

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140544.4 - phase: 0 /pseudo
(305 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
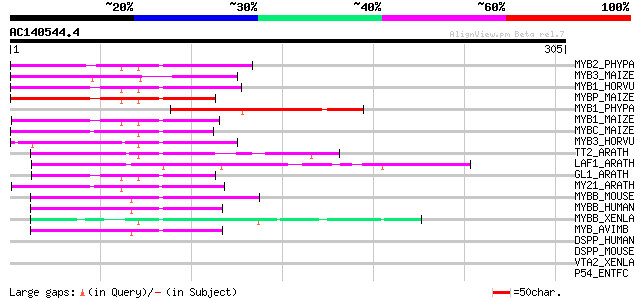
Score E
Sequences producing significant alignments: (bits) Value
MYB2_PHYPA (P80073) Myb-related protein Pp2 111 2e-24
MYB3_MAIZE (P20025) Myb-related protein Zm38 111 3e-24
MYB1_HORVU (P20026) Myb-related protein Hv1 110 3e-24
MYBP_MAIZE (P27898) Myb-related protein P 101 2e-21
MYB1_PHYPA (P80074) Myb-related protein Pp1 (Fragment) 100 8e-21
MYB1_MAIZE (P20024) Myb-related protein Zm1 88 3e-17
MYBC_MAIZE (P10290) Anthocyanin regulatory C1 protein 85 3e-16
MYB3_HORVU (P20027) Myb-related protein Hv33 75 2e-13
TT2_ARATH (Q9FJA2) TRANSPARENT TESTA 2 protein (Myb-related prot... 70 8e-12
LAF1_ARATH (Q9M0K4) Transcription factor LAF1 (Long after far-re... 68 2e-11
GL1_ARATH (P27900) Trichome differentiation protein GL1 (GLABROU... 63 8e-10
MY21_ARATH (Q9LK95) Transcription factor MYB21 (Myb-related prot... 60 9e-09
MYBB_MOUSE (P48972) Myb-related protein B (B-Myb) 53 8e-07
MYBB_HUMAN (P10244) Myb-related protein B (B-Myb) 52 2e-06
MYBB_XENLA (P52551) Myb-related protein B (B-Myb) (Myb-related p... 51 4e-06
MYB_AVIMB (P01104) Transforming protein Myb 48 3e-05
DSPP_HUMAN (Q9NZW4) Dentin sialophosphoprotein precursor [Contai... 42 0.002
DSPP_MOUSE (P97399) Dentin sialophosphoprotein precursor (Dentin... 39 0.016
VTA2_XENLA (P18709) Vitellogenin A2 precursor (VTG A2) [Contains... 38 0.036
P54_ENTFC (P13692) P54 protein precursor 38 0.036
>MYB2_PHYPA (P80073) Myb-related protein Pp2
Length = 421
Score = 111 bits (278), Expect = 2e-24
Identities = 65/143 (45%), Positives = 84/143 (58%), Gaps = 17/143 (11%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR PCCEKVGL++GPWTSEEDQKL+S+I +G WR++P L G+ R L
Sbjct: 1 MGRKPCCEKVGLRRGPWTSEEDQKLVSHITNNGLSCWRAIPK-----LAGLLRCGKSCRL 55
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG E+L+ + L N + A +LP RTDNEIKNYWNT LK
Sbjct: 56 RWTNYLRPDLKRGIFSEAEENLILDLHATLGN--RWSRIAAQLPGRTDNEIKNYWNTRLK 113
Query: 111 KRLTKMGIDPVTHKPKNETLLPE 133
KRL G+DP TH P ++ L +
Sbjct: 114 KRLRSQGLDPNTHLPLEDSKLDD 136
>MYB3_MAIZE (P20025) Myb-related protein Zm38
Length = 255
Score = 111 bits (277), Expect = 3e-24
Identities = 64/144 (44%), Positives = 77/144 (53%), Gaps = 35/144 (24%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKA---------------- 44
MGRSPCCEK +G WT EED++L++YI HG G WRSLP A
Sbjct: 1 MGRSPCCEKAHTNRGAWTKEEDERLVAYIRAHGEGCWRSLPKAAGLLRCGKSCRLRWINY 60
Query: 45 -VQDLRGVARAAD*DGLII*GRILRGE--SLVCKKNKPLFNFMHF*ATATRLPKRTDNEI 101
DL+ AD D LI+ L G SL+ A RLP RTDNEI
Sbjct: 61 LRPDLKRGNFTADEDDLIVKLHSLLGNKWSLI----------------AARLPGRTDNEI 104
Query: 102 KNYWNTHLKKRLTKMGIDPVTHKP 125
KNYWNTH++++L GIDPVTH+P
Sbjct: 105 KNYWNTHVRRKLLGRGIDPVTHRP 128
>MYB1_HORVU (P20026) Myb-related protein Hv1
Length = 267
Score = 110 bits (276), Expect = 3e-24
Identities = 64/137 (46%), Positives = 79/137 (56%), Gaps = 17/137 (12%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGRSPCCEK KG WT EED +L +YI+ HG G WRSLP A G+ R L
Sbjct: 1 MGRSPCCEKAHTNKGAWTKEEDDRLTAYIKAHGEGCWRSLPKAA-----GLLRCGKSCRL 55
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG + L+ K + L N + A RLP RTDNEIKNYWNTH++
Sbjct: 56 RWINYLRPDLKRGNFSHEEDELIIKLHSLLGN--KWSLIAGRLPGRTDNEIKNYWNTHIR 113
Query: 111 KRLTKMGIDPVTHKPKN 127
++LT GIDPVTH+ N
Sbjct: 114 RKLTSRGIDPVTHRAIN 130
>MYBP_MAIZE (P27898) Myb-related protein P
Length = 399
Score = 101 bits (252), Expect = 2e-21
Identities = 58/123 (47%), Positives = 74/123 (60%), Gaps = 17/123 (13%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR+PCCEKVGLK+G WT+EEDQ L +YI EHG GSWRSLP A G+ R L
Sbjct: 1 MGRTPCCEKVGLKRGRWTAEEDQLLANYIAEHGEGSWRSLPKNA-----GLLRCGKSCRL 55
Query: 61 ----II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLK 110
+ + RG E ++ K + L N + A+ LP RTDNEIKNYWN+HL
Sbjct: 56 RWINYLRADVKRGNISKEEEDIIIKLHATLGN--RWSLIASHLPGRTDNEIKNYWNSHLS 113
Query: 111 KRL 113
+++
Sbjct: 114 RQI 116
>MYB1_PHYPA (P80074) Myb-related protein Pp1 (Fragment)
Length = 449
Score = 99.8 bits (247), Expect = 8e-21
Identities = 55/112 (49%), Positives = 72/112 (64%), Gaps = 8/112 (7%)
Query: 89 TATRLPKRTDNEIKNYWNTHLKKRLTKMGIDPVTHKPK------NETLLPENHSKNVANL 142
+A +P+RTDNEIKNYWNTHLKKRL +MGIDPVTHK + +++P NL
Sbjct: 6 SAIAIPRRTDNEIKNYWNTHLKKRLMQMGIDPVTHKSTAAEELVHYSIIPGLRPVVSTNL 65
Query: 143 SHKAQWESARLEAEARLVRESKLRSQLQHQFGSNTSVFPSQSSSSSSNQVLN 194
+H +QW+SAR EAEARL R+S L S + TS+ +S+ S+ N V N
Sbjct: 66 THMSQWDSARAEAEARLSRQSSLTSPASDL--AQTSLEHQKSNLSTKNDVNN 115
>MYB1_MAIZE (P20024) Myb-related protein Zm1
Length = 340
Score = 87.8 bits (216), Expect = 3e-17
Identities = 51/124 (41%), Positives = 72/124 (57%), Gaps = 17/124 (13%)
Query: 2 GRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL- 60
GR+PCC KVGL +G WT +ED +L++YI++HGH +WR+LP +A G+ R L
Sbjct: 4 GRAPCCAKVGLNRGSWTPQEDMRLIAYIQKHGHTNWRALPKQA-----GLLRCGKSCRLR 58
Query: 61 ---II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKK 111
+ + RG E + + + L N + A LP RTDNEIKN WNTHLKK
Sbjct: 59 WINYLRPDLKRGNFTDEEEEAIIRLHGLLGN--KWSKIAACLPGRTDNEIKNVWNTHLKK 116
Query: 112 RLTK 115
++ +
Sbjct: 117 KVAQ 120
>MYBC_MAIZE (P10290) Anthocyanin regulatory C1 protein
Length = 273
Score = 84.7 bits (208), Expect = 3e-16
Identities = 51/119 (42%), Positives = 68/119 (56%), Gaps = 11/119 (9%)
Query: 1 MGRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL 60
MGR CC K G+K+G WTS+ED L +Y++ HG G WR +P KA LR ++ L
Sbjct: 1 MGRRACCAKEGVKRGAWTSKEDDALAAYVKAHGEGKWREVPQKA--GLRRCGKSCRLRWL 58
Query: 61 -II*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKR 112
+ I RG E L+ + ++ L N + A RLP RTDNEIKNYWN+ L +R
Sbjct: 59 NYLRPNIRRGNISYDEEDLIIRLHRLLGN--RWSLIAGRLPGRTDNEIKNYWNSTLGRR 115
>MYB3_HORVU (P20027) Myb-related protein Hv33
Length = 302
Score = 75.1 bits (183), Expect = 2e-13
Identities = 52/134 (38%), Positives = 73/134 (53%), Gaps = 13/134 (9%)
Query: 1 MGRSPCCEKVG---LKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD* 57
MGR P VG ++KG W+ EED+KL ++I HG G W S+P A + G +
Sbjct: 1 MGR-PSSGAVGQPKVRKGLWSPEEDEKLYNHIIRHGVGCWSSVPRLAALNRCGKSCRLRW 59
Query: 58 DGLII*GRILRG------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKK 111
+ + RG E + ++ L N + A+ LP RTDNEIKN+WN+ +KK
Sbjct: 60 INYLR-PDLKRGCFSQQEEDHIVALHQILGN--RWSQIASHLPGRTDNEIKNFWNSCIKK 116
Query: 112 RLTKMGIDPVTHKP 125
+L + GIDP THKP
Sbjct: 117 KLRQQGIDPATHKP 130
>TT2_ARATH (Q9FJA2) TRANSPARENT TESTA 2 protein (Myb-related protein
123) (AtMYB123) (Myb-related transcription factor
LBM2-like)
Length = 258
Score = 69.7 bits (169), Expect = 8e-12
Identities = 60/178 (33%), Positives = 81/178 (44%), Gaps = 29/178 (16%)
Query: 12 LKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGLII*GRILRG-- 69
L +G WT ED+ L YI HG G W +LP +A G + + G I RG
Sbjct: 14 LNRGAWTDHEDKILRDYITTHGEGKWSTLPNQAGLKRCGKSCRLRWKNYLRPG-IKRGNI 72
Query: 70 ----ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKRLTKMGIDPVTHKP 125
E L+ + + L N + A RLP RTDNEIKN+WN++L+KRL P
Sbjct: 73 SSDEEELIIRLHNLLGN--RWSLIAGRLPGRTDNEIKNHWNSNLRKRL-----------P 119
Query: 126 KNETLLPENHSKNVANLSHKAQWESARLEAEARLVRESK--LRSQLQHQFGSNTSVFP 181
K +T P+ + H E+ + +R SK L S L Q S+TS P
Sbjct: 120 KTQTKQPK-------RIKHSTNNENNVCVIRTKAIRCSKTLLFSDLSLQKKSSTSPLP 170
>LAF1_ARATH (Q9M0K4) Transcription factor LAF1 (Long after far-red
light protein 1) (Myb-related protein 18) (AtMYB18)
Length = 283
Score = 68.2 bits (165), Expect = 2e-11
Identities = 66/252 (26%), Positives = 107/252 (42%), Gaps = 29/252 (11%)
Query: 13 KKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGLII*GRILRGESL 72
+KG W+ EED+KL S+I +GH W ++P KA G + + G L+ + +
Sbjct: 11 RKGLWSPEEDEKLRSFILSYGHSCWTTVPIKAGLQRNGKSCRLRWINYLRPG--LKRDMI 68
Query: 73 VCKKNKPLFNF-----MHF*ATATRLPKRTDNEIKNYWNTHLKKRLTK----MGIDPVTH 123
++ + + F + A LP RTDNEIKNYW++HLKK+ K ++
Sbjct: 69 SAEEEETILTFHSSLGNKWSQIAKFLPGRTDNEIKNYWHSHLKKKWLKSQSLQDAKSISP 128
Query: 124 KPKNETLLPENHSKNVANLSHKAQWESARLEAEARLVRESKLRSQLQHQFGSNTSVFPSQ 183
+ + L +N L + RL E+K S Q G+N+ Q
Sbjct: 129 PSSSSSSLVACGKRNPETLISNHVFSFQRL-------LENKSSSPSQESNGNNS----HQ 177
Query: 184 SSSSSSNQVLNIKEEGEKEW--KGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMINVAE 241
SS+ I EW Y + + EF D + + TL M +V +
Sbjct: 178 CSSAP-----EIPRLFFSEWLSSSYPHTDYSSEFTDSKHSQAPNVEETLSAYEEMGDVDQ 232
Query: 242 EGFTNLLLDNSN 253
+ ++++NSN
Sbjct: 233 FHYNEMMINNSN 244
>GL1_ARATH (P27900) Trichome differentiation protein GL1 (GLABROUS1
protein)
Length = 228
Score = 63.2 bits (152), Expect = 8e-10
Identities = 41/111 (36%), Positives = 56/111 (49%), Gaps = 17/111 (15%)
Query: 13 KKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL----II*GRILR 68
KKG WT EED L+ Y+ HG G W + K G+ R L + + +
Sbjct: 15 KKGLWTVEEDNILMDYVLNHGTGQWNRIVRKT-----GLKRCGKSCRLRWMNYLSPNVNK 69
Query: 69 G------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKRL 113
G E L+ + +K L N + A R+P RTDN++KNYWNTHL K+L
Sbjct: 70 GNFTEQEEDLIIRLHKLLGN--RWSLIAKRVPGRTDNQVKNYWNTHLSKKL 118
>MY21_ARATH (Q9LK95) Transcription factor MYB21 (Myb-related protein
21) (AtMYB21) (Myb homolog 3) (AtMyb3)
Length = 226
Score = 59.7 bits (143), Expect = 9e-09
Identities = 40/123 (32%), Positives = 59/123 (47%), Gaps = 9/123 (7%)
Query: 2 GRSPCCEKVGLKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGL- 60
G S + ++KGPWT EED L++YI HG G W SL A L+ ++ L
Sbjct: 10 GGSGSSAEAEVRKGPWTMEEDLILINYIANHGDGVWNSLAKSA--GLKRTGKSCRLRWLN 67
Query: 61 -----II*GRILRGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKRLTK 115
+ G I E L+ + + + A LP RTDNEIKN+W T ++K + +
Sbjct: 68 YLRPDVRRGNITPEEQLIIMELHAKWG-NRWSKIAKHLPGRTDNEIKNFWRTRIQKYIKQ 126
Query: 116 MGI 118
+
Sbjct: 127 SDV 129
>MYBB_MOUSE (P48972) Myb-related protein B (B-Myb)
Length = 704
Score = 53.1 bits (126), Expect = 8e-07
Identities = 38/130 (29%), Positives = 61/130 (46%), Gaps = 6/130 (4%)
Query: 12 LKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGLII*GR----IL 67
L KGPWT EEDQK++ ++++G W + L R + L +
Sbjct: 81 LVKGPWTKEEDQKVIELVKKYGTKQWTLIAKHLKGRLGKQCRERWHNHLNPEVKKSCWTE 140
Query: 68 RGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKRLTKMGIDPVTHKPKN 127
+ ++C+ +K L N + A LP RTDN +KN+WN+ +K+++ G + K
Sbjct: 141 EEDRIICEAHKVLGN--RWAEIAKMLPGRTDNAVKNHWNSTIKRKVDTGGFPAESRDCKP 198
Query: 128 ETLLPENHSK 137
LL E K
Sbjct: 199 VYLLLELEDK 208
>MYBB_HUMAN (P10244) Myb-related protein B (B-Myb)
Length = 700
Score = 51.6 bits (122), Expect = 2e-06
Identities = 33/110 (30%), Positives = 55/110 (50%), Gaps = 6/110 (5%)
Query: 12 LKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGLII*GR----IL 67
L KGPWT EEDQK++ ++++G W + L R + L +
Sbjct: 81 LVKGPWTKEEDQKVIELVKKYGTKQWTLIAKHLKGRLGKQCRERWHNHLNPEVKKSCWTE 140
Query: 68 RGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKRLTKMG 117
+ ++C+ +K L N + A LP RTDN +KN+WN+ +K+++ G
Sbjct: 141 EEDRIICEAHKVLGN--RWAEIAKMLPGRTDNAVKNHWNSTIKRKVDTGG 188
>MYBB_XENLA (P52551) Myb-related protein B (B-Myb) (Myb-related
protein 1) (XMYB1)
Length = 743
Score = 50.8 bits (120), Expect = 4e-06
Identities = 57/238 (23%), Positives = 95/238 (38%), Gaps = 46/238 (19%)
Query: 12 LKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGLII*GRILRG-- 69
L KGPWT EED+K++ ++++G W T + LRG G+ R
Sbjct: 81 LVKGPWTKEEDEKVIELVKKYGTKHW----TLIAKQLRGRM-----------GKQCRERW 125
Query: 70 -----------------ESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKR 112
+ ++C+ +K L N + A LP RTDN +KN+WN+ +K++
Sbjct: 126 HNHLNPEVKKSSWTEEEDRIICQAHKVLGN--RWAEIAKLLPGRTDNAVKNHWNSTIKRK 183
Query: 113 LTKMGIDPVTHKPKNETLLPENH----SKNVANLSHKAQWESARLEAEARLVRESKLRSQ 168
+ G V + E + +N LS + SA + E + KL ++
Sbjct: 184 VETGGFLTVKASGQQEEREDSGYQAAEDQNHVLLSEPVE-RSANIPEEPSNILSPKLLTK 242
Query: 169 LQHQFGSNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAF 226
S S S+S I + ++W E L+ D+ME+ A+
Sbjct: 243 SP----GIRSEQESGGEGSNSESATAIVDSAPEKWM-VEYVNFLVPGSDIMESDPEAW 295
Score = 33.9 bits (76), Expect = 0.52
Identities = 11/27 (40%), Positives = 19/27 (69%)
Query: 14 KGPWTSEEDQKLLSYIEEHGHGSWRSL 40
K WT EED+ L + +++HG G W+++
Sbjct: 31 KVKWTPEEDETLKALVKKHGQGEWKTI 57
>MYB_AVIMB (P01104) Transforming protein Myb
Length = 382
Score = 47.8 bits (112), Expect = 3e-05
Identities = 29/110 (26%), Positives = 56/110 (50%), Gaps = 6/110 (5%)
Query: 12 LKKGPWTSEEDQKLLSYIEEHGHGSWRSLPTKAVQDLRGVARAAD*DGLII*GR----IL 67
L KGPWT EEDQ+++ +++++G W + + R + L +
Sbjct: 19 LNKGPWTKEEDQRVIEHVQKYGPKRWSDIAKHLKGRIGKQCRERWHNHLNPEVKKTSWTE 78
Query: 68 RGESLVCKKNKPLFNFMHF*ATATRLPKRTDNEIKNYWNTHLKKRLTKMG 117
+ ++ + +K L N + A LP RTDN +KN+WN+ +++++ + G
Sbjct: 79 EEDRIIYQAHKRLGN--RWAEIAKLLPGRTDNAVKNHWNSTMRRKVEQEG 126
>DSPP_HUMAN (Q9NZW4) Dentin sialophosphoprotein precursor [Contains:
Dentin phosphoprotein (Dentin phosphophoryn) (DPP);
Dentin sialoprotein (DSP)]
Length = 1253
Score = 41.6 bits (96), Expect = 0.002
Identities = 35/130 (26%), Positives = 59/130 (44%), Gaps = 2/130 (1%)
Query: 175 SNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEM 234
SN+S S SS SN+ N + + DS+ D ++S+ + SS +
Sbjct: 874 SNSSDSSDSSDSSDSNESSNSSDSSDSSNSSDSDSSDSSNSSDSSDSSNSSDSSESSNSS 933
Query: 235 TMINVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILN 294
N ++ ++ D+S+S + S S SG N+ D S S S + D+ N ++ +
Sbjct: 934 DNSNSSDSSNSSDSSDSSDSSNSSDSSNSGDSSNSSDSSDS--NSSDSSDSSNSSDSSDS 991
Query: 295 LVNSSPSDSS 304
+S SDSS
Sbjct: 992 SDSSDSSDSS 1001
Score = 40.0 bits (92), Expect = 0.007
Identities = 34/130 (26%), Positives = 62/130 (47%), Gaps = 2/130 (1%)
Query: 175 SNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEM 234
S++S S+SS S+ N + + DS+ D E+S+ + +S
Sbjct: 883 SDSSDSNESSNSSDSSDSSNSSDSDSSDSSNSSDSSDSSNSSDSSESSNSSDNSNSSDSS 942
Query: 235 TMINVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILN 294
+ ++ ++ D+SNSGD S S +S + N+ D S S SD + D+ + ++ +
Sbjct: 943 NSSDSSDSSDSSNSSDSSNSGDSSNSSDS-SDSNSSDSSDSSNSSD-SSDSSDSSDSSDS 1000
Query: 295 LVNSSPSDSS 304
+S+ SDSS
Sbjct: 1001 SDSSNSSDSS 1010
Score = 38.9 bits (89), Expect = 0.016
Identities = 36/132 (27%), Positives = 59/132 (44%), Gaps = 4/132 (3%)
Query: 175 SNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEM 234
SN+S S +SS SS+ + + DS++ + D ++S + SS
Sbjct: 727 SNSSDSDSSNSSDSSDSSDSSDSSNSSDSSDSSDSSNSSDSSDSSDSSDSSDSSNSSDSN 786
Query: 235 TMINVAEEGFTNLLLDNSNSGDLSLSPES--GGECNTCDGSGSGGGSDFNEDNKNYWNNI 292
N ++ ++ D+SNS D S S +S N+ D S S SD N + ++
Sbjct: 787 DSSNSSDSSDSSNSSDSSNSSDSSDSSDSSDSDSSNSSDSSNSSDSSD--SSNSSDSSDS 844
Query: 293 LNLVNSSPSDSS 304
+ +SS SDSS
Sbjct: 845 SDSSDSSDSDSS 856
Score = 38.1 bits (87), Expect = 0.027
Identities = 33/130 (25%), Positives = 60/130 (45%), Gaps = 1/130 (0%)
Query: 175 SNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEM 234
S+ S S SS SS ++ + K E + + + D +NS + SS +
Sbjct: 607 SSDSSDSSDSSDSSDSKSDSSKSESDSSDSDSKSDSSDSNSSDSSDNSDSSDSSNSSNSS 666
Query: 235 TMINVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILN 294
+ ++ ++ D+S+S D S S +S ++ + S S SD ++ + + ++ N
Sbjct: 667 DSSDSSDSSDSSSSSDSSSSSDSSNSSDSSDSSDSSNSSESSDSSDSSDSDSSDSSDSSN 726
Query: 295 LVNSSPSDSS 304
NSS SDSS
Sbjct: 727 -SNSSDSDSS 735
Score = 37.0 bits (84), Expect = 0.061
Identities = 32/123 (26%), Positives = 57/123 (46%), Gaps = 2/123 (1%)
Query: 182 SQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMINVAE 241
S SS SSN + + + DS++ + D ++S+ + SS + ++
Sbjct: 914 SSDSSDSSNSSDSSESSNSSDNSNSSDSSNSSDSSDSSDSSNSSDSSNSGDSSNSSDSSD 973
Query: 242 EGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILNLVNSSPS 301
++ D+SNS D S S +S ++ D S S SD + D+ + N+ + +S S
Sbjct: 974 SNSSD-SSDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSD-SSDSSDSSNSSDSSNSSDSS 1031
Query: 302 DSS 304
DSS
Sbjct: 1032 DSS 1034
Score = 36.6 bits (83), Expect = 0.080
Identities = 40/176 (22%), Positives = 73/176 (40%), Gaps = 12/176 (6%)
Query: 134 NHSKNVANLSHKAQWESARLEAEARLVRESKLRSQLQHQFGSNTSVFPSQSSSSSSNQVL 193
N S N ++ S + + +++ +S SN+S S+SS S+
Sbjct: 786 NDSSNSSDSSDSSNSSDSSNSSDSSDSSDSSDSDSSNSSDSSNSSDSSDSSNSSDSSDSS 845
Query: 194 NIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMINVAEEGFTNLLLDNSN 253
+ + + + DS++ + D ++S+ + SS + E ++ D+SN
Sbjct: 846 DSSDSSDSDSSNRSDSSNSSDSSDSSDSSNSSDSSDSSDSS---DSNESSNSSDSSDSSN 902
Query: 254 SGDLSLSPESGGECNTCDGSGSGGGSDFNE-----DNKNYWNNILNLVNSSPSDSS 304
S D +S N+ D S S SD +E DN N ++ + +S SDSS
Sbjct: 903 SSD----SDSSDSSNSSDSSDSSNSSDSSESSNSSDNSNSSDSSNSSDSSDSSDSS 954
Score = 35.8 bits (81), Expect = 0.14
Identities = 33/130 (25%), Positives = 60/130 (45%), Gaps = 4/130 (3%)
Query: 175 SNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEM 234
S++S S SS SSN + + DS++ + D ++S + SS +
Sbjct: 969 SDSSDSNSSDSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSNSSDSSNSS 1028
Query: 235 TMINVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILN 294
+ ++ ++ D+SNS D S +S ++ D SGS SD + D+ + ++ +
Sbjct: 1029 DSSDSSDSSDSSDSSDSSNSSD---SSDSSDSSDSSDSSGSSDSSD-SSDSSDSSDSSDS 1084
Query: 295 LVNSSPSDSS 304
+S SDSS
Sbjct: 1085 SDSSDSSDSS 1094
Score = 35.4 bits (80), Expect = 0.18
Identities = 32/130 (24%), Positives = 62/130 (47%), Gaps = 3/130 (2%)
Query: 175 SNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEM 234
SN+S S SS+S++ + + + + DS++ NSS + S+ +
Sbjct: 690 SNSSDSSDSSDSSNSSESSDSSDSSDSDSSDSSDSSNSNSSDSDSSNSSDSSDSSDSSDS 749
Query: 235 TMINVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILN 294
+ N ++ ++ ++S+S D S S +S N+ D + S SD + D+ N ++ +
Sbjct: 750 S--NSSDSSDSSDSSNSSDSSDSSDSSDSSDSSNSSDSNDSSNSSD-SSDSSNSSDSSNS 806
Query: 295 LVNSSPSDSS 304
+S SDSS
Sbjct: 807 SDSSDSSDSS 816
Score = 34.7 bits (78), Expect = 0.30
Identities = 31/125 (24%), Positives = 55/125 (43%), Gaps = 2/125 (1%)
Query: 182 SQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMINVAE 241
S SS SS+ + + DS+ + D ++S + SS + ++
Sbjct: 1084 SSDSSDSSDSSESSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSD 1143
Query: 242 EGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNE--DNKNYWNNILNLVNSS 299
++ D+S+S D S S +S ++ D S S SD NE D+ + ++ + +S
Sbjct: 1144 SSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSNESSDSSDSSDSSDSSNSSD 1203
Query: 300 PSDSS 304
SDSS
Sbjct: 1204 SSDSS 1208
Score = 33.9 bits (76), Expect = 0.52
Identities = 32/131 (24%), Positives = 53/131 (40%), Gaps = 8/131 (6%)
Query: 182 SQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDL--------MENSSMAFSSTLQHE 233
S SS SSN + DS+ + D NSS + S+ +
Sbjct: 743 SSDSSDSSNSSDSSDSSDSSNSSDSSDSSDSSDSSDSSNSSDSNDSSNSSDSSDSSNSSD 802
Query: 234 MTMINVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNIL 293
+ + + + + D+SNS D S S +S N+ D S S SD ++ + + ++
Sbjct: 803 SSNSSDSSDSSDSSDSDSSNSSDSSNSSDSSDSSNSSDSSDSSDSSDSSDSDSSNRSDSS 862
Query: 294 NLVNSSPSDSS 304
N +SS S S
Sbjct: 863 NSSDSSDSSDS 873
Score = 33.1 bits (74), Expect = 0.88
Identities = 31/125 (24%), Positives = 56/125 (44%), Gaps = 5/125 (4%)
Query: 182 SQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMINVAE 241
S SS SSN + + DS++ + D ++S + SS + + ++
Sbjct: 997 SSDSSDSSNSSDSSDSSDSSDSSNSSDSSNSSDSSDSSDSSDSSDSSDSSNSSDSSDSSD 1056
Query: 242 EGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNE--DNKNYWNNILNLVNSS 299
++ D+S S D S S +S ++ D S S SD +E D+ + ++ + +S
Sbjct: 1057 SSDSS---DSSGSSDSSDSSDSSDSSDSSDSSDSSDSSDSSESSDSSDSSDSSDSSDSSD 1113
Query: 300 PSDSS 304
SDSS
Sbjct: 1114 SSDSS 1118
Score = 30.8 bits (68), Expect = 4.4
Identities = 30/130 (23%), Positives = 60/130 (46%), Gaps = 3/130 (2%)
Query: 175 SNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEM 234
S++S S S+SSS+ + + + DS++ + D ++S + SS
Sbjct: 564 SDSSDSDSSDSNSSSDSDSSDSDSSDSSDSDSSDSSNSSDSSDSSDSSDSSDSSDSSDSK 623
Query: 235 TMINVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILN 294
+ + +E ++ D+ + S S +S ++ D S S SD + D+ + ++ +
Sbjct: 624 SDSSKSESDSSD--SDSKSDSSDSNSSDSSDNSDSSDSSNSSNSSD-SSDSSDSSDSSSS 680
Query: 295 LVNSSPSDSS 304
+SS SDSS
Sbjct: 681 SDSSSSSDSS 690
Score = 30.4 bits (67), Expect = 5.7
Identities = 39/153 (25%), Positives = 59/153 (38%), Gaps = 28/153 (18%)
Query: 175 SNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEM 234
SN+S S SS S+ N + + DS+ D NSS + S+ +
Sbjct: 762 SNSSDSSDSSDSSDSSDSSNSSDSNDSS--NSSDSSDSSNSSD-SSNSSDSSDSSDSSDS 818
Query: 235 TMINVAEEGFTNLLLDNSNSGD-----------------------LSLSPESGGECNTCD 271
N ++ ++ D+SNS D S S +S N+ D
Sbjct: 819 DSSNSSDSSNSSDSSDSSNSSDSSDSSDSSDSSDSDSSNRSDSSNSSDSSDSSDSSNSSD 878
Query: 272 GSGSGGGSDFNEDNKNYWNNILNLVNSSPSDSS 304
S S SD NE + + ++ + NSS SDSS
Sbjct: 879 SSDSSDSSDSNESSNS--SDSSDSSNSSDSDSS 909
Score = 29.6 bits (65), Expect = 9.7
Identities = 27/125 (21%), Positives = 50/125 (39%), Gaps = 15/125 (12%)
Query: 182 SQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMINVAE 241
S SS SSN + + DS+ + D ++S + SS + ++
Sbjct: 1123 SSDSSDSSNSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSD 1182
Query: 242 EGFTNLLLDNSNSGDLSLSPESGGECNTCD---------------GSGSGGGSDFNEDNK 286
++ D+S+S D S S +S ++ D G+G+ GSD + D++
Sbjct: 1183 SNESSDSSDSSDSSDSSNSSDSSDSSDSSDSTSDSNDESDSQSKSGNGNNNGSDSDSDSE 1242
Query: 287 NYWNN 291
+N
Sbjct: 1243 GSDSN 1247
Score = 29.6 bits (65), Expect = 9.7
Identities = 32/130 (24%), Positives = 53/130 (40%), Gaps = 7/130 (5%)
Query: 182 SQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLME-----NSSMAFSSTLQHEMTM 236
S SS SSN N DS+ D NSS + S
Sbjct: 923 SSDSSESSNSSDNSNSSDSSNSSDSSDSSDSSNSSDSSNSGDSSNSSDSSDSNSSDSSDS 982
Query: 237 INVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNE--DNKNYWNNILN 294
N ++ ++ D+S+S D S S +S ++ D S S S+ ++ D+ + ++ +
Sbjct: 983 SNSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSNSSDSSNSSDSSDSSDSSDSSDS 1042
Query: 295 LVNSSPSDSS 304
+S+ SDSS
Sbjct: 1043 SDSSNSSDSS 1052
>DSPP_MOUSE (P97399) Dentin sialophosphoprotein precursor (Dentin
matrix protein-3) (DMP-3) [Contains: Dentin
phosphoprotein (Dentin phosphophoryn) (DPP); Dentin
sialoprotein (DSP)]
Length = 934
Score = 38.9 bits (89), Expect = 0.016
Identities = 35/130 (26%), Positives = 59/130 (44%), Gaps = 1/130 (0%)
Query: 175 SNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEM 234
S++S S SSSSSN + DS+ + D +S + SS
Sbjct: 670 SDSSDSSSSDSSSSSNSSDSSDSSDSSSSSDSSDSSDSSDSSDSSGSSDSSDSSASSDSS 729
Query: 235 TMINVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILN 294
+ + ++ ++ D+S+S D S S ES N+ D S S SD + D+ + ++ +
Sbjct: 730 SSSDSSDSSSSSDSSDSSDSSDSSDSSESSDSSNSSDSSDSSDSSD-SSDSSDSSDSSDS 788
Query: 295 LVNSSPSDSS 304
+S+ SDSS
Sbjct: 789 SDSSNSSDSS 798
Score = 34.3 bits (77), Expect = 0.39
Identities = 31/123 (25%), Positives = 53/123 (42%), Gaps = 1/123 (0%)
Query: 182 SQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMINVAE 241
S SS SSN + + DS+ + D +S + SS N ++
Sbjct: 755 SSESSDSSNSSDSSDSSDSSDSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSDSSNSSD 814
Query: 242 EGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILNLVNSSPS 301
++ D+S+S D S S +S ++ D S S SD + D+ + ++ + +S S
Sbjct: 815 SSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSD-SSDSSDSSDSSDSSDSSDSS 873
Query: 302 DSS 304
DSS
Sbjct: 874 DSS 876
Score = 32.7 bits (73), Expect = 1.1
Identities = 29/123 (23%), Positives = 55/123 (44%), Gaps = 1/123 (0%)
Query: 182 SQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMINVAE 241
S SSS SS+ + + DS+ + + ++S + SS + ++
Sbjct: 728 SSSSSDSSDSSSSSDSSDSSDSSDSSDSSESSDSSNSSDSSDSSDSSDSSDSSDSSDSSD 787
Query: 242 EGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILNLVNSSPS 301
++ D+S+S D S S +S ++ D S S SD + D+ + ++ + +S S
Sbjct: 788 SSDSSNSSDSSDSSDSSDSSDSSNSSDSSDSSDSSDSSD-SSDSSDSSDSSDSSDSSDSS 846
Query: 302 DSS 304
DSS
Sbjct: 847 DSS 849
Score = 31.2 bits (69), Expect = 3.3
Identities = 28/108 (25%), Positives = 47/108 (42%), Gaps = 2/108 (1%)
Query: 182 SQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMINVAE 241
S +SS SS+ + + DS+ + D ++S + SS + ++
Sbjct: 809 SSNSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSDSSD 868
Query: 242 EGFTNLLLDNSNSGDLSLSPESGGECNTCDG-SGSGGG-SDFNEDNKN 287
++ D+SNS D S S ++ DG S SG G SD N D+ +
Sbjct: 869 SSDSSDSSDSSNSSDSSDSDSKDSSSDSSDGDSKSGNGNSDSNSDSNS 916
Score = 30.4 bits (67), Expect = 5.7
Identities = 30/131 (22%), Positives = 55/131 (41%), Gaps = 6/131 (4%)
Query: 174 GSNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHE 233
G S + S SS+ + E + + D + D ++S + SS
Sbjct: 519 GDGDSESEDKDESDSSDHDNSSDSESKSDSSDSSDDS-----SDSSDSSDSSDSSDSSDS 573
Query: 234 MTMINVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNIL 293
+ ++ +N D+S+S S S +S C++ D S S SD + D+ + ++
Sbjct: 574 SDSSDSSDSSDSNSSSDSSDSSGSSDSSDSSDTCDSSDSSDSSDSSD-SSDSSDSSDSSD 632
Query: 294 NLVNSSPSDSS 304
+ +S SDSS
Sbjct: 633 SSDSSDSSDSS 643
>VTA2_XENLA (P18709) Vitellogenin A2 precursor (VTG A2) [Contains:
Lipovitellin I; Lipovitellin II; Phosvitin]
Length = 1807
Score = 37.7 bits (86), Expect = 0.036
Identities = 49/182 (26%), Positives = 75/182 (40%), Gaps = 16/182 (8%)
Query: 107 THLKKRLTK-MGIDPVTHKPKNETLLPENHSKNVANLSHKAQWESARLEAEARLVRESKL 165
T + KRL +GID + K NET L + K K + + RL+AE +V K
Sbjct: 1074 TAVTKRLKMILGIDE-SRKDTNETALYRSKQKK------KNKIHNRRLDAE--VVEARKQ 1124
Query: 166 RSQLQHQFGSNTSVFPSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMA 225
+S L S++S S SSSSSS+ + Y + E + S +
Sbjct: 1125 QSSLSSSSSSSSSSSSSSSSSSSSS---SSSSPSSSSSSSYSKRSKRREHNPHHQRESSS 1181
Query: 226 FSSTLQHEMTMINVAEEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDN 285
SS Q++ + +E + S+S S S S ++ S S S+ N +
Sbjct: 1182 SSSQEQNKKRNL---QENRKHGQKGMSSSSSSSSSSSSSSSSSSSSSSSSSSSSEENRPH 1238
Query: 286 KN 287
KN
Sbjct: 1239 KN 1240
>P54_ENTFC (P13692) P54 protein precursor
Length = 516
Score = 37.7 bits (86), Expect = 0.036
Identities = 41/184 (22%), Positives = 74/184 (39%), Gaps = 24/184 (13%)
Query: 128 ETLLPENHSKNVANLSHKAQWESARLEAEARLVRES-------KLRSQLQHQFGSNTSVF 180
E E+ ++ +A+ E AR+ +ARL ++ K + + Q + +
Sbjct: 206 EQATAEDKKADLNRKKAEAEAEQARIREQARLAEQARQQAAQEKAEKEAREQAAAQAAQT 265
Query: 181 PSQSSSSSSNQVLNIKEEGEKEWKGYEDSTHLLEFKDLMENSSMAFSSTLQHEMTMINVA 240
+ SS+S++ + + + +E K E ST E S+ SST
Sbjct: 266 QALSSASTTTESSSAAQSSSEESKAPESST--------TEESTSTESST---------TT 308
Query: 241 EEGFTNLLLDNSNSGDLSLSPESGGECNTCDGSGSGGGSDFNEDNKNYWNNILNLVNSSP 300
E T S+S + S PES + +T + S S N N N+ N N+S
Sbjct: 309 ENSSTGSSSTESSSTEESTVPESSTQESTPANTESSSSSSNTNVNNNTNNSTNNSTNNST 368
Query: 301 SDSS 304
++++
Sbjct: 369 TNNN 372
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.131 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 36,099,028
Number of Sequences: 164201
Number of extensions: 1487641
Number of successful extensions: 5774
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 44
Number of HSP's successfully gapped in prelim test: 38
Number of HSP's that attempted gapping in prelim test: 5553
Number of HSP's gapped (non-prelim): 209
length of query: 305
length of database: 59,974,054
effective HSP length: 110
effective length of query: 195
effective length of database: 41,911,944
effective search space: 8172829080
effective search space used: 8172829080
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 65 (29.6 bits)
Medicago: description of AC140544.4


