

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.8 + phase: 0 /pseudo
(481 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
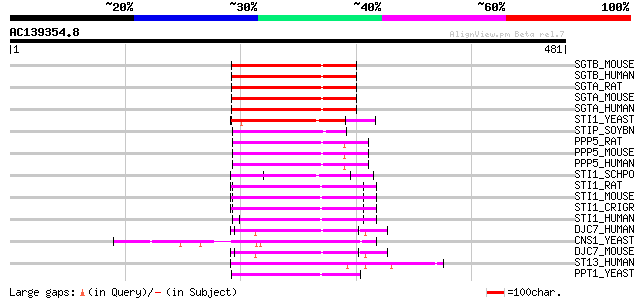
Score E
Sequences producing significant alignments: (bits) Value
SGTB_MOUSE (Q8VD33) Small glutamine-rich tetratricopeptide repea... 104 6e-22
SGTB_HUMAN (Q96EQ0) Small glutamine-rich tetratricopeptide repea... 103 1e-21
SGTA_RAT (O70593) Small glutamine-rich tetratricopeptide repeat-... 102 3e-21
SGTA_MOUSE (Q8BJU0) Small glutamine-rich tetratricopeptide repea... 102 3e-21
SGTA_HUMAN (O43765) Small glutamine-rich tetratricopeptide repea... 99 2e-20
STI1_YEAST (P15705) Heat shock protein STI1 78 6e-14
STIP_SOYBN (Q43468) Heat shock protein STI (Stress inducible pro... 68 5e-11
PPP5_RAT (P53042) Serine/threonine protein phosphatase 5 (EC 3.1... 66 2e-10
PPP5_MOUSE (Q60676) Serine/threonine protein phosphatase 5 (EC 3... 65 3e-10
PPP5_HUMAN (P53041) Serine/threonine protein phosphatase 5 (EC 3... 64 9e-10
STI1_SCHPO (Q9USI5) Heat shock protein sti1 homolog 63 2e-09
STI1_RAT (O35814) Stress-induced-phosphoprotein 1 (STI1) (Hsc70/... 63 2e-09
STI1_MOUSE (Q60864) Stress-induced-phosphoprotein 1 (STI1) (Hsc7... 63 2e-09
STI1_CRIGR (O54981) Stress-induced-phosphoprotein 1 (STI1) (Hsc7... 63 2e-09
STI1_HUMAN (P31948) Stress-induced-phosphoprotein 1 (STI1) (Hsc7... 61 6e-09
DJC7_HUMAN (Q99615) DnaJ homolog subfamily C member 7 (Tetratric... 60 1e-08
CNS1_YEAST (P33313) Cyclophilin seven suppressor 1 (STI1 stress-... 59 2e-08
DJC7_MOUSE (Q9QYI3) DnaJ homolog subfamily C member 7 (Tetratric... 58 5e-08
ST13_HUMAN (P50502) Hsc70-interacting protein (Hip) (Putative tu... 57 1e-07
PPT1_YEAST (P53043) Serine/threonine protein phosphatase T (EC 3... 57 1e-07
>SGTB_MOUSE (Q8VD33) Small glutamine-rich tetratricopeptide
repeat-containing protein B
Length = 304
Score = 104 bits (259), Expect = 6e-22
Identities = 50/108 (46%), Positives = 74/108 (68%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + Y A++ Y AI + +AVYYCNRAAA ++++ YT+AI+D ++I ID YSKA
Sbjct: 96 MKEENYAAAVDCYTQAIELDPNNAVYYCNRAAAQSKLSHYTDAIKDCEKAIAIDSKYSKA 155
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA A + +A+ ++KAL LDP N+S K N+++AE KL E
Sbjct: 156 YGRMGLALTAMNKFEEAV-TSYQKALDLDPENDSYKSNLKIAEQKLRE 202
>SGTB_HUMAN (Q96EQ0) Small glutamine-rich tetratricopeptide
repeat-containing protein B (Small glutamine-rich
protein with tetratricopeptide repeats 2)
Length = 304
Score = 103 bits (256), Expect = 1e-21
Identities = 50/108 (46%), Positives = 73/108 (67%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + Y A++ Y AI + +AVYYCNRAAA +++ YT+AI+D ++I ID YSKA
Sbjct: 96 MKEENYAAAVDCYTQAIELDPNNAVYYCNRAAAQSKLGHYTDAIKDCEKAIAIDSKYSKA 155
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA A + +A+ ++KAL LDP N+S K N+++AE KL E
Sbjct: 156 YGRMGLALTALNKFEEAV-TSYQKALDLDPENDSYKSNLKIAEQKLRE 202
>SGTA_RAT (O70593) Small glutamine-rich tetratricopeptide
repeat-containing protein A
Length = 314
Score = 102 bits (253), Expect = 3e-21
Identities = 50/108 (46%), Positives = 75/108 (69%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + + A+ LY AI + +AVY+CNRAAAY+++ Y A+QD R+I IDP YSKA
Sbjct: 102 MKLENFEAAVHLYGKAIELNPANAVYFCNRAAAYSKLGNYVGAVQDCERAIGIDPGYSKA 161
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA + + +A+ +KKAL+LDP+N++ K N+++AE KL E
Sbjct: 162 YGRMGLALSSLNKHAEAV-AYYKKALELDPDNDTYKSNLKIAELKLRE 208
>SGTA_MOUSE (Q8BJU0) Small glutamine-rich tetratricopeptide
repeat-containing protein A
Length = 315
Score = 102 bits (253), Expect = 3e-21
Identities = 50/108 (46%), Positives = 75/108 (69%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + + A+ LY AI + +AVY+CNRAAAY+++ Y A+QD R+I IDP YSKA
Sbjct: 103 MKLENFEAAVHLYGKAIELNPANAVYFCNRAAAYSKLGNYVGAVQDCERAIGIDPGYSKA 162
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA + + +A+ +KKAL+LDP+N++ K N+++AE KL E
Sbjct: 163 YGRMGLALSSLNKHAEAV-AYYKKALELDPDNDTYKSNLKIAELKLRE 209
>SGTA_HUMAN (O43765) Small glutamine-rich tetratricopeptide
repeat-containing protein A (Vpu-binding protein) (UBP)
Length = 313
Score = 99.4 bits (246), Expect = 2e-20
Identities = 50/108 (46%), Positives = 74/108 (68%), Gaps = 1/108 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
M+ + + A+ Y AI + +AVY+CNRAAAY+++ Y A+QD R+I IDP YSKA
Sbjct: 102 MKVENFEAAVHFYGKAIELNPANAVYFCNRAAAYSKLGNYAGAVQDCERAICIDPAYSKA 161
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLME 300
Y R+GLA + + +A+ +KKAL+LDP+NE+ K N+++AE KL E
Sbjct: 162 YGRMGLALSSLNKHVEAV-AYYKKALELDPDNETYKSNLKIAELKLRE 208
>STI1_YEAST (P15705) Heat shock protein STI1
Length = 589
Score = 77.8 bits (190), Expect = 6e-14
Identities = 39/101 (38%), Positives = 66/101 (64%), Gaps = 2/101 (1%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEK-SAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYS 250
A +K Y AIEL+ AI + E + V Y NR+A YT + ++++A+ D+ ++I+P++S
Sbjct: 15 AFTAKDYDKAIELFTKAIEVSETPNHVLYSNRSACYTSLKKFSDALNDANECVKINPSWS 74
Query: 251 KAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENI 291
K Y+RLG A+ G+ D + +KKAL+LD +N++ KE +
Sbjct: 75 KGYNRLGAAHLGLGDL-DEAESNYKKALELDASNKAAKEGL 114
Score = 52.4 bits (124), Expect = 3e-06
Identities = 36/132 (27%), Positives = 62/132 (46%), Gaps = 7/132 (5%)
Query: 193 MQSKQYF------DAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEID 246
++ K+YF +A++ Y I + A Y NRAAA ++ + EAI D ++IE D
Sbjct: 401 LEGKEYFTKSDWPNAVKAYTEMIKRAPEDARGYSNRAAALAKLMSFPEAIADCNKAIEKD 460
Query: 247 PNYSKAYSRLGLAYYAQGNYRDAIDK-GFKKALQLDPNNESVKENIRVAEHKLMEERHRA 305
PN+ +AY R A A Y A++ + + NN S I +K ++R +
Sbjct: 461 PNFVRAYIRKATAQIAVKEYASALETLDAARTKDAEVNNGSSAREIDQLYYKASQQRFQP 520
Query: 306 DHNQNSRSQEFQ 317
+ + + +Q
Sbjct: 521 GTSNETPEETYQ 532
Score = 39.3 bits (90), Expect = 0.023
Identities = 34/119 (28%), Positives = 62/119 (51%), Gaps = 10/119 (8%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIE----IDPNY 249
+++Q+ +AIE YN A ++ K Y NRAAA + Y AI ++E + +Y
Sbjct: 274 KARQFDEAIEHYNKAWELH-KDITYLNNRAAAEYEKGEYETAISTLNDAVEQGREMRADY 332
Query: 250 ---SKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRA 305
SK+++R+G AY+ G+ + I+ ++K+L + + +R AE +L + A
Sbjct: 333 KVISKSFARIGNAYHKLGDLKKTIEY-YQKSL-TEHRTADILTKLRNAEKELKKAEAEA 389
>STIP_SOYBN (Q43468) Heat shock protein STI (Stress inducible
protein) (GmSTI)
Length = 569
Score = 68.2 bits (165), Expect = 5e-11
Identities = 34/99 (34%), Positives = 59/99 (59%), Gaps = 1/99 (1%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
+ ++Y +A + Y AI K A Y NRAA YT++ E ++D+ + IE+DP +SK Y
Sbjct: 393 KQQKYPEATKHYTEAIKRNPKDAKAYSNRAACYTKLGAMPEGLKDAEKCIELDPTFSKGY 452
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIR 292
+R G ++ Y A++ +++ L+ DPNN+ + + IR
Sbjct: 453 TRKGAVQFSMKEYDKALET-YREGLKHDPNNQELLDGIR 490
Score = 43.9 bits (102), Expect = 0.001
Identities = 30/104 (28%), Positives = 53/104 (50%), Gaps = 2/104 (1%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + + A+ ++ AIA+ + V Y NR+AA T +++++ P++ K
Sbjct: 12 AFSAGDFAAAVRHFSDAIALSPSNHVLYSNRSAA-TLPPELRGGPSRRQKTVDLKPDWPK 70
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAE 295
AYSRLG A+ +RDA K A +P+N ++K + A+
Sbjct: 71 AYSRLGAAHLGLRRHRDA-SPPTKPASNSNPDNAALKSGLADAQ 113
Score = 39.3 bits (90), Expect = 0.023
Identities = 43/201 (21%), Positives = 85/201 (41%), Gaps = 14/201 (6%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A + K + AI Y+ A+ + ++ Y NRAA Y ++ ++ + I+D +++E
Sbjct: 252 AYKKKDFETAIGHYSKALELDDEDISYLTNRAAVYLEMGKFEDCIKDCEKAVERGKELRS 311
Query: 252 AYSRLGLAYYAQG-----------NYRDAIDKGFKKALQLDPNNESVKE-NIRVAEHKLM 299
Y + A +G ++ AI+ F+KAL + N +++K+ N K +
Sbjct: 312 DYKMIARALTRKGTALAKMAKCSKDFEPAIEI-FQKALTENRNPDTLKKLNEAEKAKKEL 370
Query: 300 EERHRADHNQNSRSQEFQNHYARGSRSHAAPASF-GSMPFNPSNLASMFMAAANGGQGSH 358
E++ D ++E N + + A + ++ NP + + AA +
Sbjct: 371 EQQEYFDPKLADEAREKGNELFKQQKYPEATKHYTEAIKRNPKDAKAYSNRAACYTKLGA 430
Query: 359 SQEGQEDANSSGANEPEIRFG 379
EG +DA +P G
Sbjct: 431 MPEGLKDAEKCIELDPTFSKG 451
>PPP5_RAT (P53042) Serine/threonine protein phosphatase 5 (EC
3.1.3.16) (PP5) (Protein phosphatase T) (PPT)
Length = 499
Score = 66.2 bits (160), Expect = 2e-10
Identities = 37/122 (30%), Positives = 67/122 (54%), Gaps = 5/122 (4%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
++K Y +AI+ Y+ AI + +A+YY NR+ AY + Y A+ D+ R+IE+D Y K Y
Sbjct: 40 KAKDYENAIKFYSQAIELNPSNAIYYGNRSLAYLRTECYGYALGDATRAIELDKKYIKGY 99
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVK----ENIRVAEHKLMEERHRADHNQ 309
R + A G +R A+ + ++ +++ PN++ K E ++ + K E D ++
Sbjct: 100 YRRAASNMALGKFRAAL-RDYETVVKVKPNDKDAKMKYQECSKIVKQKAFERAIAGDEHR 158
Query: 310 NS 311
S
Sbjct: 159 RS 160
>PPP5_MOUSE (Q60676) Serine/threonine protein phosphatase 5 (EC
3.1.3.16) (PP5) (Protein phosphatase T) (PPT)
Length = 499
Score = 65.5 bits (158), Expect = 3e-10
Identities = 37/122 (30%), Positives = 67/122 (54%), Gaps = 5/122 (4%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
++K Y +AI+ Y+ AI + +A+YY NR+ AY + Y A+ D+ R+IE+D Y K Y
Sbjct: 40 KAKDYENAIKFYSQAIELNPGNAIYYGNRSLAYLRTECYGYALGDATRAIELDKKYIKGY 99
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVK----ENIRVAEHKLMEERHRADHNQ 309
R + A G +R A+ + ++ +++ PN++ K E ++ + K E D ++
Sbjct: 100 YRRAASNMALGKFRAAL-RDYETVVKVKPNDKDAKMKYQECSKIVKQKAFERAIAGDEHR 158
Query: 310 NS 311
S
Sbjct: 159 RS 160
>PPP5_HUMAN (P53041) Serine/threonine protein phosphatase 5 (EC
3.1.3.16) (PP5) (Protein phosphatase T) (PP-T) (PPT)
Length = 499
Score = 63.9 bits (154), Expect = 9e-10
Identities = 36/122 (29%), Positives = 67/122 (54%), Gaps = 5/122 (4%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
++K Y +AI+ Y+ AI + +A+YY NR+ AY + Y A+ D+ R+IE+D Y K Y
Sbjct: 40 KAKDYENAIKFYSQAIELNPSNAIYYGNRSLAYLRTECYGYALGDATRAIELDKKYIKGY 99
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVK----ENIRVAEHKLMEERHRADHNQ 309
R + A G +R A+ + ++ +++ P+++ K E ++ + K E D ++
Sbjct: 100 YRRAASNMALGKFRAAL-RDYETVVKVKPHDKDAKMKYQECNKIVKQKAFERAIAGDEHK 158
Query: 310 NS 311
S
Sbjct: 159 RS 160
>STI1_SCHPO (Q9USI5) Heat shock protein sti1 homolog
Length = 591
Score = 62.8 bits (151), Expect = 2e-09
Identities = 31/104 (29%), Positives = 57/104 (54%), Gaps = 1/104 (0%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A K Y AI+ + AI + E++ + Y NR+A Y Y +A++D+ + E+ P+++K
Sbjct: 12 AFSKKDYKTAIDYFTQAIGLDERNHILYSNRSACYASEKDYADALKDATKCTELKPDWAK 71
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAE 295
+SR G A + G+ DA +++ L+ D NN + ++ E
Sbjct: 72 GWSRKGAALHGLGDL-DAARSAYEEGLKHDANNAQLLNGLKSVE 114
Score = 53.1 bits (126), Expect = 2e-06
Identities = 32/96 (33%), Positives = 51/96 (52%), Gaps = 2/96 (2%)
Query: 221 NRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQL 280
NRAAAY ++ E I+D ++IE+DPN++KAY R A + +Y ID +A ++
Sbjct: 438 NRAAAYLKVMAPAECIRDCNKAIELDPNFAKAYVRKAQALFMLKDYNKCID-ACNEASEV 496
Query: 281 DPNNESVKENIRVAEHKLME-ERHRADHNQNSRSQE 315
D + +N+R E +L + A QN +E
Sbjct: 497 DRREPNTGKNLREIESQLSKCMSAMASQRQNETEEE 532
>STI1_RAT (O35814) Stress-induced-phosphoprotein 1 (STI1)
(Hsc70/Hsp90-organizing protein) (Hop)
Length = 543
Score = 62.8 bits (151), Expect = 2e-09
Identities = 34/127 (26%), Positives = 69/127 (53%), Gaps = 1/127 (0%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A+ + DA++ Y+ AI + ++ V Y NR+AAY + Y +A +D +++++ P++ K
Sbjct: 14 ALSAGNIDDALQCYSEAIKLDPQNHVLYSNRSAAYAKKGDYQKAYEDGCKTVDLKPDWGK 73
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNS 311
YSR A + +A + +++ L+ + NN +KE ++ E +L E + N +
Sbjct: 74 GYSRKAAALEFLNRFEEA-KRTYEEGLKHEANNLQLKEGLQNMEARLAERKFMNPFNLPN 132
Query: 312 RSQEFQN 318
Q+ +N
Sbjct: 133 LYQKLEN 139
Score = 54.7 bits (130), Expect = 5e-07
Identities = 37/113 (32%), Positives = 58/113 (50%), Gaps = 4/113 (3%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
Q Y A++ Y AI + A Y NRAA YT++ + A++D I+++P + K Y
Sbjct: 372 QKGDYPQAMKHYTEAIKRNPRDAKLYSNRAACYTKLLEFQLALKDCEECIQLEPTFIKGY 431
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRAD 306
+R A A +Y A+D ++KAL LD S KE + +M + +R D
Sbjct: 432 TRKAAALEAMKDYTKAMDV-YQKALDLD---SSCKEAADGYQRCMMAQYNRHD 480
Score = 38.1 bits (87), Expect = 0.051
Identities = 31/131 (23%), Positives = 59/131 (44%), Gaps = 9/131 (6%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEID----- 246
A + K + A++ Y+ A + + Y N+AA + + Y + + ++IE+
Sbjct: 235 AYKKKDFDKALKHYDKAKELDPTNMTYITNQAAVHFEKGDYNKCRELCEKAIEVGRENRE 294
Query: 247 --PNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHR 304
+KAY+R+G +Y+ + Y+DAI F + V + + AE L E+
Sbjct: 295 DYRQIAKAYARIGNSYFKEERYKDAIH--FYNKSLAEHRTPDVLKKCQQAEKILKEQERL 352
Query: 305 ADHNQNSRSQE 315
A N + +E
Sbjct: 353 AYINPDLALEE 363
>STI1_MOUSE (Q60864) Stress-induced-phosphoprotein 1 (STI1)
(Hsc70/Hsp90-organizing protein) (Hop) (mSTI1)
Length = 543
Score = 62.8 bits (151), Expect = 2e-09
Identities = 34/127 (26%), Positives = 69/127 (53%), Gaps = 1/127 (0%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A+ + DA++ Y+ AI + ++ V Y NR+AAY + Y +A +D +++++ P++ K
Sbjct: 14 ALSAGNIDDALQCYSEAIKLDPQNHVLYSNRSAAYAKKGDYQKAYEDGCKTVDLKPDWGK 73
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNS 311
YSR A + +A + +++ L+ + NN +KE ++ E +L E + N +
Sbjct: 74 GYSRKAAALEFLNRFEEA-KRTYEEGLKHEANNLQLKEGLQNMEARLAERKFMNPFNLPN 132
Query: 312 RSQEFQN 318
Q+ +N
Sbjct: 133 LYQKLEN 139
Score = 54.7 bits (130), Expect = 5e-07
Identities = 37/113 (32%), Positives = 58/113 (50%), Gaps = 4/113 (3%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
Q Y A++ Y AI + A Y NRAA YT++ + A++D I+++P + K Y
Sbjct: 372 QKGDYPQAMKHYTEAIKRNPRDAKLYSNRAACYTKLLEFQLALKDCEECIQLEPTFIKGY 431
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRAD 306
+R A A +Y A+D ++KAL LD S KE + +M + +R D
Sbjct: 432 TRKAAALEAMKDYTKAMDV-YQKALDLD---SSCKEAADGYQRCMMAQYNRHD 480
Score = 38.1 bits (87), Expect = 0.051
Identities = 31/131 (23%), Positives = 59/131 (44%), Gaps = 9/131 (6%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEID----- 246
A + K + A++ Y+ A + + Y N+AA + + Y + + ++IE+
Sbjct: 235 AYKKKDFDKALKHYDRAKELDPTNMTYITNQAAVHFEKGDYNKCRELCEKAIEVGRENRE 294
Query: 247 --PNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHR 304
+KAY+R+G +Y+ + Y+DAI F + V + + AE L E+
Sbjct: 295 DYRQIAKAYARIGNSYFKEEKYKDAIH--FYNKSLAEHRTPDVLKKCQQAEKILKEQERL 352
Query: 305 ADHNQNSRSQE 315
A N + +E
Sbjct: 353 AYINPDLALEE 363
>STI1_CRIGR (O54981) Stress-induced-phosphoprotein 1 (STI1)
(Hsc70/Hsp90-organizing protein) (Hop)
Length = 543
Score = 62.8 bits (151), Expect = 2e-09
Identities = 34/127 (26%), Positives = 69/127 (53%), Gaps = 1/127 (0%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A+ + DA++ Y+ AI + ++ V Y NR+AAY + Y +A +D +++++ P++ K
Sbjct: 14 ALSAGNIDDALQCYSEAIKLDPQNHVLYSNRSAAYAKKGDYQKAYEDGCKTVDLKPDWGK 73
Query: 252 AYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNS 311
YSR A + +A + +++ L+ + NN +KE ++ E +L E + N +
Sbjct: 74 GYSRKAAALEFLNRFEEA-KRTYEEGLKHEANNLQLKEGLQNMEARLAERKFMNPFNLPN 132
Query: 312 RSQEFQN 318
Q+ +N
Sbjct: 133 LYQKLEN 139
Score = 56.6 bits (135), Expect = 1e-07
Identities = 38/113 (33%), Positives = 59/113 (51%), Gaps = 4/113 (3%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
Q Y A++ Y AI K A Y NRAA YT++ + A++D I+++P + K Y
Sbjct: 372 QKGDYPQAMKHYTEAIKRNPKDAKLYSNRAACYTKLLEFQLALKDCEECIQLEPTFIKGY 431
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRAD 306
+R A A +Y A+D ++KAL+LD S KE + +M + +R D
Sbjct: 432 TRKAAALEAMKDYTKAMDV-YQKALELD---SSCKEAADGYQRCMMAQYNRHD 480
Score = 37.4 bits (85), Expect = 0.086
Identities = 31/131 (23%), Positives = 59/131 (44%), Gaps = 9/131 (6%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPN--- 248
A + K + A++ Y+ A + + Y N+AA + + Y + + ++IE+
Sbjct: 235 AYKKKDFDMALKHYDRAKELDPTNMTYITNQAAVHFEKGDYNKCRELCEKAIEVGRENRE 294
Query: 249 ----YSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHR 304
+KAY+R+G +Y+ + Y+DAI F + V + + AE L E+
Sbjct: 295 DYRQIAKAYARIGNSYFKEERYKDAIH--FYNKSLAEHRTPDVLKKCQQAEKILKEQERL 352
Query: 305 ADHNQNSRSQE 315
A N + +E
Sbjct: 353 AYINPDLALEE 363
>STI1_HUMAN (P31948) Stress-induced-phosphoprotein 1 (STI1)
(Hsc70/Hsp90-organizing protein) (Hop)
(Transformation-sensitive protein IEF SSP 3521)
Length = 543
Score = 61.2 bits (147), Expect = 6e-09
Identities = 32/119 (26%), Positives = 65/119 (53%), Gaps = 1/119 (0%)
Query: 200 DAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLA 259
DA++ Y+ AI + + V Y NR+AAY + Y +A +D +++++ P++ K YSR A
Sbjct: 22 DALQCYSEAIKLDPHNHVLYSNRSAAYAKKGDYQKAYEDGCKTVDLKPDWGKGYSRKAAA 81
Query: 260 YYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNSRSQEFQN 318
+ +A + +++ L+ + NN +KE ++ E +L E + N + Q+ ++
Sbjct: 82 LEFLNRFEEA-KRTYEEGLKHEANNPQLKEGLQNMEARLAERKFMNPFNMPNLYQKLES 139
Score = 55.8 bits (133), Expect = 2e-07
Identities = 38/113 (33%), Positives = 58/113 (50%), Gaps = 4/113 (3%)
Query: 194 QSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAY 253
Q Y A++ Y AI K A Y NRAA YT++ + A++D I+++P + K Y
Sbjct: 372 QKGDYPQAMKHYTEAIKRNPKDAKLYSNRAACYTKLLEFQLALKDCEECIQLEPTFIKGY 431
Query: 254 SRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRAD 306
+R A A +Y A+D ++KAL LD S KE + +M + +R D
Sbjct: 432 TRKAAALEAMKDYTKAMDV-YQKALDLD---SSCKEAADGYQRCMMAQYNRHD 480
Score = 40.0 bits (92), Expect = 0.013
Identities = 32/131 (24%), Positives = 59/131 (44%), Gaps = 9/131 (6%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEID----- 246
A + K + A++ Y+ A + + Y N+AA Y + Y + + ++IE+
Sbjct: 235 AYKKKDFDTALKHYDKAKELDPTNMTYITNQAAVYFEKGDYNKCRELCEKAIEVGRENRE 294
Query: 247 --PNYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHR 304
+KAY+R+G +Y+ + Y+DAI F + V + + AE L E+
Sbjct: 295 DYRQIAKAYARIGNSYFKEEKYKDAIH--FYNKSLAEHRTPDVLKKCQQAEKILKEQERL 352
Query: 305 ADHNQNSRSQE 315
A N + +E
Sbjct: 353 AYINPDLALEE 363
>DJC7_HUMAN (Q99615) DnaJ homolog subfamily C member 7
(Tetratricopeptide repeat protein 2) (TPR repeat protein
2)
Length = 494
Score = 60.1 bits (144), Expect = 1e-08
Identities = 33/107 (30%), Positives = 58/107 (53%), Gaps = 1/107 (0%)
Query: 196 KQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSR 255
K Y +A Y AI + K+A YY NRAA + R+ EA+ D+ +S+ +D ++ + + R
Sbjct: 42 KDYNEAYNYYTKAIDMCPKNASYYGNRAATLMMLGRFREALGDAQQSVRLDDSFVRGHLR 101
Query: 256 LGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER 302
G + + GN A + F++AL+LD N ++ + A + E+
Sbjct: 102 EGKCHLSLGNAMAAC-RSFQRALELDHKNAQAQQEFKNANAVMEYEK 147
Score = 52.4 bits (124), Expect = 3e-06
Identities = 37/146 (25%), Positives = 71/146 (48%), Gaps = 13/146 (8%)
Query: 192 AMQSKQYFDAIELYNCAIAI----YEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDP 247
A + Y A ELY A+ I + +A YCNR +++ + +AI+D ++++D
Sbjct: 266 AFKEGNYKLAYELYTEALGIDPNNIKTNAKLYCNRGTVNSKLRKLDDAIEDCTNAVKLDD 325
Query: 248 NYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADH 307
Y KAY R Y Y +A+ + ++K Q + E K+ ++ A+ +L + + R D+
Sbjct: 326 TYIKAYLRRAQCYMDTEQYEEAV-RDYEKVYQTEKTKEH-KQLLKNAQLELKKSK-RKDY 382
Query: 308 ------NQNSRSQEFQNHYARGSRSH 327
++N+ E + Y + + H
Sbjct: 383 YKILGVDKNASEDEIKKAYRKRALMH 408
>CNS1_YEAST (P33313) Cyclophilin seven suppressor 1 (STI1
stress-inducible protein homolog)
Length = 385
Score = 59.3 bits (142), Expect = 2e-08
Identities = 57/238 (23%), Positives = 100/238 (41%), Gaps = 26/238 (10%)
Query: 91 VSAQNAADAKTHPDESKPMDEDWTQGPHTSAVSKDELCGQFFAVLEKKHYFRTNID---- 146
+S+ NA T P + P D P S KD+ + + + +F T +D
Sbjct: 1 MSSVNANGGYTKPQKYVPGPGDPELPPQLSEF-KDKTSDEILKEMNRMPFFMTKLDETDG 59
Query: 147 GGDDIVQLEKASRLFDDG--FTLYRRWKDLGVSSLI*RIWLNH*KH*AMQSKQYFDAIEL 204
G + V+LE L +G + +K G ++K++ DA EL
Sbjct: 60 AGGENVELEALKALAYEGEPHEIAENFKKQGNE--------------LYKAKRFKDAREL 105
Query: 205 YNCAIAIY--EKSA--VYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSRLGLAY 260
Y+ +A+ +KS Y NRAA ++ Y I+D +++ I+P K Y R A+
Sbjct: 106 YSKGLAVECEDKSINESLYANRAACELELKNYRRCIEDCSKALTINPKNVKCYYRTSKAF 165
Query: 261 YAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADHNQNSRSQEFQN 318
+ +A ++DP N+S+ + V + K E + + + Q +QE +N
Sbjct: 166 FQLNKLEEAKSAATFANQRIDPENKSILNMLSVIDRKEQELKAK-EEKQQREAQEREN 222
>DJC7_MOUSE (Q9QYI3) DnaJ homolog subfamily C member 7
(Tetratricopeptide repeat protein 2) (TPR repeat protein
2) (MDj11)
Length = 494
Score = 58.2 bits (139), Expect = 5e-08
Identities = 32/107 (29%), Positives = 57/107 (52%), Gaps = 1/107 (0%)
Query: 196 KQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKAYSR 255
K Y +A Y AI + +A YY NRAA + R+ EA+ D+ +S+ +D ++ + + R
Sbjct: 42 KDYNEAYNYYTKAIDMCPNNASYYGNRAATLMMLGRFREALGDAQQSVRLDDSFVRGHLR 101
Query: 256 LGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEER 302
G + + GN A + F++AL+LD N ++ + A + E+
Sbjct: 102 EGKCHLSLGNAMAAC-RSFQRALELDHKNAQAQQEFKNANAVMEYEK 147
Score = 50.4 bits (119), Expect = 1e-05
Identities = 36/146 (24%), Positives = 71/146 (47%), Gaps = 13/146 (8%)
Query: 192 AMQSKQYFDAIELYNCAIAI----YEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDP 247
A + Y A ELY A+ I + +A YCNR +++ + +AI+D ++++D
Sbjct: 266 AFKEGNYKLAYELYTEALGIDPNNIKTNAKLYCNRGTVNSKLRQLEDAIEDCTNAVKLDD 325
Query: 248 NYSKAYSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHRADH 307
Y KAY R Y + +A+ + ++K Q + E K+ ++ A+ +L + + R D+
Sbjct: 326 TYIKAYLRRAQCYMDTEQFEEAV-RDYEKVYQTEKTKEH-KQLLKNAQLELKKSK-RKDY 382
Query: 308 ------NQNSRSQEFQNHYARGSRSH 327
++N+ E + Y + + H
Sbjct: 383 YKILGVDKNASEDEIKKAYRKRALMH 408
>ST13_HUMAN (P50502) Hsc70-interacting protein (Hip) (Putative tumor
suppressor ST13) (Progesterone receptor-associated p48
protein)
Length = 369
Score = 56.6 bits (135), Expect = 1e-07
Identities = 49/200 (24%), Positives = 90/200 (44%), Gaps = 16/200 (8%)
Query: 192 AMQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSK 251
A+ + AI+L+ AI + + A+ Y RA+ + ++ + AI+D R+IEI+P+ ++
Sbjct: 124 ALNDGELQKAIDLFTDAIKLNPRLAILYAKRASVFVKLQKPNAAIRDCDRAIEINPDSAQ 183
Query: 252 AYSRLGLAYYAQGNYRDAI-DKGFKKALQLDPNNESVKENI-----RVAEHKLMEERHRA 305
Y G A+ G++ +A D L D + ++ + + ++AEH+ ER R
Sbjct: 184 PYKWRGKAHRLLGHWEEAAHDLALACKLDYDEDASAMLKEVQPRAQKIAEHRRKYERKRE 243
Query: 306 DH---NQNSRSQEFQNHYARGSRSHAA----PASFGSMPFN-PSNLASMFMAAANG-GQG 356
+ + R ++ + + R R A A +GS P P + F G G G
Sbjct: 244 EREIKERIERVKKAREEHERAQREEEARRQSGAQYGSFPGGFPGGMPGNFPGGMPGMGGG 303
Query: 357 SHSQEGQEDANSSGANEPEI 376
G N ++PE+
Sbjct: 304 MPGMAGMPGLNEI-LSDPEV 322
>PPT1_YEAST (P53043) Serine/threonine protein phosphatase T (EC
3.1.3.16) (PPT)
Length = 513
Score = 56.6 bits (135), Expect = 1e-07
Identities = 32/112 (28%), Positives = 58/112 (51%), Gaps = 1/112 (0%)
Query: 193 MQSKQYFDAIELYNCAIAIYEKSAVYYCNRAAAYTQINRYTEAIQDSLRSIEIDPNYSKA 252
++ K + AIE Y AI + ++Y+ NRA A+ +++ + A+ D +I++DP KA
Sbjct: 23 VKEKHFLKAIEKYTEAIDLDSTQSIYFSNRAFAHFKVDNFQSALNDCDEAIKLDPKNIKA 82
Query: 253 YSRLGLAYYAQGNYRDAIDKGFKKALQLDPNNESVKENIRVAEHKLMEERHR 304
Y R L+ A ++ A K L+ PN+ + + + + + EER R
Sbjct: 83 YHRRALSCMALLEFKKA-RKDLNVLLKAKPNDPAATKALLTCDRFIREERFR 133
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.317 0.132 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 56,844,532
Number of Sequences: 164201
Number of extensions: 2406716
Number of successful extensions: 7195
Number of sequences better than 10.0: 157
Number of HSP's better than 10.0 without gapping: 83
Number of HSP's successfully gapped in prelim test: 76
Number of HSP's that attempted gapping in prelim test: 6856
Number of HSP's gapped (non-prelim): 334
length of query: 481
length of database: 59,974,054
effective HSP length: 114
effective length of query: 367
effective length of database: 41,255,140
effective search space: 15140636380
effective search space used: 15140636380
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 68 (30.8 bits)
Medicago: description of AC139354.8


