

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139356.2 + phase: 0
(180 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
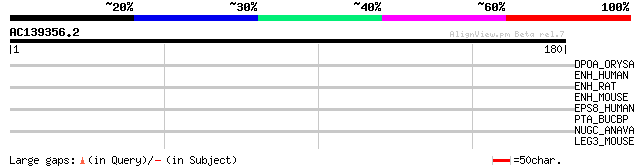
Score E
Sequences producing significant alignments: (bits) Value
DPOA_ORYSA (O48653) DNA polymerase alpha catalytic subunit (EC 2... 32 1.0
ENH_HUMAN (Q96HC4) Enigma homolog (Enigma-like PDZ and LIM domai... 30 3.9
ENH_RAT (Q62920) Enigma homolog (Enigma-like PDZ and LIM domains... 29 5.2
ENH_MOUSE (Q8CI51) Enigma homolog (Enigma-like PDZ and LIM domai... 29 5.2
EPS8_HUMAN (Q12929) Epidermal growth factor receptor kinase subs... 29 6.7
PTA_BUCBP (Q89AS7) Phosphate acetyltransferase (EC 2.3.1.8) (Pho... 28 8.8
NUGC_ANAVA (Q9XBL6) NAD(P)H-quinone oxidoreductase subunit J (EC... 28 8.8
LEG3_MOUSE (P16110) Galectin-3 (Galactose-specific lectin 3) (Ma... 28 8.8
>DPOA_ORYSA (O48653) DNA polymerase alpha catalytic subunit (EC
2.7.7.7)
Length = 1243
Score = 31.6 bits (70), Expect = 1.0
Identities = 27/100 (27%), Positives = 40/100 (40%), Gaps = 7/100 (7%)
Query: 53 VQCKSCQTKFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKILMNPRLPRCAHCNQENSVK 112
+ C SC T F P + + S V+N N +D + R PRC E+ V
Sbjct: 1047 LSCPSCSTTFDCP-PVSSLIIGSSSGNVSNPNEGNDASINFWRRMRCPRCPDDTDESRVS 1105
Query: 113 PVIA--EKKKNINWLFLLLGQMLGCCKLEQLKYFCKYNNH 150
P + + K+ + L + L C E CKY+ H
Sbjct: 1106 PAVLANQMKRQADSFINLYYKGLLMCDDEG----CKYSTH 1141
>ENH_HUMAN (Q96HC4) Enigma homolog (Enigma-like PDZ and LIM domains
protein)
Length = 596
Score = 29.6 bits (65), Expect = 3.9
Identities = 13/33 (39%), Positives = 18/33 (54%)
Query: 75 ISQYLVTNMNTMHDRAPKILMNPRLPRCAHCNQ 107
+ Q ++ +T+ RA I R P CAHCNQ
Sbjct: 393 LGQTQPSDQDTLVQRAEHIPAGKRTPMCAHCNQ 425
>ENH_RAT (Q62920) Enigma homolog (Enigma-like PDZ and LIM domains
protein)
Length = 591
Score = 29.3 bits (64), Expect = 5.2
Identities = 12/27 (44%), Positives = 16/27 (58%)
Query: 81 TNMNTMHDRAPKILMNPRLPRCAHCNQ 107
++ +T+ RA I R P CAHCNQ
Sbjct: 394 SDQDTLVQRAEHIPAGKRTPMCAHCNQ 420
>ENH_MOUSE (Q8CI51) Enigma homolog (Enigma-like PDZ and LIM domains
protein)
Length = 591
Score = 29.3 bits (64), Expect = 5.2
Identities = 12/27 (44%), Positives = 16/27 (58%)
Query: 81 TNMNTMHDRAPKILMNPRLPRCAHCNQ 107
++ +T+ RA I R P CAHCNQ
Sbjct: 394 SDQDTLVQRAEHIPAGKRTPMCAHCNQ 420
>EPS8_HUMAN (Q12929) Epidermal growth factor receptor kinase
substrate EPS8
Length = 822
Score = 28.9 bits (63), Expect = 6.7
Identities = 33/147 (22%), Positives = 56/147 (37%), Gaps = 10/147 (6%)
Query: 9 PPRRRSAPRSGPIPIPFIWATNHRAKIHNL--NHLLQNKIFSITGDVQCKSCQTKFQMPF 66
PP P P+P+P +T + + N QN S +G + Q Q+P
Sbjct: 621 PPAPSPPPTPAPVPVPLPPSTPAPVPVSKVPANITRQNSSSSDSGGSIVRDSQRHKQLPV 680
Query: 67 DLR-TKFDVISQYLVTNMNTMHDRAPKILMNPR----LPRCAHCNQENSVKPVIAEKKKN 121
D R ++ + + L+ + A K PR + + + VK + K N
Sbjct: 681 DRRKSQMEEVQDELIHRLTIGRSAAQKKFHVPRQNVPVINITYDSTPEDVKTWLQSKGFN 740
Query: 122 ---INWLFLLLGQMLGCCKLEQLKYFC 145
+N L +L G L ++L+ C
Sbjct: 741 PVTVNSLGVLNGAQLFSLNKDELRTVC 767
>PTA_BUCBP (Q89AS7) Phosphate acetyltransferase (EC 2.3.1.8)
(Phosphotransacetylase)
Length = 715
Score = 28.5 bits (62), Expect = 8.8
Identities = 12/43 (27%), Positives = 22/43 (50%)
Query: 74 VISQYLVTNMNTMHDRAPKILMNPRLPRCAHCNQENSVKPVIA 116
+IS L+TN +HD+ + ++ C C+ N V V++
Sbjct: 301 LISAILLTNFLNVHDKCTDFIDQLKIANCTICSTPNDVFTVVS 343
>NUGC_ANAVA (Q9XBL6) NAD(P)H-quinone oxidoreductase subunit J (EC
1.6.5.-) (NAD(P)H dehydrogenase I, subunit J) (NDH-1,
subunit J)
Length = 175
Score = 28.5 bits (62), Expect = 8.8
Identities = 19/64 (29%), Positives = 30/64 (46%), Gaps = 6/64 (9%)
Query: 61 KFQMPFDLRTKFDVISQYLVTNMNTMHDRAPKILMNPRLPRCAHCNQENSVKPVIAEKKK 120
+FQ DLR D++S Y + ++ D+ +I + LPR EN V P + K
Sbjct: 69 QFQRRIDLRPGQDLVSVYHLVKVSDNADKPEEIRVKVFLPR------ENPVVPSVYWIGK 122
Query: 121 NINW 124
+W
Sbjct: 123 TADW 126
>LEG3_MOUSE (P16110) Galectin-3 (Galactose-specific lectin 3) (Mac-2
antigen) (IgE-binding protein) (35 kDa lectin)
(Carbohydrate binding protein 35) (CBP 35)
(Laminin-binding protein) (Lectin L-29) (L-34
galactoside-binding lectin)
Length = 263
Score = 28.5 bits (62), Expect = 8.8
Identities = 18/66 (27%), Positives = 34/66 (51%), Gaps = 5/66 (7%)
Query: 64 MPFDLRTKFDVISQYLVTNMNTMHDRAPKILMNPRLPR--CAHCN---QENSVKPVIAEK 118
+P+DL V+ + L+T M T+ A +I+++ R H N EN+ + ++
Sbjct: 129 VPYDLPLPGGVMPRMLITIMGTVKPNANRIVLDFRRGNDVAFHFNPRFNENNRRVIVCNT 188
Query: 119 KKNINW 124
K++ NW
Sbjct: 189 KQDNNW 194
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.325 0.138 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,894,475
Number of Sequences: 164201
Number of extensions: 835390
Number of successful extensions: 2217
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 4
Number of HSP's that attempted gapping in prelim test: 2213
Number of HSP's gapped (non-prelim): 8
length of query: 180
length of database: 59,974,054
effective HSP length: 103
effective length of query: 77
effective length of database: 43,061,351
effective search space: 3315724027
effective search space used: 3315724027
T: 11
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC139356.2


