

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139743.2 - phase: 0 /pseudo
(461 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
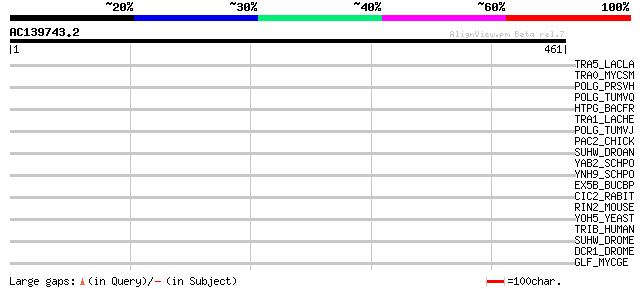
Score E
Sequences producing significant alignments: (bits) Value
TRA5_LACLA (P35881) Transposase for insertion sequence element I... 40 0.016
TRA0_MYCSM (P35883) Transposase for insertion sequence element I... 37 0.11
POLG_PRSVH (Q01901) Genome polyprotein [Contains: P1 proteinase ... 34 0.69
POLG_TUMVQ (Q02597) Genome polyprotein [Contains: P1 proteinase ... 34 0.90
HTPG_BACFR (P58476) Chaperone protein htpG (Heat shock protein h... 34 0.90
TRA1_LACHE (P35880) Transposase for insertion sequence element I... 33 1.5
POLG_TUMVJ (P89509) Genome polyprotein [Contains: P1 proteinase ... 33 2.0
PAC2_CHICK (O13154) Protein kinase C and casein kinase substrate... 33 2.0
SUHW_DROAN (Q08875) Suppressor of hairy wing protein 32 2.6
YAB2_SCHPO (Q09804) Hypothetical protein C2G11.02 in chromosome I 32 3.4
YNH9_SCHPO (Q96WW0) Hypothetical WD-repeat protein C32H8.09 in c... 32 4.5
EX5B_BUCBP (Q89AB3) Exodeoxyribonuclease V beta chain (EC 3.1.11.5) 32 4.5
CIC2_RABIT (P13806) Dihydropyridine-sensitive L-type, calcium ch... 32 4.5
RIN2_MOUSE (Q9D684) Ras and Rab interactor 2 (Ras interaction/in... 31 5.9
YOH5_YEAST (Q08234) Probable ATP-dependent transporter YOL074C/Y... 31 7.6
TRIB_HUMAN (Q14669) Thyroid receptor interacting protein 12 (TRI... 31 7.6
SUHW_DROME (P08970) Suppressor of hairy wing protein 31 7.6
DCR1_DROME (Q9VCU9) Endoribonuclease Dcr-1 (EC 3.1.26.-) (Dicer-... 31 7.6
GLF_MYCGE (Q49398) UDP-galactopyranose mutase (EC 5.4.99.9) 30 10.0
>TRA5_LACLA (P35881) Transposase for insertion sequence element
IS905
Length = 391
Score = 39.7 bits (91), Expect = 0.016
Identities = 31/152 (20%), Positives = 70/152 (45%), Gaps = 10/152 (6%)
Query: 237 KSLQFLISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTY----K 292
+ + +I ++ H+Y+ + + + T E++ H S++ N+ +VL +D TY +
Sbjct: 115 REISDIIERMYGHHYSPATISNISKATQENVATFHERSLEA--NY-SVLFLDGTYLPLRR 171
Query: 293 TNMYRMPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTD 352
+ + + +G+T +G+ +E N W T+L KL + + ++VTD
Sbjct: 172 GTVSKECIHIALGITPEGQKAVLGYEIAPNEN--NASWS-TLLDKLQNQGIQQVSLVVTD 228
Query: 353 RDMSLMKAVAHVFPESYALNCFFHVQANVKQR 384
L + ++ +P + C H+ N+ +
Sbjct: 229 GFKGLEQIISQAYPLAKQQRCLIHISRNLASK 260
>TRA0_MYCSM (P35883) Transposase for insertion sequence element
IS6120
Length = 323
Score = 37.0 bits (84), Expect = 0.11
Identities = 32/114 (28%), Positives = 46/114 (40%), Gaps = 3/114 (2%)
Query: 334 MLRKLLSSKMNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQRCVLNCKYPL 393
+LR M P + + D + KAV VFP + C+FH QANV + +P
Sbjct: 113 LLRDCKRRGMTAPVLAIGDGALGFWKAVREVFPATKEQRCWFHKQANV-LAALPKSAHPS 171
Query: 394 GFKKDGKEVSNRDVVKKIMNAWKAMVESPNQQLYANALVEFKDSCSDFSIFVDY 447
KE+ N + + K A KA E+ Y A+ + D F Y
Sbjct: 172 ALAAI-KEIYNAEDIDKAQIAVKAF-EADFGAKYPKAVAKITDDLDVLLEFYKY 223
>POLG_PRSVH (Q01901) Genome polyprotein [Contains: P1 proteinase
(N-terminal protein); Helper component proteinase (EC
3.4.22.45) (HC-pro); Protein P3; 6 kDa protein 1 (6K1);
Cytoplasmic inclusion protein (CI); 6 kDa protein 2
(6K2); Viral genome-linked p
Length = 3344
Score = 34.3 bits (77), Expect = 0.69
Identities = 14/55 (25%), Positives = 31/55 (55%)
Query: 401 EVSNRDVVKKIMNAWKAMVESPNQQLYANALVEFKDSCSDFSIFVDYAMTTLDEV 455
+V D V +I+N +K ++ S N +Y +L + +D D + +D+ +T +++
Sbjct: 1373 DVDKSDCVYRILNKFKGVINSSNTNVYHQSLDDIRDFYEDKQLTIDFDITGENQI 1427
>POLG_TUMVQ (Q02597) Genome polyprotein [Contains: P1 proteinase
(N-terminal protein); Helper component proteinase (EC
3.4.22.45) (HC-pro); Protein P3; 6 kDa protein 1 (6K1);
Cytoplasmic inclusion protein (CI); 6 kDa protein 2
(6K2); Viral genome-linked p
Length = 3163
Score = 33.9 bits (76), Expect = 0.90
Identities = 15/59 (25%), Positives = 32/59 (53%)
Query: 401 EVSNRDVVKKIMNAWKAMVESPNQQLYANALVEFKDSCSDFSIFVDYAMTTLDEVKDKI 459
+ D V KI+N K +V + +Y L E +D ++ ++FVD+ +++ E+ ++
Sbjct: 1199 DAERSDCVTKILNKLKGLVATVEPTVYHQTLNEIEDDLNERNLFVDFELSSDSEMLQQL 1257
>HTPG_BACFR (P58476) Chaperone protein htpG (Heat shock protein
htpG) (High temperature protein G)
Length = 681
Score = 33.9 bits (76), Expect = 0.90
Identities = 16/51 (31%), Positives = 26/51 (50%)
Query: 408 VKKIMNAWKAMVESPNQQLYANALVEFKDSCSDFSIFVDYAMTTLDEVKDK 458
VKKI V Q ++ N +F++ +D IF++Y M T ++ DK
Sbjct: 335 VKKISTYISKKVSDRLQSIFKNDRAQFEEKWNDLKIFINYGMLTQEDFYDK 385
>TRA1_LACHE (P35880) Transposase for insertion sequence element
IS1201
Length = 369
Score = 33.1 bits (74), Expect = 1.5
Identities = 40/211 (18%), Positives = 90/211 (41%), Gaps = 15/211 (7%)
Query: 242 LISKLEEHNYTYYSRTQLESNTIEDIFWAHPTSIKLFNNFPTVLVMDSTY----KTNMYR 297
LI K+ +Y+ + + I + H KL + F V +D+TY + R
Sbjct: 120 LIEKMYGSHYSPAQVSNISKQMIPKVEAYHKR--KLSDKFFCVY-LDATYVPLRRETFER 176
Query: 298 MPMFEVVGVTSTDLTYSVGFGFMTHEKEENFVWVLTMLRKLLSSKMNMPKVIVTDRDMSL 357
++ +G+ + + +E E VW +L+ + S + ++ ++D + +
Sbjct: 177 EAVYIAIGIKPNGHKEVIDYCIAPNENIE--VWT-ELLQSMKSRGLEQVELFLSDGVVGM 233
Query: 358 MKAVAHVFPESYALNCFFHVQANVKQRC-VLNCKYPLGFKKDGKEVSNRDVVKKIMNAWK 416
A+A +P+++ C HV N+ + V + + + K + +N+ +++A+
Sbjct: 234 KTALAKTYPQAHFQRCLVHVMRNICAKVRVEDREAIMNEFKQIHQQANKAAAVDVLHAFY 293
Query: 417 AMVESPNQQLYANALVEFKDSCSDFSIFVDY 447
A + + Y + + KD D +F +Y
Sbjct: 294 AKWD----KSYNHVIRNLKDIEPDLLVFYNY 320
>POLG_TUMVJ (P89509) Genome polyprotein [Contains: P1 proteinase
(N-terminal protein); Helper component proteinase (EC
3.4.22.45) (HC-pro); Protein P3; 6 kDa protein 1 (6K1);
Cytoplasmic inclusion protein (CI); 6 kDa protein 2
(6K2); Viral genome-linked p
Length = 3164
Score = 32.7 bits (73), Expect = 2.0
Identities = 14/51 (27%), Positives = 28/51 (54%)
Query: 401 EVSNRDVVKKIMNAWKAMVESPNQQLYANALVEFKDSCSDFSIFVDYAMTT 451
+ D V KI+N K +V + +Y L + +D S+ ++FVD+ +++
Sbjct: 1199 DAERSDCVTKILNKLKGLVATVEPTVYHQTLNDIEDDLSERNLFVDFELSS 1249
>PAC2_CHICK (O13154) Protein kinase C and casein kinase substrate in
neurons protein 2 (Focal adhesion protein of 52 kDa)
(FAP52)
Length = 448
Score = 32.7 bits (73), Expect = 2.0
Identities = 31/118 (26%), Positives = 56/118 (47%), Gaps = 21/118 (17%)
Query: 166 NHPMEPALEGHILAGRLKEDDKKIVRDLTKSKMLPRNILIHLKNQRPHCMTNVKQVYNER 225
N +PAL L +L++ ++ +D+ K+K L L N P M N++QV+ +
Sbjct: 175 NSKADPALNPEQLK-KLQDKVERSKQDVLKTKAKYEKSLKELDNATPQYMENMEQVFEQC 233
Query: 226 QQIWKANRGDKKSLQF---LISKLEEH----NYTYYSR--TQLESN-----TIEDIFW 269
QQ ++K L+F ++ ++++H N Y +LE N +ED+ W
Sbjct: 234 QQF------EEKRLRFFREVLLEVQKHLDLSNVASYKNIYRELEQNIKTADAVEDLRW 285
>SUHW_DROAN (Q08875) Suppressor of hairy wing protein
Length = 886
Score = 32.3 bits (72), Expect = 2.6
Identities = 26/124 (20%), Positives = 48/124 (37%), Gaps = 10/124 (8%)
Query: 115 VCERSGSHIVPEKKPKHANTGSRKCGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALE 174
+C++ S +V KK + +TG + C + K+ + G H E +
Sbjct: 446 LCDKKLSALVALKKHRRYHTGEKPYSCTVCNQAFAVKEVLNRHMKRHTGERPHKCEECGK 505
Query: 175 GHILAGRLKEDDKKIVRDLT---------KSKMLPRNILIHLKNQRPH-CMTNVKQVYNE 224
I A +L+ K +R K L R++ H + +RP+ T K+ +
Sbjct: 506 SFIQATQLRTHSKTHIRPFACDMCEEKFKTEKQLERHVKEHSRQKRPYFSCTECKRHFRN 565
Query: 225 RQQI 228
Q+
Sbjct: 566 TAQL 569
>YAB2_SCHPO (Q09804) Hypothetical protein C2G11.02 in chromosome I
Length = 1318
Score = 32.0 bits (71), Expect = 3.4
Identities = 18/60 (30%), Positives = 32/60 (53%), Gaps = 2/60 (3%)
Query: 72 FFSDDVKSKNRDELLE--WVRRQANKAGFTIVTQRSSLINPMFRLVCERSGSHIVPEKKP 129
+F D + + +D +LE W R ++ + + + R SLI+ + RL+C S + I P P
Sbjct: 666 YFIDSLNLETKDSVLESNWERFISSTETYELDSIRDSLIDCLIRLLCYSSNNAIDPVSAP 725
>YNH9_SCHPO (Q96WW0) Hypothetical WD-repeat protein C32H8.09 in
chromosome II
Length = 483
Score = 31.6 bits (70), Expect = 4.5
Identities = 16/78 (20%), Positives = 37/78 (46%), Gaps = 6/78 (7%)
Query: 228 IWKANRGDKKSLQFLISKLEEH------NYTYYSRTQLESNTIEDIFWAHPTSIKLFNNF 281
+WK N G +K Q ++ +++ + YY+ Q + + + I W+ + L+++F
Sbjct: 61 LWKPNLGAEKCHQICVASVDKVFVLDIVQHDYYASIQCDQDPLSSISWSPSGELLLWSSF 120
Query: 282 PTVLVMDSTYKTNMYRMP 299
+ + + S Y +P
Sbjct: 121 DSKITVWSLNTQKGYLLP 138
>EX5B_BUCBP (Q89AB3) Exodeoxyribonuclease V beta chain (EC 3.1.11.5)
Length = 1180
Score = 31.6 bits (70), Expect = 4.5
Identities = 18/83 (21%), Positives = 38/83 (45%)
Query: 196 SKMLPRNILIHLKNQRPHCMTNVKQVYNERQQIWKANRGDKKSLQFLISKLEEHNYTYYS 255
+K L + I KN+ ++ + + + QI N + + + +S+ +NY + S
Sbjct: 851 TKKLYTELSILKKNKNIEIISEIPNIIKKNFQIPTLNTNSQSLMHYQVSRKFNYNYNFTS 910
Query: 256 RTQLESNTIEDIFWAHPTSIKLF 278
+QL+ N ++ + KLF
Sbjct: 911 YSQLKKNIKPSTMYSTLNTKKLF 933
>CIC2_RABIT (P13806) Dihydropyridine-sensitive L-type, calcium
channel alpha-2/delta subunits precursor
Length = 1106
Score = 31.6 bits (70), Expect = 4.5
Identities = 26/83 (31%), Positives = 38/83 (45%), Gaps = 10/83 (12%)
Query: 7 VMVKNAKDVPGETKENFVENSGDVKP-----PNFEAKTNIDEASVHVEASKYVGGSGVLP 61
V+ NAKD K + S +KP NF + + A+VH+ Y G + VL
Sbjct: 124 VVYYNAKDDLDPEKNDSEPGSQRIKPVFIDDANFRRQVSYQHAAVHIPTDIYEGSTIVL- 182
Query: 62 LQPHEVDTAKFFSDDVKSKNRDE 84
+E++ DDV KNR+E
Sbjct: 183 ---NELNWTSAL-DDVFKKNREE 201
>RIN2_MOUSE (Q9D684) Ras and Rab interactor 2 (Ras
interaction/interference protein 2)
Length = 903
Score = 31.2 bits (69), Expect = 5.9
Identities = 32/150 (21%), Positives = 60/150 (39%), Gaps = 12/150 (8%)
Query: 13 KDVPGETKENFVENSGDVKPPNFEAKTNIDEASVHVEASKYVGGSGVLPLQPHEVDTAKF 72
+D PG+ S PP A+ + + S+ +S + +PL +E DT
Sbjct: 410 RDAPGDCTRAPPPGSESQPPPCHGARQRLSDMSLSTSSSDSLEFDRSMPLYGYEADTTSS 469
Query: 73 FSDDVKSKNRDELLEWVRRQANKAGFTIVTQRSSLINPMFRLVCERSGSHIVPEKK--PK 130
D +++ + ++ + + ++ + L+ R + S + PEK+ +
Sbjct: 470 LEDYEGESDQETMAPPIKSKKKRNSSFVLPK---LVKSQLRKMSGVFSSFMTPEKRMVRR 526
Query: 131 HANTGSRKC---GCLFMISGYQS--KQTKE 155
A KC GCL + Y S K+ KE
Sbjct: 527 IAELSRDKCTYFGCL--VQDYVSFLKENKE 554
>YOH5_YEAST (Q08234) Probable ATP-dependent transporter
YOL074C/YOL075C
Length = 1294
Score = 30.8 bits (68), Expect = 7.6
Identities = 18/53 (33%), Positives = 31/53 (57%), Gaps = 3/53 (5%)
Query: 396 KKDGKEVSNRDVVKKIMNAWKAMVESPNQQLYANALVEFKDSCSDFSIFVDYA 448
+ + E+S+R V+KI++AWKA ++ N+ L + E K S S F +Y+
Sbjct: 958 QNEQNEISSRARVEKILSAWKANMD--NESLSPTPISE-KQQYSQESFFTEYS 1007
>TRIB_HUMAN (Q14669) Thyroid receptor interacting protein 12 (TRIP12)
Length = 1992
Score = 30.8 bits (68), Expect = 7.6
Identities = 17/63 (26%), Positives = 31/63 (48%), Gaps = 10/63 (15%)
Query: 33 PNFEAKTNIDEASVHVEASKYVGGSGVLPLQPHEVDTAKFFSDDVKSKNRDELLEWVRRQ 92
P + + S ++E ++ GGSG+ A+ S D S NR+++ W++ Q
Sbjct: 1047 PKTWGRLSTQSNSNNIEPARTAGGSGL----------ARAASKDTISNNREKIKGWIKEQ 1096
Query: 93 ANK 95
A+K
Sbjct: 1097 AHK 1099
>SUHW_DROME (P08970) Suppressor of hairy wing protein
Length = 941
Score = 30.8 bits (68), Expect = 7.6
Identities = 22/107 (20%), Positives = 41/107 (37%), Gaps = 9/107 (8%)
Query: 115 VCERSGSHIVPEKKPKHANTGSRKCGCLFMISGYQSKQTKEWGLNILNGVHNHPMEPALE 174
+C++ S +V KK + +TG + C + K+ + G H + +
Sbjct: 445 LCDKKFSALVALKKHRRYHTGEKPYSCTVCNQAFAVKEVLNRHMKRHTGERPHKCDECGK 504
Query: 175 GHILAGRLKEDDKKIVR---------DLTKSKMLPRNILIHLKNQRP 212
I A +L+ K +R K L R++ H + +RP
Sbjct: 505 SFIQATQLRTHSKTHIRPFPCEQCDEKFKTEKQLERHVKTHSRTKRP 551
>DCR1_DROME (Q9VCU9) Endoribonuclease Dcr-1 (EC 3.1.26.-) (Dicer-1
protein)
Length = 2249
Score = 30.8 bits (68), Expect = 7.6
Identities = 16/63 (25%), Positives = 29/63 (45%)
Query: 398 DGKEVSNRDVVKKIMNAWKAMVESPNQQLYANALVEFKDSCSDFSIFVDYAMTTLDEVKD 457
D K +S+ + + + A+ N L + DSC+DFS F+ Y + + + D
Sbjct: 1841 DIKNLSSVQICEMVREKADALGLEQNGGAQNGQLDDSNDSCNDFSCFIPYNLVSQHSIPD 1900
Query: 458 KIV 460
K +
Sbjct: 1901 KSI 1903
>GLF_MYCGE (Q49398) UDP-galactopyranose mutase (EC 5.4.99.9)
Length = 404
Score = 30.4 bits (67), Expect = 10.0
Identities = 18/68 (26%), Positives = 33/68 (48%)
Query: 343 MNMPKVIVTDRDMSLMKAVAHVFPESYALNCFFHVQANVKQRCVLNCKYPLGFKKDGKEV 402
+N K++++++ A+ P+S N F + AN LNCK L +D K
Sbjct: 192 INRVKIVLSEQSSYFPDAIIQGLPKSGYTNSFLKMLANPLIDVQLNCKDNLLVYQDEKLF 251
Query: 403 SNRDVVKK 410
N ++++K
Sbjct: 252 FNNNLIEK 259
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.319 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 53,230,066
Number of Sequences: 164201
Number of extensions: 2165228
Number of successful extensions: 5243
Number of sequences better than 10.0: 19
Number of HSP's better than 10.0 without gapping: 8
Number of HSP's successfully gapped in prelim test: 11
Number of HSP's that attempted gapping in prelim test: 5233
Number of HSP's gapped (non-prelim): 21
length of query: 461
length of database: 59,974,054
effective HSP length: 114
effective length of query: 347
effective length of database: 41,255,140
effective search space: 14315533580
effective search space used: 14315533580
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC139743.2


