

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
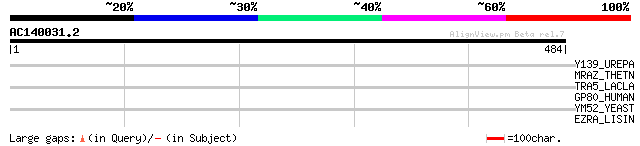
Query= AC140031.2 + phase: 0 /pseudo
(484 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
Y139_UREPA (Q9PR05) Hypothetical protein UU139 38 0.051
MRAZ_THETN (Q8R9F8) Protein mraZ 34 0.96
TRA5_LACLA (P35881) Transposase for insertion sequence element I... 32 3.7
GP80_HUMAN (Q96P68) Probable G protein-coupled receptor GPR80 (P... 32 3.7
YM52_YEAST (Q04322) Hypothetical 82.1 kDa protein in SGS1-MRPL24... 32 4.8
EZRA_LISIN (Q92BB4) Septation ring formation regulator ezrA 31 8.2
>Y139_UREPA (Q9PR05) Hypothetical protein UU139
Length = 236
Score = 38.1 bits (87), Expect = 0.051
Identities = 20/67 (29%), Positives = 39/67 (57%), Gaps = 1/67 (1%)
Query: 30 NFPTVLVMDSTYQTNMYKMPMFEIVGVTSTDLTYSVGF-EFVTHEKEENFVWVLQMMCKL 88
NF ++D ++ ++K+ E GV ST+L ++ GF +FV K +NF+ ++ ++
Sbjct: 136 NFNNSYLIDPNWKKYVFKIKKDESKGVISTNLLHNYGFVDFVVQSKSQNFLKAYRVQDEI 195
Query: 89 LTSRMNV 95
L S +N+
Sbjct: 196 LKSVLNL 202
>MRAZ_THETN (Q8R9F8) Protein mraZ
Length = 143
Score = 33.9 bits (76), Expect = 0.96
Identities = 20/63 (31%), Positives = 34/63 (53%), Gaps = 4/63 (6%)
Query: 370 EEWNGIQERLKKIPNRMNLEMKETKSTSQGSQEKSGYCEVYRKGHQLVESHLFGSVLIPK 429
+EW I+E+LK +P L K+ ++ ++ + CEV ++G L+ SHL I K
Sbjct: 46 DEWKNIEEKLKTLP----LTKKDARAFTRFFLAGAVECEVDKQGRILIPSHLREHAKIEK 101
Query: 430 ILI 432
+I
Sbjct: 102 DVI 104
>TRA5_LACLA (P35881) Transposase for insertion sequence element
IS905
Length = 391
Score = 32.0 bits (71), Expect = 3.7
Identities = 27/113 (23%), Positives = 48/113 (41%), Gaps = 8/113 (7%)
Query: 23 HPLSFFNNFPTVLVMDSTY----QTNMYKMPMFEIVGVTSTDLTYSVGFEFVTHEKEENF 78
H S N+ +VL +D TY + + K + +G+T +G+E +E ++
Sbjct: 149 HERSLEANY-SVLFLDGTYLPLRRGTVSKECIHIALGITPEGQKAVLGYEIAPNENNASW 207
Query: 79 VWVLQMMCKLLTSRMNVPKVVVTDKDTTLMNAVSNVLPESFAMNCYFHVQKNV 131
+L KL + +VVTD L +S P + C H+ +N+
Sbjct: 208 STLLD---KLQNQGIQQVSLVVTDGFKGLEQIISQAYPLAKQQRCLIHISRNL 257
>GP80_HUMAN (Q96P68) Probable G protein-coupled receptor GPR80
(P2Y-like nucleotide receptor) (P2Y-like GPCR)
Length = 337
Score = 32.0 bits (71), Expect = 3.7
Identities = 21/79 (26%), Positives = 34/79 (42%), Gaps = 3/79 (3%)
Query: 14 LKTYFALIQHPLSFFNNFPTVLVMDSTYQTNM--YKMPMFEIVGVTSTDLTYSVGFEFVT 71
LK ++ + + + F FP V+ STY M +K ++ + TDL Y F+
Sbjct: 31 LKMHYLPVIYGIIFLVGFPGNAVVISTYIFKMRPWKSSTIIMLNLACTDLLYLTSLPFLI 90
Query: 72 HEKEENFVWVL-QMMCKLL 89
H W+ MCK +
Sbjct: 91 HYYASGENWIFGDFMCKFI 109
>YM52_YEAST (Q04322) Hypothetical 82.1 kDa protein in SGS1-MRPL24
intergenic region
Length = 720
Score = 31.6 bits (70), Expect = 4.8
Identities = 14/66 (21%), Positives = 33/66 (49%)
Query: 103 KDTTLMNAVSNVLPESFAMNCYFHVQKNVKVRCILDCRYPMAKKDGKEVKHGDVVKKIMR 162
+D + A + L +F N F+ ++ ++C LD K+D K++ + ++ +++
Sbjct: 211 EDPLDLAAQKHFLASTFKRNMLFYKSEDNSIKCDLDKNILNLKEDSKKINNNEIPEEVSS 270
Query: 163 VW*VII 168
W +I
Sbjct: 271 FWLKVI 276
>EZRA_LISIN (Q92BB4) Septation ring formation regulator ezrA
Length = 571
Score = 30.8 bits (68), Expect = 8.2
Identities = 18/62 (29%), Positives = 36/62 (58%), Gaps = 2/62 (3%)
Query: 360 ESNEDCFSVEEEWNGIQERLKKIPNRMNLEMKETKSTSQGSQEKSGYCEVYRKGHQLVES 419
E+ E SV++E I+ ++++IP+ L +T + ++ ++GY E+ RKG+ L +
Sbjct: 194 EAREIVISVQKEMAVIEAQMERIPSL--LHETDTILPEEMNKLRAGYEEMVRKGYYLAQM 251
Query: 420 HL 421
L
Sbjct: 252 EL 253
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.342 0.150 0.517
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 53,637,294
Number of Sequences: 164201
Number of extensions: 2139265
Number of successful extensions: 7294
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 1
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 7292
Number of HSP's gapped (non-prelim): 6
length of query: 484
length of database: 59,974,054
effective HSP length: 114
effective length of query: 370
effective length of database: 41,255,140
effective search space: 15264401800
effective search space used: 15264401800
T: 11
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 68 (30.8 bits)
Medicago: description of AC140031.2


