

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139600.10 + phase: 0
(81 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
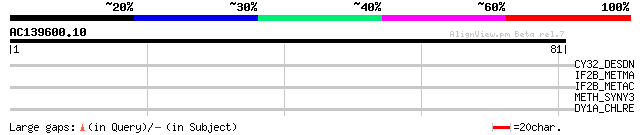
Score E
Sequences producing significant alignments: (bits) Value
CY32_DESDN (P38554) Cytochrome c3, 26 kDa 28 3.2
IF2B_METMA (Q8PWV1) Probable translation initiation factor 2 bet... 28 4.2
IF2B_METAC (Q8TU91) Probable translation initiation factor 2 bet... 28 4.2
METH_SYNY3 (Q55786) Methionine synthase (EC 2.1.1.13) (5-methylt... 27 7.2
DY1A_CHLRE (Q9SMH3) Dynein 1-alpha heavy chain, flagellar inner ... 27 7.2
>CY32_DESDN (P38554) Cytochrome c3, 26 kDa
Length = 111
Score = 28.5 bits (62), Expect = 3.2
Identities = 15/46 (32%), Positives = 22/46 (47%), Gaps = 1/46 (2%)
Query: 20 KYGECQKNHAANVGGYAVDGCREFMPSTNGSLTCAACGCHRNFHKR 65
K G+ NHA+++ A C +P T +C GCH N +R
Sbjct: 22 KKGDVTFNHASHMD-IACQQCHHTVPDTYTIESCMTEGCHDNIKER 66
>IF2B_METMA (Q8PWV1) Probable translation initiation factor 2 beta
subunit (eIF-2-beta)
Length = 202
Score = 28.1 bits (61), Expect = 4.2
Identities = 13/31 (41%), Positives = 16/31 (50%)
Query: 51 LTCAACGCHRNFHKREVEVVSESSSPHSNGT 81
L C+ACG HR KR+V V + GT
Sbjct: 121 LKCSACGAHRPVKKRKVSNVVVREAIEEGGT 151
>IF2B_METAC (Q8TU91) Probable translation initiation factor 2 beta
subunit (eIF-2-beta)
Length = 202
Score = 28.1 bits (61), Expect = 4.2
Identities = 13/31 (41%), Positives = 16/31 (50%)
Query: 51 LTCAACGCHRNFHKREVEVVSESSSPHSNGT 81
L C+ACG HR KR+V V + GT
Sbjct: 121 LKCSACGAHRPVKKRKVSNVVVRDAIEEGGT 151
>METH_SYNY3 (Q55786) Methionine synthase (EC 2.1.1.13)
(5-methyltetrahydrofolate--homocysteine
methyltransferase) (Methionine synthase, vitamin-B12
dependent isozyme) (MS)
Length = 1195
Score = 27.3 bits (59), Expect = 7.2
Identities = 19/68 (27%), Positives = 29/68 (41%), Gaps = 10/68 (14%)
Query: 10 RSEDPQRSVVKYG-------ECQKNHAANVGGYAVDGCREFMPSTNGSLTCAACGCHRNF 62
R P R V+ + + Q AA+ GG +GC E++ T A HR F
Sbjct: 8 RIHSPDRPVLVFDGAMGTNLQVQNLTAADFGGAEYEGCNEYLVHTKPE---AVATVHRAF 64
Query: 63 HKREVEVV 70
++ +VV
Sbjct: 65 YEAGADVV 72
>DY1A_CHLRE (Q9SMH3) Dynein 1-alpha heavy chain, flagellar inner arm
I1 complex (1-alpha DHC) (Dynein 1, subspecies f)
Length = 4625
Score = 27.3 bits (59), Expect = 7.2
Identities = 13/32 (40%), Positives = 18/32 (55%)
Query: 48 NGSLTCAACGCHRNFHKREVEVVSESSSPHSN 79
+G + C A GC F++ E EV+S SS N
Sbjct: 1998 SGLVQCGAWGCFDEFNRIEAEVLSVVSSQIKN 2029
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.316 0.129 0.395
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,551,101
Number of Sequences: 164201
Number of extensions: 315894
Number of successful extensions: 752
Number of sequences better than 10.0: 5
Number of HSP's better than 10.0 without gapping: 3
Number of HSP's successfully gapped in prelim test: 2
Number of HSP's that attempted gapping in prelim test: 749
Number of HSP's gapped (non-prelim): 5
length of query: 81
length of database: 59,974,054
effective HSP length: 57
effective length of query: 24
effective length of database: 50,614,597
effective search space: 1214750328
effective search space used: 1214750328
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC139600.10


