

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140034.3 + phase: 0
(199 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
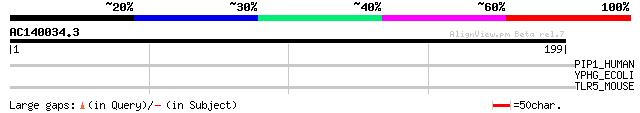
Score E
Sequences producing significant alignments: (bits) Value
PIP1_HUMAN (Q9H074) Polyadenylate-binding protein-interacting pr... 31 1.6
YPHG_ECOLI (P76585) Hypothetical protein yphG 31 2.1
TLR5_MOUSE (Q9JLF7) Toll-like receptor 5 precursor 30 3.7
>PIP1_HUMAN (Q9H074) Polyadenylate-binding protein-interacting
protein 1 (Poly(A) binding protein-interacting protein
1) (PABP-interacting protein 1) (PAIP-1)
Length = 479
Score = 31.2 bits (69), Expect = 1.6
Identities = 35/143 (24%), Positives = 60/143 (41%), Gaps = 21/143 (14%)
Query: 62 GYISYPRLRDFYTSYLGRANVLAGTEDPEELEELARVRTYCVRWYLMYLVGCLLFDDRSN 121
G YP L ++ +L G+ + E +E+ A L GC+ DD
Sbjct: 151 GCEDYPTLSEYVQDFLNHLTEQPGSFETE-IEQFAET-----------LNGCVTTDDALQ 198
Query: 122 KRIELIY--LTTMAD-GYAGMR--NYSWGGMTLAYLYGE----LADACRPGHRALGGSVT 172
+ +ELIY T++ + Y G R NY +T++ G L CR + +
Sbjct: 199 ELVELIYQQATSIPNFSYMGARLCNYLSHHLTISPQSGNFRQLLLQRCRTEYEVKDQAAK 258
Query: 173 LLTVRKLKFYIYVNFIFLLYFSV 195
V + +F+ +V F+ LY ++
Sbjct: 259 GDEVTRKRFHAFVLFLGELYLNL 281
>YPHG_ECOLI (P76585) Hypothetical protein yphG
Length = 1124
Score = 30.8 bits (68), Expect = 2.1
Identities = 18/62 (29%), Positives = 31/62 (49%), Gaps = 6/62 (9%)
Query: 89 PEELEELARVRTYCVRWYLMYLVGCLLFDDRS-NKRIE-----LIYLTTMADGYAGMRNY 142
P LEE+A + + W+ +L+ C ++ RS NK I + ADG+ G+ +
Sbjct: 759 PNTLEEVAALESIEECWFARHLLACFYYNKRSYNKAIAFWQRCVEMSPEFADGWRGLAIH 818
Query: 143 SW 144
+W
Sbjct: 819 AW 820
>TLR5_MOUSE (Q9JLF7) Toll-like receptor 5 precursor
Length = 859
Score = 30.0 bits (66), Expect = 3.7
Identities = 16/45 (35%), Positives = 25/45 (55%), Gaps = 8/45 (17%)
Query: 89 PEELEELARVRTYCVRWYLMYLVGCLLFDDRSNKRIELIYLTTMA 133
PE+L++ V W+L L GC+L +++ KR I L T+A
Sbjct: 820 PEDLQD--------VGWFLDKLSGCILKEEKGKKRSSSIQLRTIA 856
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.142 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,328,582
Number of Sequences: 164201
Number of extensions: 1002081
Number of successful extensions: 2458
Number of sequences better than 10.0: 3
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 2457
Number of HSP's gapped (non-prelim): 3
length of query: 199
length of database: 59,974,054
effective HSP length: 105
effective length of query: 94
effective length of database: 42,732,949
effective search space: 4016897206
effective search space used: 4016897206
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC140034.3


