLHNNGIMHRDIKGANILVDNKGCIRLADFGASKKVVELATVNGAKSM----KGTP 237
Query: 180 LWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMA 239
WMAPEV+L +S +ADIWS+GCTVIEMATG+PPW + A
Sbjct: 238 YWMAPEVILQTGHS-----------FSADIWSVGCTVIEMATGKPPWSEQ-----YQQFA 281
Query: 240 AMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
A+ I P P S E DFL +CL ++P R +A ELL HPF+
Sbjct: 282 AVLHIGRTKAHPPIPEDLSPEAKDFLMKCLHKEPSLRLSATELLQHPFV 330
>M3K3_MOUSE (Q61084) Mitogen-activated protein kinase kinase kinase
3 (EC 2.7.1.37) (MAPK/ERK kinase kinase 3) (MEK kinase
3) (MEKK 3)
Length = 626
Score = 182 bits (462), Expect = 1e-45
Identities = 115/285 (40%), Positives = 157/285 (54%), Gaps = 32/285 (11%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGR-------DALENEVNILNTLNS 62
W +GK++G G+FG V+L + TG K+ + ALE E+ +L L
Sbjct: 362 WRRGKLLGQGAFGRVYLCYDVDTGRELASKQVQFDPDSPETSKEVSALECEIQLLKNLQH 421
Query: 63 SPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGL 122
IVQ G D + L +FMEYM GGS+ D +G +L E V R YTRQI+ G+
Sbjct: 422 ER---IVQYYGCLRDRAEKILTIFMEYMPGGSVKDQLKAYG-ALTESVTRKYTRQILEGM 477
Query: 123 YHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWM 182
+LH + IVH D+K N+L S+GNVKL DFG +KR++ + S + I + GTP WM
Sbjct: 478 SYLHSNMIVHRDIKGANILRDSAGNVKLGDFGASKRLQ---TICMSGTGIRSVTGTPYWM 534
Query: 183 APEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMF 242
+PEV+ + AD+WSLGCTV+EM T +PPW + + MAA+F
Sbjct: 535 SPEVISGEGYG-----------RKADVWSLGCTVVEMLTEKPPWAEYE------AMAAIF 577
Query: 243 KIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPF 287
KIA PQ P H S+ G DFLRR V + ++R +A ELL H F
Sbjct: 578 KIATQPTNPQLPSHISEHGRDFLRRIFV-EARQRPSAEELLTHHF 621
>M3K3_HUMAN (Q99759) Mitogen-activated protein kinase kinase kinase
3 (EC 2.7.1.37) (MAPK/ERK kinase kinase 3) (MEK kinase
3) (MEKK 3)
Length = 626
Score = 182 bits (461), Expect = 2e-45
Identities = 114/285 (40%), Positives = 157/285 (55%), Gaps = 32/285 (11%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGR-------DALENEVNILNTLNS 62
W +GK++G G+FG V+L + TG K+ + ALE E+ +L L
Sbjct: 362 WRRGKLLGQGAFGRVYLCYDVDTGRELASKQVQFDPDSPETSKEVSALECEIQLLKNLQH 421
Query: 63 SPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGL 122
IVQ G D + L +FMEYM GGS+ D +G +L E V R YTRQI+ G+
Sbjct: 422 ER---IVQYYGCLRDRAEKTLTIFMEYMPGGSVKDQLKAYG-ALTESVTRKYTRQILEGM 477
Query: 123 YHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWM 182
+LH + IVH D+K N+L S+GNVKL DFG +KR++ + S + + + GTP WM
Sbjct: 478 SYLHSNMIVHRDIKGANILRDSAGNVKLGDFGASKRLQ---TICMSGTGMRSVTGTPYWM 534
Query: 183 APEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMF 242
+PEV+ + AD+WSLGCTV+EM T +PPW + + MAA+F
Sbjct: 535 SPEVISGEGYG-----------RKADVWSLGCTVVEMLTEKPPWAEYE------AMAAIF 577
Query: 243 KIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPF 287
KIA PQ P H S+ G DFLRR V + ++R +A ELL H F
Sbjct: 578 KIATQPTNPQLPSHISEHGRDFLRRIFV-EARQRPSAEELLTHHF 621
>M3K2_HUMAN (Q9Y2U5) Mitogen-activated protein kinase kinase kinase
2 (EC 2.7.1.37) (MAPK/ERK kinase kinase 2) (MEK kinase
2) (MEKK 2)
Length = 618
Score = 172 bits (437), Expect = 9e-43
Identities = 111/290 (38%), Positives = 154/290 (52%), Gaps = 36/290 (12%)
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNT----LNSS 63
+ W GK++G G+FG V+L + TG VK+ + EVN L L +
Sbjct: 353 TNWRLGKLLGQGAFGRVYLCYDVDTGRELAVKQVQFDPDSPETSKEVNALECEIQLLKNF 412
Query: 64 PSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLY 123
IVQ G D Q+ L +FMEYM GGS+ D +G +L E+ R YTRQI+ G++
Sbjct: 413 LHERIVQYYGCLRDPQEKTLSIFMEYMPGGSIKDQLKAYG-ALTENGTRKYTRQILEGVH 471
Query: 124 HLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANG-----GT 178
+LH + I+H D+K N+L S+GNVKL DFG +KR++ + C+ G GT
Sbjct: 472 YLHSNMILHRDIKGANILRDSTGNVKLGDFGASKRLQ--------TICLSGTGMKSVTGT 523
Query: 179 PLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPM 238
P WM+PEV+ + ADIWS+ CTV+EM T +PPW + + M
Sbjct: 524 PYWMSPEVISGQGYG-----------RKADIWSVACTVVEMLTEKPPWAEFE------AM 566
Query: 239 AAMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
AA+FKIA P+ P H S DFL+R V + K R +A ELL H F+
Sbjct: 567 AAIFKIATQPTNPKLPPHVSDYTRDFLKRIFV-EAKLRPSADELLRHMFV 615
>BYR2_SCHPO (P28829) Protein kinase byr2 (EC 2.7.1.-) (Protein
kinase ste8) (MAPK kinase kinase) (MAPKKK)
Length = 659
Score = 171 bits (432), Expect = 4e-42
Identities = 104/291 (35%), Positives = 163/291 (55%), Gaps = 36/291 (12%)
Query: 9 EWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGR----------DALENEVNILN 58
+W++G ++G GSFG V+L MN S+G L VK+ ++ DAL E+ +L
Sbjct: 393 KWIRGALIGSGSFGQVYLGMNASSGELMAVKQVILDSVSESKDRHAKLLDALAGEIALLQ 452
Query: 59 TLNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQI 118
L+ +IVQ LG++ + + L++F+EY+ GGS+A + +G S E +V+ + +Q
Sbjct: 453 ELSHE---HIVQYLGSNLN--SDHLNIFLEYVPGGSVAGLLTMYG-SFEETLVKNFIKQT 506
Query: 119 VHGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGT 178
+ GL +LH GIVH D+K N+L+ + G +K++DFG +K+++ N K+ + G+
Sbjct: 507 LKGLEYLHSRGIVHRDIKGANILVDNKGKIKISDFGISKKLELNSTSTKTGGARPSFQGS 566
Query: 179 PLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPM 238
WMAPEV+ + + DIWSLGC VIEM T + P+ + D M
Sbjct: 567 SFWMAPEVV-----------KQTMHTEKTDIWSLGCLVIEMLTSKHPYPNCD------QM 609
Query: 239 AAMFKIACGDGI-PQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
A+F+I G+ I P+FP + S DFL + D R TA ELL+HPF+
Sbjct: 610 QAIFRI--GENILPEFPSNISSSAIDFLEKTFAIDCNLRPTASELLSHPFV 658
>M3K2_MOUSE (Q61083) Mitogen-activated protein kinase kinase kinase
2 (EC 2.7.1.37) (MAPK/ERK kinase kinase 2) (MEK kinase
2) (MEKK 2)
Length = 619
Score = 170 bits (430), Expect = 6e-42
Identities = 110/290 (37%), Positives = 152/290 (51%), Gaps = 36/290 (12%)
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNT----LNSS 63
+ W GK++G G+FG V+L + TG VK+ EVN L L +
Sbjct: 354 TNWRLGKLLGQGAFGRVYLCYDVDTGRELAVKQVQFNPESPETSKEVNALECEIQLLKNL 413
Query: 64 PSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLY 123
IVQ G D Q+ L +FME GGS+ D +G +L E+V R YTRQI+ G++
Sbjct: 414 LHERIVQYYGCLRDPQEKTLSIFMELSPGGSIKDQLKAYG-ALTENVTRKYTRQILEGVH 472
Query: 124 HLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANG-----GT 178
+LH + IVH D+K N+L S+GN+KL DFG +KR++ + C+ G GT
Sbjct: 473 YLHSNMIVHRDIKGANILRDSTGNIKLGDFGASKRLQ--------TICLSGTGMKSVTGT 524
Query: 179 PLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPM 238
P WM+PEV+ + ADIWS+ CTV+EM T +PPW + + M
Sbjct: 525 PYWMSPEVISGEGYG-----------RKADIWSVACTVVEMLTEKPPWAEFE------AM 567
Query: 239 AAMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
AA+FKIA P+ P H S DFL+R V + K R +A ELL H F+
Sbjct: 568 AAIFKIATQPTNPKLPPHVSDYTRDFLKRIFV-EAKLRPSAEELLRHMFV 616
>SSK2_YEAST (P53599) MAP kinase kinase kinase SSK2 (EC 2.7.1.-)
(Suppressor of sensor kinase 2)
Length = 1579
Score = 169 bits (427), Expect = 1e-41
Identities = 113/314 (35%), Positives = 168/314 (52%), Gaps = 54/314 (17%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDA------LENEVNILNTLNSS 63
W K +G G+FG V+ A++ G + VK+ + + + ++ E+++L LN
Sbjct: 1266 WQKRNFIGGGTFGRVYSAVDLDNGEILAVKEINIQDSKSMQKIFPLIKEEMSVLEILNH- 1324
Query: 64 PSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLY 123
P IV G + H+D ++++FMEY GGSLA + + G +E V +VYT Q++ GL
Sbjct: 1325 --PNIVSYYGVEV-HRD-KVNIFMEYCEGGSLAALL-EHGRIEDEMVTQVYTLQLLEGLA 1379
Query: 124 HLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKN----MNMNK------------ 167
+LH+ GIVH D+K +N+LL +G +K DFG AK++ N +MNK
Sbjct: 1380 YLHESGIVHRDVKPENILLDFNGVIKYVDFGAAKKIANNGTRLASMNKIENADGEHEDVT 1439
Query: 168 --SSSCIIANG--------GTPLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVI 217
S S + N GTP++MAPE SI + A D+WSLGC V+
Sbjct: 1440 HVSDSKAVKNNENALLDMMGTPMYMAPE--------SITGSTTKGKLGADDVWSLGCVVL 1491
Query: 218 EMATGRPPWVDDDLISISNPMAAMFKIACGDGIPQFPI--HFSQEGFDFLRRCLVRDPKK 275
EM TGR PW ++ N A M+ +A G PQFP S G FL RCL+++P K
Sbjct: 1492 EMITGRRPWA-----NLDNEWAIMYHVAAGH-TPQFPTKDEVSSAGMKFLERCLIQNPSK 1545
Query: 276 RSTALELLNHPFLV 289
R++A+ELL P++V
Sbjct: 1546 RASAVELLMDPWIV 1559
>BCK1_YEAST (Q01389) Serine/threonine-protein kinase BCK1/SLK1/SSP31
(EC 2.7.1.37)
Length = 1478
Score = 166 bits (421), Expect = 7e-41
Identities = 107/296 (36%), Positives = 163/296 (54%), Gaps = 35/296 (11%)
Query: 3 GIIKGSEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKK-------AHSEAGRDALENEVN 55
G K WMKG+M+G GSFG+V+L +N +TG + VK+ + +EA +E +
Sbjct: 1168 GEYKEFAWMKGEMIGKGSFGAVYLCLNVTTGEMMAVKQVEVPKYSSQNEAILSTVEALRS 1227
Query: 56 ILNTLNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYT 115
++TL IVQ LG +++++N +F+EY++GGS+ + +G +E +++ T
Sbjct: 1228 EVSTLKDLDHLNIVQYLG--FENKNNIYSLFLEYVAGGSVGSLIRMYG-RFDEPLIKHLT 1284
Query: 116 RQIVHGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIAN 175
Q++ GL +LH GI+H D+K N+LL G K++DFG +++ K + S+ +
Sbjct: 1285 TQVLKGLAYLHSKGILHRDMKADNLLLDQDGICKISDFGISRKSK-----DIYSNSDMTM 1339
Query: 176 GGTPLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISIS 235
GT WMAPE++ K S A DIWSLGC V+EM G+ PW + +++
Sbjct: 1340 RGTVFWMAPEMVDTKQGYS----------AKVDIWSLGCIVLEMFAGKRPWSNLEVV--- 1386
Query: 236 NPMAAMFKIACGDGIPQFPIH----FSQEGFDFLRRCLVRDPKKRSTALELLNHPF 287
AAMFKI P P SQ G +FL C +P+KR TA ELL+HPF
Sbjct: 1387 ---AAMFKIGKSKSAPPIPEDTLPLISQIGRNFLDACFEINPEKRPTANELLSHPF 1439
>M3K4_HUMAN (Q9Y6R4) Mitogen-activated protein kinase kinase kinase 4
(EC 2.7.1.37) (MAPK/ERK kinase kinase 4) (MEK kinase 4)
(MEKK 4) (MAP three kinase 1)
Length = 1607
Score = 165 bits (418), Expect = 2e-40
Identities = 98/281 (34%), Positives = 145/281 (50%), Gaps = 22/281 (7%)
Query: 9 EWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAG-RDALENEVNILNTLNSSPSPY 67
+W +G +G G +G V+ ++ TG L +K+ + ++ + L P
Sbjct: 1341 KWQRGNKIGEGQYGKVYTCISVDTGELMAMKEIRFQPNDHKTIKETADELKIFEGIKHPN 1400
Query: 68 IVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQ 127
+V+ G + ++ +++FMEY G+L +VS L E V+R+Y++QI + LH+
Sbjct: 1401 LVRYFGVELHREE--MYIFMEYCDEGTLEEVSRL---GLQEHVIRLYSKQITIAINVLHE 1455
Query: 128 HGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVL 187
HGIVH D+K N+ L SSG +KL DFGC+ ++K N + + GT +MAPEV+
Sbjct: 1456 HGIVHRDIKGANIFLTSSGLIKLGDFGCSVKLKNNAQTMPGE--VNSTLGTAAYMAPEVI 1513
Query: 188 LMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACG 247
AADIWSLGC VIEM TG+ PW + + M+K+ G
Sbjct: 1514 TRAKGEGHG--------RAADIWSLGCVVIEMVTGKRPWHE-----YEHNFQIMYKVGMG 1560
Query: 248 DGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
P P S EG DFL CL DPK R TA +LL+H F+
Sbjct: 1561 HK-PPIPERLSPEGKDFLSHCLESDPKMRWTASQLLDHSFV 1600
>SS22_YEAST (P25390) Serine/threonine-protein kinase SSK22 (EC
2.7.1.37) (MAP kinase kinase kinase SSK22) (Suppressor of
sensor kinase 22)
Length = 1331
Score = 164 bits (414), Expect = 4e-40
Identities = 109/298 (36%), Positives = 163/298 (54%), Gaps = 38/298 (12%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVK--KAHSEAGRDAL----ENEVNILNTLNSS 63
W K +G G+FG V+ A+N G + VK K H + + E+ +L LN
Sbjct: 1034 WQKRSFIGGGTFGQVYSAINLENGEILAVKEIKIHDTTTMKKIFPLIKEEMTVLEMLNH- 1092
Query: 64 PSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLY 123
P IVQ G + H+D ++++FMEY GGSLA + G +E V +VYT +++ GL
Sbjct: 1093 --PNIVQYYGVEV-HRD-KVNIFMEYCEGGSLASLLDH-GRIEDEMVTQVYTFELLEGLA 1147
Query: 124 HLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANG------- 176
+LHQ G+VH D+K +N+LL +G +K DFG A+ V + ++ + G
Sbjct: 1148 YLHQSGVVHRDIKPENILLDFNGIIKYVDFGTARTVVGSRTRTVRNAAVQDFGVETKSLN 1207
Query: 177 ---GTPLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLIS 233
GTP++MAPE + + S++ + A D+W+LGC V+EMATGR PW +
Sbjct: 1208 EMMGTPMYMAPETI---SGSAVKGK-----LGADDVWALGCVVLEMATGRRPW-----SN 1254
Query: 234 ISNPMAAMFKIACGDGIPQFP--IHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLV 289
+ N A M+ +A G IPQ P + G FL RCLV+DP R+TA+ELL P+++
Sbjct: 1255 LDNEWAIMYHVAAG-RIPQLPNRDEMTAAGRAFLERCLVQDPTMRATAVELLIDPWMI 1311
>M3K4_MOUSE (O08648) Mitogen-activated protein kinase kinase kinase 4
(EC 2.7.1.37) (MAPK/ERK kinase kinase 4) (MEK kinase 4)
(MEKK 4)
Length = 1597
Score = 163 bits (412), Expect = 7e-40
Identities = 98/281 (34%), Positives = 144/281 (50%), Gaps = 22/281 (7%)
Query: 9 EWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAG-RDALENEVNILNTLNSSPSPY 67
+W +G +G G +G V+ ++ TG L +K+ + ++ + L P
Sbjct: 1331 KWQRGNKIGEGQYGKVYTCISVDTGELMAMKEIRFQPNDHKTIKETADELKIFEGIKHPN 1390
Query: 68 IVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQ 127
+V+ G + ++ +++FMEY G+L +VS L E V+R+YT+QI + LH+
Sbjct: 1391 LVRYFGVELHREE--MYIFMEYCDEGTLEEVSRL---GLQEHVIRLYTKQITVAINVLHE 1445
Query: 128 HGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVL 187
HGIVH D+K N+ L SSG +KL DFGC+ ++K N + + GT +MAPEV+
Sbjct: 1446 HGIVHRDIKGANIFLTSSGLIKLGDFGCSVKLKNNAQTMPGE--VNSTLGTAAYMAPEVI 1503
Query: 188 LMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACG 247
AADIWSLGC VIEM TG+ PW + + M+K+ G
Sbjct: 1504 TRAKGEGHG--------RAADIWSLGCVVIEMVTGKRPWHE-----YEHNFQIMYKVGMG 1550
Query: 248 DGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
P P S EG FL CL DPK R TA +LL+H F+
Sbjct: 1551 HK-PPIPERLSPEGKAFLSHCLESDPKIRWTASQLLDHAFV 1590
>MKH1_SCHPO (Q10407) MAP kinase kinase kinase mkh1 (EC 2.7.1.-)
Length = 1116
Score = 162 bits (410), Expect = 1e-39
Identities = 103/293 (35%), Positives = 153/293 (52%), Gaps = 35/293 (11%)
Query: 9 EWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKA---------HSEAGRDALENEVNILNT 59
+WMKG+++G G++G V LAMN +TG L VK+ H + +D +++ ++
Sbjct: 824 KWMKGELIGNGTYGKVFLAMNINTGELIAVKQVEIPQTINGRHDQLRKDIVDSINAEISM 883
Query: 60 LNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIV 119
+ IVQ LG ++ + + +F+EY+SGGS+ +G E +VR +RQ++
Sbjct: 884 IADLDHLNIVQYLG--FEKTETDISIFLEYVSGGSIGRCLRNYG-PFEEQLVRFVSRQVL 940
Query: 120 HGLYHLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTP 179
+GL +LH GI+H DLK N+L+ G K++DFG +K N+ N ++ ++ G+
Sbjct: 941 YGLSYLHSKGIIHRDLKADNLLIDFDGVCKISDFGISKH-SDNVYDNDAN---LSMQGSI 996
Query: 180 LWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMA 239
WMAPEV+ ND A D+WSLGC V+EM GR PW D+ I
Sbjct: 997 FWMAPEVI-------HNDHQGY--SAKVDVWSLGCVVLEMLAGRRPWSTDEAIQ------ 1041
Query: 240 AMFKIACGDGIPQFPIHF----SQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
AMFK+ P P S E FL C + R TA ELLNHPF+
Sbjct: 1042 AMFKLGTEKKAPPIPSELVSQVSPEAIQFLNACFTVNADVRPTAEELLNHPFM 1094
>STK4_HUMAN (Q13043) Serine/threonine-protein kinase 4 (EC 2.7.1.37)
(STE20-like kinase MST1) (MST-1) (Mammalian STE20-like
protein kinase 1) (Serine/threonine-protein kinase
Krs-2)
Length = 487
Score = 160 bits (405), Expect = 5e-39
Identities = 107/321 (33%), Positives = 168/321 (52%), Gaps = 31/321 (9%)
Query: 16 VGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIVQCLGTD 75
+G GS+GSV+ A++K TG + +K+ E+ + E++I+ +S P++V+ G+
Sbjct: 36 LGEGSYGSVYKAIHKETGQIVAIKQVPVESDLQEIIKEISIMQQCDS---PHVVKYYGSY 92
Query: 76 YDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQHGIVHCDL 135
+ + D L + MEY GS++D+ +L ED + + + GL +LH +H D+
Sbjct: 93 FKNTD--LWIVMEYCGAGSVSDIIRLRNKTLTEDEIATILQSTLKGLEYLHFMRKIHRDI 150
Query: 136 KCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLMKNNSSI 195
K N+LL + G+ KLADFG A ++ M K ++ I GTP WMAPEV+
Sbjct: 151 KAGNILLNTEGHAKLADFGVAGQLTD--TMAKRNTVI----GTPFWMAPEVI-------- 196
Query: 196 NDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACGDGIPQF-- 253
+ ADIWSLG T IEMA G+PP+ D +PM A+F I P F
Sbjct: 197 ---QEIGYNCVADIWSLGITAIEMAEGKPPYAD------IHPMRAIFMIPTNPP-PTFRK 246
Query: 254 PIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHYSASSPASVLEVHQF 313
P +S DF+++CLV+ P++R+TA +LL HPF+ S + V Q
Sbjct: 247 PELWSDNFTDFVKQCLVKSPEQRATATQLLQHPFVRSAKGVSILRDLINEAMDVKLKRQE 306
Query: 314 EDTYDDDDDDDDSDDNDELIS 334
+ D DD+++ + DE+ S
Sbjct: 307 SQQREVDQDDEENSEEDEMDS 327
>STK3_MOUSE (Q9JI10) Serine/threonine-protein kinase 3 (EC 2.7.1.37)
(STE20-like kinase MST2) (MST-2) (Mammalian STE20-like
protein kinase 2)
Length = 497
Score = 158 bits (399), Expect = 2e-38
Identities = 105/321 (32%), Positives = 172/321 (52%), Gaps = 32/321 (9%)
Query: 16 VGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIVQCLGTD 75
+G GS+GSV A++K +G + +K+ E+ + E++I+ +S PY+V+ G+
Sbjct: 33 LGEGSYGSVFKAIHKESGQVVAIKQVPVESDLQEIIKEISIMQQCDS---PYVVKYYGSY 89
Query: 76 YDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQHGIVHCDL 135
+ + D L + MEY GS++D+ +L ED + + + GL +LH +H D+
Sbjct: 90 FKNTD--LWIVMEYCGAGSVSDIIRLRNKTLTEDEIATILKSTLKGLEYLHFMRKIHRDI 147
Query: 136 KCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLMKNNSSI 195
K N+LL + G+ KLADFG A ++ M K ++ I GTP WMAPEV+
Sbjct: 148 KAGNILLNTEGHAKLADFGVAGQLTD--TMAKRNTVI----GTPFWMAPEVI-------- 193
Query: 196 NDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACGDGIPQF-- 253
+ ADIWSLG T IEMA G+PP+ D +PM A+F I P F
Sbjct: 194 ---QEIGYNCVADIWSLGITSIEMAEGKPPYAD------IHPMRAIFMIPTNPP-PTFRK 243
Query: 254 PIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHYSASSPASVLEVHQF 313
P +S + DF+++CLV+ P++R+TA +LL HPF+ + + A ++ +
Sbjct: 244 PELWSDDFTDFVKKCLVKSPEQRATATQLLQHPFIKNAKPVSILR-DLIAEAMEIKAKRH 302
Query: 314 EDTYDDDDDDDDSDDNDELIS 334
E+ + ++++++ D DEL S
Sbjct: 303 EEQQRELEEEEENSDEDELDS 323
>STK3_HUMAN (Q13188) Serine/threonine-protein kinase 3 (EC 2.7.1.37)
(STE20-like kinase MST2) (MST-2) (Mammalian STE20-like
protein kinase 2) (Serine/threonine-protein kinase
Krs-1)
Length = 491
Score = 157 bits (397), Expect = 4e-38
Identities = 105/321 (32%), Positives = 173/321 (53%), Gaps = 32/321 (9%)
Query: 16 VGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIVQCLGTD 75
+G GS+GSV A++K +G + +K+ E+ + E++I+ +S PY+V+ G+
Sbjct: 33 LGEGSYGSVFKAIHKESGQVVAIKQVPVESDLQEIIKEISIMQQCDS---PYVVKYYGSY 89
Query: 76 YDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQHGIVHCDL 135
+ + D L + MEY GS++D+ +L ED + + + GL +LH +H D+
Sbjct: 90 FKNTD--LWIVMEYCGAGSVSDIIRLRNKTLIEDEIATILKSTLKGLEYLHFMRKIHRDI 147
Query: 136 KCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLMKNNSSI 195
K N+LL + G+ KLADFG A ++ M K ++ I GTP WMAPEV+
Sbjct: 148 KAGNILLNTEGHAKLADFGVAGQLTD--TMAKRNTVI----GTPFWMAPEVI-------- 193
Query: 196 NDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACGDGIPQF-- 253
+ ADIWSLG T IEMA G+PP+ D +PM A+F I P F
Sbjct: 194 ---QEIGYNCVADIWSLGITSIEMAEGKPPYAD------IHPMRAIFMIPTNPP-PTFRK 243
Query: 254 PIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHHKHYSASSPASVLEVHQF 313
P +S + DF+++CLV++P++R+TA +LL HPF+ + + A ++ +
Sbjct: 244 PELWSDDFTDFVKKCLVKNPEQRATATQLLQHPFIKNAKPVSILR-DLITEAMEIKAKRH 302
Query: 314 EDTYDDDDDDDDSDDNDELIS 334
E+ + ++++++ D DEL S
Sbjct: 303 EEQQRELEEEEENSDEDELDS 323
>WIS4_SCHPO (O14299) MAP kinase kinase kinase wis4 (EC 2.7.1.-) (MAP
kinase kinase kinase wak1) (MAP kinase kinase kinase
wik1)
Length = 1401
Score = 157 bits (396), Expect = 5e-38
Identities = 101/291 (34%), Positives = 151/291 (51%), Gaps = 33/291 (11%)
Query: 10 WMKGKMVGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGR------DALENEVNILNTLNSS 63
W +G V G FG V+ +N TG L VK+ + R D + NE+ +L LN
Sbjct: 1037 WQQGHFVRSGMFGDVYTGVNMETGDLLAVKEIKLQDSRTFRSTVDQIHNEMTVLERLNH- 1095
Query: 64 PSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLY 123
P +V G + + ++++FME+ GGSLAD+ G +E+V++VY Q++ GL
Sbjct: 1096 --PNVVTYYGVEVHRE--KVYIFMEFCQGGSLADLL-AHGRIEDENVLKVYVVQLLEGLA 1150
Query: 124 HLHQHGIVHCDLKCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIAN----GGTP 179
++H I+H D+K N+LL G +K +DFG A V + I GTP
Sbjct: 1151 YIHSQHILHRDIKPANILLDHRGMIKYSDFGSALYVSPPTDPEVRYEDIQPELQHLAGTP 1210
Query: 180 LWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMA 239
++MAPE++L ++ DF A DIWSLGC ++EM TG PW + D N A
Sbjct: 1211 MYMAPEIIL---------GTKKGDFGAMDIWSLGCVILEMMTGSTPWSEMD-----NEWA 1256
Query: 240 AMFKIAC--GDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFL 288
M+ +A IPQ S DF+ +C RDP++R A++LL HP++
Sbjct: 1257 IMYHVAAMHTPSIPQNE-KISSLARDFIEQCFERDPEQRPRAVDLLTHPWI 1306
>STKA_HUMAN (O94804) Serine/threonine-protein kinase 10 (EC
2.7.1.37) (Lymphocyte-oriented kinase)
Length = 968
Score = 156 bits (395), Expect = 7e-38
Identities = 118/350 (33%), Positives = 176/350 (49%), Gaps = 30/350 (8%)
Query: 16 VGCGSFGSVHLAMNKSTGGLFVVKKAHSEAGRDALENEVNILNTLNSSPSPYIVQCLGTD 75
+G G+FG V+ A NK TG L K +++ + LE+ + + L + PYIV+ LG
Sbjct: 42 LGDGAFGKVYKAKNKETGALAAAKVIETKS-EEELEDYIVEIEILATCDHPYIVKLLGAY 100
Query: 76 YDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIVHGLYHLHQHGIVHCDL 135
Y D +L + +E+ GG++ + + L E ++V RQ++ L LH I+H DL
Sbjct: 101 Y--HDGKLWIMIEFCPGGAVDAIMLELDRGLTEPQIQVVCRQMLEALNFLHSKRIIHRDL 158
Query: 136 KCKNVLLASSGNVKLADFGCAKRVKKNMNMNKSSSCIIANGGTPLWMAPEVLLMKNNSSI 195
K NVL+ G+++LADFG + K + K S I GTP WMAPEV++ +
Sbjct: 159 KAGNVLMTLEGDIRLADFGVS--AKNLKTLQKRDSFI----GTPYWMAPEVVMCETMKDT 212
Query: 196 NDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNPMAAMFKIACGDGIPQF-P 254
+ + ADIWSLG T+IEMA PP + NPM + KIA D P
Sbjct: 213 PYDYK------ADIWSLGITLIEMAQIEPPHHE------LNPMRVLLKIAKSDPPTLLTP 260
Query: 255 IHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVSTTLTHH-KHYSASSPASVLEVHQF 313
+S E DFL+ L ++P+ R +A +LL HPF+ S T + A + A V+E +
Sbjct: 261 SKWSVEFRDFLKIALDKNPETRPSAAQLLEHPFVSSITSNKALRELVAEAKAEVME--EI 318
Query: 314 EDTYDDDDDDDDSDDNDELISPAGGNHFFITKELLCPQGTTMWQLEDSAS 363
ED D+ +++D D L NH + E+ P LE+S S
Sbjct: 319 EDGRDEGEEEDAVDAASTL-----ENHTQNSSEVSPPSLNADKPLEESPS 363
>M3K1_HUMAN (Q13233) Mitogen-activated protein kinase kinase kinase 1
(EC 2.7.1.37) (MAPK/ERK kinase kinase 1) (MEK kinase 1)
(MEKK 1) (Fragment)
Length = 1495
Score = 155 bits (393), Expect = 1e-37
Identities = 102/294 (34%), Positives = 157/294 (52%), Gaps = 33/294 (11%)
Query: 8 SEWMKGKMVGCGSFGSVHLAMNKSTGGLFVVKK--------AHSEAGRDALENEVNILNT 59
+EW+KG+ +G G+F S + A + TG L VK+ + E +AL E+ +++
Sbjct: 1224 TEWLKGQQIGLGAFSSCYQAQDVGTGTLMAVKQVTYVRNTSSEQEEVVEALREEIRMMSH 1283
Query: 60 LNSSPSPYIVQCLGTDYDHQDNQLHVFMEYMSGGSLADVSHKFGGSLNEDVVRVYTRQIV 119
LN P I++ LG + + L F+E+M+GGS+A + K+G + E VV YT Q++
Sbjct: 1284 LNH---PNIIRMLGATCEKSNYNL--FIEWMAGGSVAHLLSKYG-AFKESVVINYTEQLL 1337
Query: 120 HGLYHLHQHGIVHCDLKCKNVLLASSG-NVKLADFGCAKRV-KKNMNMNKSSSCIIANGG 177
GL +LH++ I+H D+K N+L+ S+G +++ADFG A R+ K + ++ G
Sbjct: 1338 RGLSYLHENQIIHRDVKGANLLIDSTGQRLRIADFGAAARLASKGTGAGEFQGQLL---G 1394
Query: 178 TPLWMAPEVLLMKNNSSINDESRVVDFAAADIWSLGCTVIEMATGRPPWVDDDLISISNP 237
T +MAPEVL + + D+WS+GC +IEMA +PPW + SN
Sbjct: 1395 TIAFMAPEVLRGQQYG-----------RSCDVWSVGCAIIEMACAKPPW---NAEKHSNH 1440
Query: 238 MAAMFKIACGDGIPQFPIHFSQEGFDFLRRCLVRDPKKRSTALELLNHPFLVST 291
+A +FKIA P P H S D RCL P+ R + ELL HP +T
Sbjct: 1441 LALIFKIASATTAPSIPSHLSPGLRDVALRCLELQPQDRPPSRELLKHPVFRTT 1494
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.135 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,272,978
Number of Sequences: 164201
Number of extensions: 2115272
Number of successful extensions: 16023
Number of sequences better than 10.0: 1784
Number of HSP's better than 10.0 without gapping: 1531
Number of HSP's successfully gapped in prelim test: 257
Number of HSP's that attempted gapping in prelim test: 9499
Number of HSP's gapped (non-prelim): 2917
length of query: 373
length of database: 59,974,054
effective HSP length: 112
effective length of query: 261
effective length of database: 41,583,542
effective search space: 10853304462
effective search space used: 10853304462
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 66 (30.0 bits)
Medicago: description of AC140549.19


