

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC140034.13 + phase: 0 /partial
(433 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
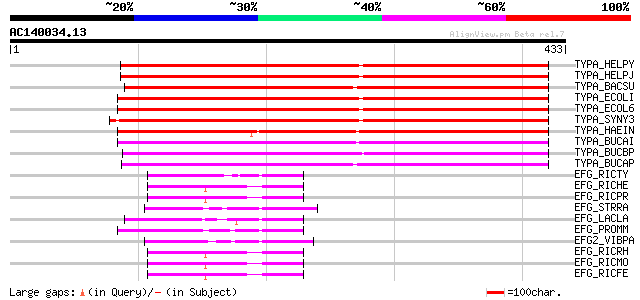
Score E
Sequences producing significant alignments: (bits) Value
TYPA_HELPY (O25225) GTP-binding protein TypA/BipA homolog 289 8e-78
TYPA_HELPJ (Q9ZLZ3) GTP-binding protein TypA/BipA homolog 286 5e-77
TYPA_BACSU (O07631) GTP-binding protein typA/bipA homolog 283 5e-76
TYPA_ECOLI (P32132) GTP-binding protein typA/BipA (Tyrosine phos... 280 7e-75
TYPA_ECOL6 (Q9EXN7) GTP-binding protein typA/BipA (Tyrosine phos... 280 7e-75
TYPA_SYNY3 (P72749) GTP-binding protein TypA/BipA homolog 275 1e-73
TYPA_HAEIN (P44910) GTP-binding protein typA/bipA homolog 270 7e-72
TYPA_BUCAI (P57508) GTP-binding protein TypA/BipA homolog 243 9e-64
TYPA_BUCBP (Q89AC9) GTP-binding protein TypA/BipA homolog 237 4e-62
TYPA_BUCAP (Q8K9C8) GTP-binding protein TypA/BipA homolog 233 7e-61
EFG_RICTY (Q8KTB2) Elongation factor G (EF-G) 51 5e-06
EFG_RICHE (Q8KTB4) Elongation factor G (EF-G) 51 5e-06
EFG_RICPR (P41084) Elongation factor G (EF-G) 51 7e-06
EFG_STRRA (P29541) Elongation factor G (EF-G) (Fragment) 50 1e-05
EFG_LACLA (Q9CDG1) Elongation factor G (EF-G) 50 1e-05
EFG_PROMM (Q7V501) Elongation factor G (EF-G) 50 1e-05
EFG2_VIBPA (Q87M30) Elongation factor G 2 (EF-G 2) 50 1e-05
EFG_RICRH (Q8KTB7) Elongation factor G (EF-G) 49 2e-05
EFG_RICMO (Q8KTB6) Elongation factor G (EF-G) 49 2e-05
EFG_RICFE (Q8KTA8) Elongation factor G (EF-G) 49 2e-05
>TYPA_HELPY (O25225) GTP-binding protein TypA/BipA homolog
Length = 599
Score = 289 bits (740), Expect = 8e-78
Identities = 144/334 (43%), Positives = 217/334 (64%), Gaps = 2/334 (0%)
Query: 87 EDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPT 146
E+G++ K++ G + A AGDI++IAG ++ +G +V L + L+ PT
Sbjct: 246 ENGRITKLIGFLGLARTEIENAYAGDIVAIAGFNAMDVGDSVVDPANPMPLDPMHLEEPT 305
Query: 147 ISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGELQLG 206
+S+ F VNDSPLAG +G H+T K+ DRL+ E +TN+A+ F+V GRGELQ+
Sbjct: 306 MSVYFAVNDSPLAGLEGKHVTANKLKDRLLKEMQTNIAMKCEEMGEGKFKVSGRGELQIT 365
Query: 207 ILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVMEALSHRRAEIT 266
IL EN+RREGFE S+S P+V+ K + G K EP E + I+ + G ++E L R+AE+
Sbjct: 366 ILAENLRREGFEFSISRPEVIIKEENGVKCEPFEHLVIDTPQDFSGAIIERLGKRKAEMK 425
Query: 267 DMGPVAGTVGRTRLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKFRGPLGNVRKG 326
M P++ G TRL P+RGL+GYRS F ++T+G G M+ +FL++ F G + + + G
Sbjct: 426 AMNPMSD--GYTRLEFEIPARGLIGYRSEFLTDTKGEGVMNHSFLEFRPFSGSVESRKNG 483
Query: 327 VLVSVGFGSITLHALMSLEARGTLFVSPGMEAYDGMIVGEHSRDTDLDVNPVRAKQLTNV 386
L+S+ G T +L +++ RGTLF++P + Y GM++GEHSRD DLDVNP+++K LTN+
Sbjct: 484 ALISMENGEATAFSLFNIQERGTLFINPQTKVYVGMVIGEHSRDNDLDVNPIKSKHLTNM 543
Query: 387 RSVNKDDTVKLIPPRLMTLEEAIGYVASDELIEV 420
R+ DD +KL PPR M LE A+ ++ DE++EV
Sbjct: 544 RASGSDDAIKLTPPRTMVLERALEWIEEDEILEV 577
>TYPA_HELPJ (Q9ZLZ3) GTP-binding protein TypA/BipA homolog
Length = 599
Score = 286 bits (733), Expect = 5e-77
Identities = 142/334 (42%), Positives = 216/334 (64%), Gaps = 2/334 (0%)
Query: 87 EDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPT 146
E+G++ K++ G + A AGDI+++AG ++ +G +V L + L+ PT
Sbjct: 246 ENGRITKLIGFLGLARTEIENAYAGDIVALAGFNAMDVGDSVVDPTNPMPLDPMHLEEPT 305
Query: 147 ISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGELQLG 206
+S+ F VNDSPLAG +G H+T K+ DRL+ E +TN+A+ F+V GRGELQ+
Sbjct: 306 MSVYFAVNDSPLAGLEGKHVTANKLKDRLLKEMQTNIAMKCEEMGEGKFKVSGRGELQIT 365
Query: 207 ILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVMEALSHRRAEIT 266
IL EN+RREGFE S+S P+V+ K + G K EP E + I+ + G ++E L R+AE+
Sbjct: 366 ILAENLRREGFEFSISRPEVIIKEENGVKCEPFEHLVIDTPQDFSGAIIERLGKRKAEMK 425
Query: 267 DMGPVAGTVGRTRLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKFRGPLGNVRKG 326
M P++ G TRL P+RGL+GYRS F ++T+G G M+ +FL++ F G + + + G
Sbjct: 426 AMNPMSD--GYTRLEFEIPARGLIGYRSEFLTDTKGEGVMNHSFLEFRPFSGSVESRKNG 483
Query: 327 VLVSVGFGSITLHALMSLEARGTLFVSPGMEAYDGMIVGEHSRDTDLDVNPVRAKQLTNV 386
L+S+ G T +L +++ RG LF++P + Y GM++GEHSRD DLDVNP+++K LTN+
Sbjct: 484 ALISMENGEATAFSLFNIQERGALFINPQTKVYVGMVIGEHSRDNDLDVNPIKSKHLTNM 543
Query: 387 RSVNKDDTVKLIPPRLMTLEEAIGYVASDELIEV 420
R+ DD +KL PPR M LE A+ ++ DE++EV
Sbjct: 544 RASGSDDAIKLTPPRTMVLERALEWIEEDEILEV 577
>TYPA_BACSU (O07631) GTP-binding protein typA/bipA homolog
Length = 612
Score = 283 bits (725), Expect = 5e-76
Identities = 150/333 (45%), Positives = 215/333 (64%), Gaps = 4/333 (1%)
Query: 90 KVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISM 149
+V K+ +G V + A AGD+++++G+ ++G TV V+ LP + +D PT+ M
Sbjct: 252 RVTKIFGFQGLKRVEIEEAKAGDLVAVSGMEDINVGETVCPVDHQDPLPVLRIDEPTLQM 311
Query: 150 TFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMS-ETFEVQGRGELQLGIL 208
TF VN+SP AGR+G ++T KI +RL ++ +T++++ V P S + + V GRGEL L IL
Sbjct: 312 TFVVNNSPFAGREGKYVTARKIEERLQSQLQTDVSLRVEPTASPDAWVVSGRGELHLSIL 371
Query: 209 IENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVMEALSHRRAEITDM 268
IENMRREG+EL VS P+V+ K G + EP+E V I+V +EH G VME++ R+ E+ DM
Sbjct: 372 IENMRREGYELQVSKPEVIIKEIDGVRCEPVERVQIDVPEEHTGSVMESMGARKGEMVDM 431
Query: 269 GPVAGTVGRTRLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKFR-GPLGNVRKGV 327
+ G+ RL T PSRGL+GY + F S TRG G ++ F Y+ + G +G R+GV
Sbjct: 432 --INNGNGQVRLIFTVPSRGLIGYSTEFLSLTRGFGILNHTFDSYQPMQAGQVGGRRQGV 489
Query: 328 LVSVGFGSITLHALMSLEARGTLFVSPGMEAYDGMIVGEHSRDTDLDVNPVRAKQLTNVR 387
LVS+ G T + + +E RG +FV PG E Y+GMIVGEH+RD DL VN + KQ TNVR
Sbjct: 490 LVSMENGKATSYGIQGIEDRGVIFVEPGTEVYEGMIVGEHNRDNDLVVNVSKMKQQTNVR 549
Query: 388 SVNKDDTVKLIPPRLMTLEEAIGYVASDELIEV 420
S KD T + R+M+LEE++ Y+ DE EV
Sbjct: 550 SATKDQTTTIKKARIMSLEESLEYLNEDEYCEV 582
>TYPA_ECOLI (P32132) GTP-binding protein typA/BipA (Tyrosine
phosphorylated protein A)
Length = 607
Score = 280 bits (715), Expect = 7e-75
Identities = 153/338 (45%), Positives = 211/338 (62%), Gaps = 4/338 (1%)
Query: 85 KIEDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDP 144
K + KV KV+ G + TD A AGDI++I GL +I TV + + ALP + +D
Sbjct: 245 KTRNAKVGKVLGHLGLERIETDLAEAGDIVAITGLGELNISDTVCDTQNVEALPALSVDE 304
Query: 145 PTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGM-SETFEVQGRGEL 203
PT+SM F VN SP G++G +T +I DRL E N+A+ V ++ F V GRGEL
Sbjct: 305 PTVSMFFCVNTSPFCGKEGKFVTSRQILDRLNKELVHNVALRVEETEDADAFRVSGRGEL 364
Query: 204 QLGILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVMEALSHRRA 263
L +LIENMRREGFEL+VS PKV+++ G+K EP E VT++V ++H G VM+AL R+
Sbjct: 365 HLSVLIENMRREGFELAVSRPKVIFREIDGRKQEPYENVTLDVEEQHQGSVMQALGERKG 424
Query: 264 EITDMGPVAGTVGRTRLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKFR-GPLGN 322
++ +M P GR RL PSRGL+G+RS F + T GTG ++ F Y+ R G +G
Sbjct: 425 DLKNMNPDGK--GRVRLDYVIPSRGLIGFRSEFMTMTSGTGLLYSTFSHYDDVRPGEVGQ 482
Query: 323 VRKGVLVSVGFGSITLHALMSLEARGTLFVSPGMEAYDGMIVGEHSRDTDLDVNPVRAKQ 382
+ GVL+S G G AL L+ RG LF+ G E Y+G I+G HSR DL VN + K+
Sbjct: 483 RQNGVLISNGQGKAVAFALFGLQDRGKLFLGHGAEVYEGQIIGIHSRSNDLTVNCLTGKK 542
Query: 383 LTNVRSVNKDDTVKLIPPRLMTLEEAIGYVASDELIEV 420
LTN+R+ D+ V L+PP MTLE+A+ ++ DEL+EV
Sbjct: 543 LTNMRASGTDEAVVLVPPIRMTLEQALEFIDDDELVEV 580
>TYPA_ECOL6 (Q9EXN7) GTP-binding protein typA/BipA (Tyrosine
phosphorylated protein A)
Length = 607
Score = 280 bits (715), Expect = 7e-75
Identities = 153/338 (45%), Positives = 211/338 (62%), Gaps = 4/338 (1%)
Query: 85 KIEDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDP 144
K + KV KV+ G + TD A AGDI++I GL +I TV + + ALP + +D
Sbjct: 245 KTRNAKVGKVLGHLGLERIETDLAEAGDIVAITGLGELNISDTVCDTQNVEALPALSVDE 304
Query: 145 PTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGM-SETFEVQGRGEL 203
PT+SM F VN SP G++G +T +I DRL E N+A+ V ++ F V GRGEL
Sbjct: 305 PTVSMFFCVNTSPFCGKEGKFVTSRQILDRLNKELVHNVALRVEETEDADAFRVSGRGEL 364
Query: 204 QLGILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVMEALSHRRA 263
L +LIENMRREGFEL+VS PKV+++ G+K EP E VT++V ++H G VM+AL R+
Sbjct: 365 HLSVLIENMRREGFELAVSRPKVIFREIDGRKQEPYENVTLDVEEQHQGSVMQALGERKG 424
Query: 264 EITDMGPVAGTVGRTRLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKFR-GPLGN 322
++ +M P GR RL PSRGL+G+RS F + T GTG ++ F Y+ R G +G
Sbjct: 425 DLKNMNPDGK--GRVRLDYVIPSRGLIGFRSEFMTMTSGTGLLYSTFSHYDDVRPGEVGQ 482
Query: 323 VRKGVLVSVGFGSITLHALMSLEARGTLFVSPGMEAYDGMIVGEHSRDTDLDVNPVRAKQ 382
+ GVL+S G G AL L+ RG LF+ G E Y+G I+G HSR DL VN + K+
Sbjct: 483 RQNGVLISNGQGKAVAFALFGLQDRGKLFLGHGAEVYEGQIIGIHSRSNDLTVNCLTGKK 542
Query: 383 LTNVRSVNKDDTVKLIPPRLMTLEEAIGYVASDELIEV 420
LTN+R+ D+ V L+PP MTLE+A+ ++ DEL+EV
Sbjct: 543 LTNMRASGTDEAVVLVPPIRMTLEQALEFIDDDELVEV 580
>TYPA_SYNY3 (P72749) GTP-binding protein TypA/BipA homolog
Length = 597
Score = 275 bits (704), Expect = 1e-73
Identities = 151/343 (44%), Positives = 212/343 (61%), Gaps = 5/343 (1%)
Query: 79 KDSGAEKIEDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALP 138
K+ G+ I GKV K++ +G + A AG I++IAG + +IG T+T + ALP
Sbjct: 241 KEDGS--IAKGKVSKLLGFEGLNRIELPEASAGYIVAIAGFADANIGETLTCPDEPQALP 298
Query: 139 TVELDPPTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMS-ETFEV 197
+++D PT+ MTF VNDSP AG++G +T +I DRL E ETN+A+ V G S E F V
Sbjct: 299 LIKVDEPTLQMTFSVNDSPFAGQEGKFVTSRQIRDRLNRELETNVALRVEDGESAEQFLV 358
Query: 198 QGRGELQLGILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVMEA 257
GRGEL LGILIE MRREG+E V+ P+V+Y+ GQ EP+E + ++V + VG +E
Sbjct: 359 SGRGELHLGILIETMRREGYEFQVAQPQVIYREVNGQPCEPVEYLVLDVPEAAVGACIER 418
Query: 258 LSHRRAEITDMGPVAGTVGRTRLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKFR 317
L RR E+ DM GRT+L P+RGL+G+R F TRG G M+ +FL+Y
Sbjct: 419 LGQRRGEMQDMQTSVN--GRTQLEFVIPARGLLGFRGDFIRITRGEGIMNHSFLEYRPMS 476
Query: 318 GPLGNVRKGVLVSVGFGSITLHALMSLEARGTLFVSPGMEAYDGMIVGEHSRDTDLDVNP 377
G L GV+V+ G T +A+ + E RG F++PG + Y GMI+GEH+R D+++N
Sbjct: 477 GDLETRYNGVMVAFEEGVATFYAMKNAEDRGVFFITPGTKVYKGMIIGEHNRPQDIELNV 536
Query: 378 VRAKQLTNVRSVNKDDTVKLIPPRLMTLEEAIGYVASDELIEV 420
+ KQLTN RS D+ V+L P M LE A+ Y+ DEL+E+
Sbjct: 537 CKTKQLTNHRSATGDELVQLQAPEDMNLERALEYIGPDELVEI 579
>TYPA_HAEIN (P44910) GTP-binding protein typA/bipA homolog
Length = 616
Score = 270 bits (689), Expect = 7e-72
Identities = 148/339 (43%), Positives = 212/339 (61%), Gaps = 6/339 (1%)
Query: 85 KIEDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDP 144
K G++ +V+ G D A AGDI++I GL +I T+ + + ALP++ +D
Sbjct: 251 KTRQGRIGQVLGHLGLQRYEEDVAYAGDIVAITGLGELNISDTICDINTVEALPSLTVDE 310
Query: 145 PTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINV--LPGMSETFEVQGRGE 202
PT++M F VN SP AG++G ++T +I +RL E N+A+ V P E F V GRGE
Sbjct: 311 PTVTMFFCVNTSPFAGQEGKYVTSRQILERLNKELVHNVALRVEETPNPDE-FRVSGRGE 369
Query: 203 LQLGILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVMEALSHRR 262
L L +LIENMRREG+EL+VS PKV+Y+ G+K EP E+VTI+V ++H G VMEAL R+
Sbjct: 370 LHLSVLIENMRREGYELAVSRPKVIYRDIDGKKQEPYEQVTIDVEEQHQGSVMEALGIRK 429
Query: 263 AEITDMGPVAGTVGRTRLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKFR-GPLG 321
E+ DM P GR RL PSRGL+G+R F + T GTG ++ +F Y++ + G +G
Sbjct: 430 GEVRDMLP--DGKGRVRLEYIIPSRGLIGFRGDFMTMTSGTGLLYSSFSHYDEIKGGEIG 487
Query: 322 NVRKGVLVSVGFGSITLHALMSLEARGTLFVSPGMEAYDGMIVGEHSRDTDLDVNPVRAK 381
+ GVL+S G +AL L+ RG L + +E Y+G I+G HSR DL VN ++ K
Sbjct: 488 QRKNGVLISNATGKALGYALFGLQERGKLMIDANIEVYEGQIIGIHSRSNDLTVNCLQGK 547
Query: 382 QLTNVRSVNKDDTVKLIPPRLMTLEEAIGYVASDELIEV 420
+LTN+R+ KDD + L P +LE+AI ++ DEL+EV
Sbjct: 548 KLTNMRASGKDDAIVLTTPVKFSLEQAIEFIDDDELVEV 586
>TYPA_BUCAI (P57508) GTP-binding protein TypA/BipA homolog
Length = 607
Score = 243 bits (619), Expect = 9e-64
Identities = 135/338 (39%), Positives = 201/338 (58%), Gaps = 4/338 (1%)
Query: 85 KIEDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDP 144
K GKV KV+ G + + AGDII+I GL+ I T+ + + LP + +D
Sbjct: 245 KHRSGKVNKVLNYFGLKRMEINQGYAGDIIAITGLNKLKISDTICHPDNLQPLPALSIDE 304
Query: 145 PTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMSET-FEVQGRGEL 203
PT++M F VN SP +G++G ++T +I +RL E+ N+A+ + F V GRGEL
Sbjct: 305 PTVNMFFSVNTSPFSGKEGKYITSRQILERLKKETMHNVALQIKETKDPNIFSVSGRGEL 364
Query: 204 QLGILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVMEALSHRRA 263
L ILIENMRREGFEL VS PK++++ G + EP E VT+++ +++ G VM+ + R+
Sbjct: 365 HLSILIENMRREGFELEVSRPKIIFREINGIQKEPFENVTLDIEEKNQGSVMQFIGIRKG 424
Query: 264 EITDMGPVAGTVGRTRLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKFRGP-LGN 322
E+ +M P + GR RL SR L+G+RS F S T GTG + +F Y+ + +G
Sbjct: 425 ELKNMIP--DSKGRVRLEYILSSRALIGFRSEFMSITSGTGLCYSSFSHYDNLQNSNIGQ 482
Query: 323 VRKGVLVSVGFGSITLHALMSLEARGTLFVSPGMEAYDGMIVGEHSRDTDLDVNPVRAKQ 382
+ GVL+S G +L +L+ RG LF+ G + Y+G I+G H+R DL VN + K+
Sbjct: 483 RKNGVLISNSTGMAVGFSLFNLQERGKLFIGHGAQVYEGQIIGLHNRSNDLTVNCLTGKK 542
Query: 383 LTNVRSVNKDDTVKLIPPRLMTLEEAIGYVASDELIEV 420
LTN+R+ D+ + L TLEEAIG++ DEL+EV
Sbjct: 543 LTNMRASGTDEAIVLTTAIHFTLEEAIGFINDDELVEV 580
>TYPA_BUCBP (Q89AC9) GTP-binding protein TypA/BipA homolog
Length = 611
Score = 237 bits (605), Expect = 4e-62
Identities = 130/334 (38%), Positives = 201/334 (59%), Gaps = 3/334 (0%)
Query: 89 GKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTIS 148
GK+ K+++ G + + A +GDII+I G+ +I T+ + +SALP +++D PT+
Sbjct: 253 GKIGKILQYLGLNKIEINEAQSGDIIAITGIDKLNISDTICDPQYISALPMLKIDEPTVE 312
Query: 149 MTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLP-GMSETFEVQGRGELQLGI 207
M F VN SP +G +G ++T +I +RL E N+A+ + + TF V GRGEL L I
Sbjct: 313 MLFSVNKSPFSGTEGKYITSRQIFNRLKKEENYNVALKIKETNDTNTFSVSGRGELHLSI 372
Query: 208 LIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVMEALSHRRAEITD 267
LIENMRREGFEL VS P+V+ KT EP+E V +++ +++ G +M+ + R+ I++
Sbjct: 373 LIENMRREGFELEVSRPQVILKTINELIQEPMETVVLDIENKYQGTIMKTIGQRKGTISN 432
Query: 268 MGPVAGTVGRTRLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKFRG-PLGNVRKG 326
+ P RTRL SR L+G+R+ FS+ T G+G + F Y+K + R G
Sbjct: 433 ITPDQNNE-RTRLDCIISSRSLIGFRTEFSTLTSGSGLFYSTFSHYQKIESNKIKRHRNG 491
Query: 327 VLVSVGFGSITLHALMSLEARGTLFVSPGMEAYDGMIVGEHSRDTDLDVNPVRAKQLTNV 386
VL++ G +L +L+ RG LF++ G + Y+G IVG H+R DL VN + K+LTN+
Sbjct: 492 VLIANKTGQAIGFSLFNLQNRGKLFITHGTKVYEGQIVGIHNRVNDLTVNCLSGKKLTNM 551
Query: 387 RSVNKDDTVKLIPPRLMTLEEAIGYVASDELIEV 420
R+ D+ + L P MTLE AI ++ DEL+E+
Sbjct: 552 RASGSDEAITLTTPIKMTLEYAISFINDDELVEI 585
>TYPA_BUCAP (Q8K9C8) GTP-binding protein TypA/BipA homolog
Length = 609
Score = 233 bits (594), Expect = 7e-61
Identities = 127/335 (37%), Positives = 198/335 (58%), Gaps = 4/335 (1%)
Query: 88 DGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTI 147
+GK+ KV+ G + AGDII+I GL+ +I T+ + + LP + +D PT+
Sbjct: 248 NGKINKVLNYFGLKRLEIQKGDAGDIIAITGLNKLNISDTICHPDNLCPLPPLIIDEPTV 307
Query: 148 SMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGM-SETFEVQGRGELQLG 206
+M F VN SP +G++G ++T +I +RL E+ N+A+ V + F V GRGEL L
Sbjct: 308 NMFFSVNTSPFSGKEGKYITSRQILERLKKETIHNVALQVKETKDANIFSVSGRGELHLS 367
Query: 207 ILIENMRREGFELSVSPPKVMYKTDKGQKLEPIEEVTIEVNDEHVGFVMEALSHRRAEIT 266
ILIENMRREGFEL VS PK++++ K EP E + +++ ++H G +M+ + R+ E+
Sbjct: 368 ILIENMRREGFELEVSRPKIIFREINDIKQEPFENIILDIEEKHQGNIMKFIGERKGELK 427
Query: 267 DMGPVAGTVGRTRLCLTCPSRGLVGYRSVFSSETRGTGFMHRAFLKYEKF-RGPLGNVRK 325
++ R RL SR L+G+R+ F S T GTG + +F Y+K+ + +G +
Sbjct: 428 NI--TVDPNKRIRLEYLLSSRALIGFRTEFMSITSGTGLFYSSFSHYQKYQKNDIGQRKN 485
Query: 326 GVLVSVGFGSITLHALMSLEARGTLFVSPGMEAYDGMIVGEHSRDTDLDVNPVRAKQLTN 385
GVL+S G AL +L+ RG LF+ G + Y+G I+G H+R DL VN + K+LTN
Sbjct: 486 GVLISNSTGMAVAFALFNLQERGKLFLGHGTQVYEGQIIGLHNRSNDLTVNCLTGKKLTN 545
Query: 386 VRSVNKDDTVKLIPPRLMTLEEAIGYVASDELIEV 420
+R+ D+ + L TLEEA+ ++ DEL+E+
Sbjct: 546 MRASGTDEAIILTTFTKFTLEEALSFINDDELVEI 580
>EFG_RICTY (Q8KTB2) Elongation factor G (EF-G)
Length = 699
Score = 51.2 bits (121), Expect = 5e-06
Identities = 37/123 (30%), Positives = 62/123 (50%), Gaps = 9/123 (7%)
Query: 108 AGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISMTFGVNDSPLAGRDGTHLT 167
A AGDI+++AGL S G T++ ++ L +E P I + + + G L+
Sbjct: 369 ASAGDIVALAGLKDTSTGDTLSDIDTQVVLERMEFPEPVIELAVEPKSTADQEKMGLALS 428
Query: 168 GGKIGDRLMAESETNLAINVLPGMSETFEVQGRGELQLGILIENMRRE-GFELSVSPPKV 226
RL AE + + ++ +T ++G GEL L I+I+ MRRE E ++ P+V
Sbjct: 429 ------RLAAE-DPSFKVSTDHETGQTV-IKGMGELHLEIIIDRMRREFKVEANIGAPQV 480
Query: 227 MYK 229
Y+
Sbjct: 481 AYR 483
Score = 33.5 bits (75), Expect = 1.1
Identities = 23/90 (25%), Positives = 40/90 (43%), Gaps = 3/90 (3%)
Query: 236 LEPIEEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAGTVGRTRLCLTCPSRGLVGYRSV 295
LEPI +V + DE++G ++ L+ RR +I M P T P + GY +
Sbjct: 610 LEPIMKVEVITPDEYMGDIIGDLNSRRGQIQSMDPRGNAQVVTAY---VPLAEMFGYVNT 666
Query: 296 FSSETRGTGFMHRAFLKYEKFRGPLGNVRK 325
S ++G F Y++ + ++ K
Sbjct: 667 LRSLSQGRAQFSMIFSHYDQVPSQVADIIK 696
>EFG_RICHE (Q8KTB4) Elongation factor G (EF-G)
Length = 699
Score = 51.2 bits (121), Expect = 5e-06
Identities = 36/126 (28%), Positives = 57/126 (44%), Gaps = 15/126 (11%)
Query: 108 AGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISMTF---GVNDSPLAGRDGT 164
A AGDI+++AGL S G T++ ++ L +E P I + D G +
Sbjct: 369 ASAGDIVALAGLKDTSTGDTLSDIDTQVVLERMEFPEPVIELAVEPKSTADQEKMGLALS 428
Query: 165 HLTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGELQLGILIENMRRE-GFELSVSP 223
L R+ + ET + ++G GEL L I+I+ MRRE E ++
Sbjct: 429 RLAAEDPSFRVSTDHETGQTV-----------IKGMGELHLEIIIDRMRREFKVEANIGV 477
Query: 224 PKVMYK 229
P+V Y+
Sbjct: 478 PQVAYR 483
Score = 34.3 bits (77), Expect = 0.64
Identities = 23/90 (25%), Positives = 41/90 (45%), Gaps = 3/90 (3%)
Query: 236 LEPIEEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAGTVGRTRLCLTCPSRGLVGYRSV 295
LEPI +V + DE++G ++ L+ RR +I M P T P + GY ++
Sbjct: 610 LEPIMKVEVITPDEYMGDIIGDLNSRRGQIQSMDPRGNAQVVT---ANVPLAEMFGYVNM 666
Query: 296 FSSETRGTGFMHRAFLKYEKFRGPLGNVRK 325
S ++G F Y++ + ++ K
Sbjct: 667 LRSLSQGRAQFSMIFSHYDQVPSQVADIIK 696
>EFG_RICPR (P41084) Elongation factor G (EF-G)
Length = 699
Score = 50.8 bits (120), Expect = 7e-06
Identities = 36/126 (28%), Positives = 57/126 (44%), Gaps = 15/126 (11%)
Query: 108 AGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISMTF---GVNDSPLAGRDGT 164
A AGDI+++AGL S G T++ ++ L +E P I + D G +
Sbjct: 369 ASAGDIVALAGLKDTSTGDTLSDIDKQVVLERMEFPEPVIELAVEPKSTADQEKMGLALS 428
Query: 165 HLTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGELQLGILIENMRRE-GFELSVSP 223
L R+ + ET + ++G GEL L I+I+ MRRE E ++
Sbjct: 429 RLAAEDPSFRVSTDHETGQTV-----------IKGMGELHLEIIIDRMRREFKVEANIGA 477
Query: 224 PKVMYK 229
P+V Y+
Sbjct: 478 PQVAYR 483
Score = 32.7 bits (73), Expect = 1.9
Identities = 23/90 (25%), Positives = 41/90 (45%), Gaps = 3/90 (3%)
Query: 236 LEPIEEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAGTVGRTRLCLTCPSRGLVGYRSV 295
LEPI +V + DE++G ++ L+ RR +I +M P T P + GY +
Sbjct: 610 LEPIMKVEVITPDEYMGDIIGDLNSRRGQIQNMDPRGNAQVVT---AHVPLAEMFGYVNT 666
Query: 296 FSSETRGTGFMHRAFLKYEKFRGPLGNVRK 325
S ++G F Y++ + ++ K
Sbjct: 667 LRSLSQGRAQFSMIFSHYDQVPSQVADMIK 696
>EFG_STRRA (P29541) Elongation factor G (EF-G) (Fragment)
Length = 341
Score = 50.1 bits (118), Expect = 1e-05
Identities = 36/136 (26%), Positives = 66/136 (48%), Gaps = 9/136 (6%)
Query: 106 DCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISMTFGVNDSPLAGRDGTH 165
+ GAGDI+++ GL + G T++ + L +++ P I + P + D
Sbjct: 13 ESVGAGDIVAVMGLKQTTTGETLSDEKSPVILESMDFPAPVIQVAI----EPKSKGDQEK 68
Query: 166 LTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGELQLGILIENMRRE-GFELSVSPP 224
L + + +AE + + ++ +T + G GEL L +L++ MRRE E +V P
Sbjct: 69 L---GVAIQRLAEEDPSFQVHSDEETGQTI-IGGMGELHLEVLVDRMRREFKVEANVGKP 124
Query: 225 KVMYKTDKGQKLEPIE 240
+V Y+ Q +E +E
Sbjct: 125 QVAYRETIRQAVEKVE 140
>EFG_LACLA (Q9CDG1) Elongation factor G (EF-G)
Length = 709
Score = 50.1 bits (118), Expect = 1e-05
Identities = 38/143 (26%), Positives = 67/143 (46%), Gaps = 13/143 (9%)
Query: 90 KVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISM 149
++ ++++ T AGDI + GL + + G ++T + L ++E+ P I +
Sbjct: 365 RIGRILQMHANTRKEIQTVYAGDIAAAVGLKNTTTGDSLTDEKAKIILESIEVPEPVIQL 424
Query: 150 TFGVNDSPLAGRDGTHLTGGKIGDRL--MAESETNLAINVLPGMSETFEVQGRGELQLGI 207
V A +D K+G L +AE + + P ET + G GEL L +
Sbjct: 425 M--VEPKTKADQD-------KMGVALQKLAEEDPTFRVETNPETGETV-ISGMGELHLDV 474
Query: 208 LIENMRRE-GFELSVSPPKVMYK 229
L++ M+RE E +V P+V Y+
Sbjct: 475 LVDRMKREFKVEANVGAPQVAYR 497
Score = 38.1 bits (87), Expect = 0.044
Identities = 26/86 (30%), Positives = 43/86 (49%), Gaps = 5/86 (5%)
Query: 229 KTDKGQKLEPIEEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAGTVGRTRLC-LTCPSR 287
KT K LEP+ +VTI V +E++G +M ++ RR ++ M G++++ P
Sbjct: 608 KTAKPAILEPMMKVTITVPEENLGDIMGHVTARRGQVNSM----EAHGKSQIVNAFVPLA 663
Query: 288 GLVGYRSVFSSETRGTGFMHRAFLKY 313
+ GY + S T+G G F Y
Sbjct: 664 EMFGYATTLRSSTQGRGTFMMVFDHY 689
>EFG_PROMM (Q7V501) Elongation factor G (EF-G)
Length = 691
Score = 49.7 bits (117), Expect = 1e-05
Identities = 39/146 (26%), Positives = 68/146 (45%), Gaps = 9/146 (6%)
Query: 85 KIEDGKVVKVMKKKGTTMVVTDCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDP 144
K E ++ +++ K D AGD+ ++ GL + + G T+ T + L T+ +
Sbjct: 343 KNEKERISRLIILKADDREEVDALRAGDLGAVLGLKNTTTGDTLCTTDDPIVLETLYIPE 402
Query: 145 PTISMTFGVNDSPLAGRDGTHLTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGELQ 204
P IS+ P D L+ + +AE + ++ P S+T + G GEL
Sbjct: 403 PVISVAV----EPKTKGDMEKLSKALLS---LAEEDPTFRVSTDPETSQTV-IAGMGELH 454
Query: 205 LGILIENMRRE-GFELSVSPPKVMYK 229
L IL++ M RE E ++ P+V Y+
Sbjct: 455 LEILVDRMLREFKVEANIGAPQVSYR 480
Score = 35.8 bits (81), Expect = 0.22
Identities = 21/80 (26%), Positives = 40/80 (49%), Gaps = 3/80 (3%)
Query: 236 LEPIEEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAGTVGRTRLCLTCPSRGLVGYRSV 295
LEP+ +V +E+ ++ +G ++ LS RR ++ + G +++ P + GY +
Sbjct: 598 LEPMMKVEVEIPEDFLGAIIGDLSSRRGQVEGQ---SIDDGLSKVQSKVPLAEMFGYATQ 654
Query: 296 FSSETRGTGFMHRAFLKYEK 315
S T+G G F YE+
Sbjct: 655 LRSMTQGRGIFSMEFSHYEE 674
>EFG2_VIBPA (Q87M30) Elongation factor G 2 (EF-G 2)
Length = 696
Score = 49.7 bits (117), Expect = 1e-05
Identities = 39/133 (29%), Positives = 61/133 (45%), Gaps = 9/133 (6%)
Query: 106 DCAGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISMTFGVNDSPLAGRDGTH 165
D A AGDII++ G+ + GHT+ + L + P IS+ D + G+
Sbjct: 366 DSAQAGDIIAVVGMKNVQTGHTLCDPKHECTLEPMIFPEPVISIAVKPKD-----KGGSE 420
Query: 166 LTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGELQLGILIENMRRE-GFELSVSPP 224
G IG M + + + ET ++G GEL L I ++ ++R G EL V P
Sbjct: 421 KMGIAIGK--MVAEDPSFQVETDEESGETI-LKGMGELHLDIKVDILKRTYGVELEVGAP 477
Query: 225 KVMYKTDKGQKLE 237
+V Y+ Q +E
Sbjct: 478 QVAYRETITQAIE 490
Score = 38.1 bits (87), Expect = 0.044
Identities = 28/80 (35%), Positives = 37/80 (46%), Gaps = 3/80 (3%)
Query: 234 QKLEPIEEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAGTVGRTRLCLTCPSRGLVGYR 293
Q LEPI +V + ++HVG V+ L+ RR I D AGT G R+ P + GY
Sbjct: 598 QLLEPIMKVDVFTPEDHVGDVIGDLNRRRGMIKDQ--QAGTTG-VRIKGDVPLSEMFGYI 654
Query: 294 SVFSSETRGTGFMHRAFLKY 313
+ T G G F Y
Sbjct: 655 GTLRTMTSGRGQFSMEFSHY 674
>EFG_RICRH (Q8KTB7) Elongation factor G (EF-G)
Length = 697
Score = 49.3 bits (116), Expect = 2e-05
Identities = 35/126 (27%), Positives = 57/126 (44%), Gaps = 15/126 (11%)
Query: 108 AGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISMTF---GVNDSPLAGRDGT 164
A AGDI+++AGL + G T++ ++ L +E P I + D G +
Sbjct: 369 ASAGDIVALAGLKDTTTGDTLSDIDQQVILEKMEFPEPVIELAVEPKSTADQEKMGLALS 428
Query: 165 HLTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGELQLGILIENMRRE-GFELSVSP 223
L R+ + ET + ++G GEL L I+I+ MRRE E ++
Sbjct: 429 RLAAEDPSFRVSTDHETGQTV-----------IKGMGELHLEIIIDRMRREFKVEANIGA 477
Query: 224 PKVMYK 229
P+V Y+
Sbjct: 478 PQVAYR 483
Score = 34.3 bits (77), Expect = 0.64
Identities = 23/90 (25%), Positives = 40/90 (43%), Gaps = 3/90 (3%)
Query: 236 LEPIEEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAGTVGRTRLCLTCPSRGLVGYRSV 295
LEPI +V + DE++G ++ L+ RR +I M P T P + GY +
Sbjct: 608 LEPIMQVEVITPDEYMGDIIGDLNSRRGQIQSMDPRGNAQVVT---ANVPLAEMFGYVNT 664
Query: 296 FSSETRGTGFMHRAFLKYEKFRGPLGNVRK 325
S ++G F Y++ + ++ K
Sbjct: 665 LRSLSQGRAQFSMIFSHYDQVPSQVADIIK 694
>EFG_RICMO (Q8KTB6) Elongation factor G (EF-G)
Length = 699
Score = 49.3 bits (116), Expect = 2e-05
Identities = 35/126 (27%), Positives = 57/126 (44%), Gaps = 15/126 (11%)
Query: 108 AGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISMTF---GVNDSPLAGRDGT 164
A AGDI+++AGL + G T++ ++ L +E P I + D G +
Sbjct: 369 ASAGDIVALAGLKDTTTGDTLSDIDQQVILERMEFPEPVIELAVEPKSTADQEKMGLALS 428
Query: 165 HLTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGELQLGILIENMRRE-GFELSVSP 223
L R+ + ET + ++G GEL L I+I+ MRRE E ++
Sbjct: 429 RLAAEDPSFRVSTDHETGQTV-----------IKGMGELHLEIIIDRMRREFKVEANIGA 477
Query: 224 PKVMYK 229
P+V Y+
Sbjct: 478 PQVAYR 483
Score = 34.3 bits (77), Expect = 0.64
Identities = 23/91 (25%), Positives = 43/91 (46%), Gaps = 5/91 (5%)
Query: 236 LEPIEEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAGTVGRTRLCL-TCPSRGLVGYRS 294
LEPI +V + DE++G ++ L+ RR +I M P G ++ + P + GY +
Sbjct: 610 LEPIMQVEVITPDEYMGDIIGDLNSRRGQIQSMDP----RGNAQVVIANVPLAEMFGYVN 665
Query: 295 VFSSETRGTGFMHRAFLKYEKFRGPLGNVRK 325
S ++G F Y++ + ++ K
Sbjct: 666 TLRSLSQGRAQFSMIFSHYDQVPSQVADIIK 696
>EFG_RICFE (Q8KTA8) Elongation factor G (EF-G)
Length = 699
Score = 49.3 bits (116), Expect = 2e-05
Identities = 35/126 (27%), Positives = 57/126 (44%), Gaps = 15/126 (11%)
Query: 108 AGAGDIISIAGLSSPSIGHTVTTVEIMSALPTVELDPPTISMTF---GVNDSPLAGRDGT 164
A AGDI+++AGL + G T++ ++ L +E P I + D G +
Sbjct: 369 ASAGDIVALAGLKDTTTGDTLSDIDKQVILERMEFPEPVIELAVEPKSTADQEKMGLALS 428
Query: 165 HLTGGKIGDRLMAESETNLAINVLPGMSETFEVQGRGELQLGILIENMRRE-GFELSVSP 223
L R+ + ET + ++G GEL L I+I+ MRRE E ++
Sbjct: 429 RLAAEDPSFRVSTDHETGQTV-----------IKGMGELHLEIIIDRMRREFKVEANIGA 477
Query: 224 PKVMYK 229
P+V Y+
Sbjct: 478 PQVAYR 483
Score = 33.9 bits (76), Expect = 0.84
Identities = 23/90 (25%), Positives = 40/90 (43%), Gaps = 3/90 (3%)
Query: 236 LEPIEEVTIEVNDEHVGFVMEALSHRRAEITDMGPVAGTVGRTRLCLTCPSRGLVGYRSV 295
LEPI +V + DE++G ++ L+ RR +I M P T P + GY +
Sbjct: 610 LEPIMKVEVITPDEYMGDIIGDLNSRRGQIQSMDPRGNAQVVT---ANVPLAEMFGYVNT 666
Query: 296 FSSETRGTGFMHRAFLKYEKFRGPLGNVRK 325
S ++G F Y++ + ++ K
Sbjct: 667 LRSLSQGRAQFSMIFSHYDQVPSQVADIIK 696
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.321 0.138 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 50,418,594
Number of Sequences: 164201
Number of extensions: 2153080
Number of successful extensions: 5506
Number of sequences better than 10.0: 211
Number of HSP's better than 10.0 without gapping: 45
Number of HSP's successfully gapped in prelim test: 166
Number of HSP's that attempted gapping in prelim test: 5198
Number of HSP's gapped (non-prelim): 320
length of query: 433
length of database: 59,974,054
effective HSP length: 113
effective length of query: 320
effective length of database: 41,419,341
effective search space: 13254189120
effective search space used: 13254189120
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC140034.13


