

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.6 - phase: 0
(342 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
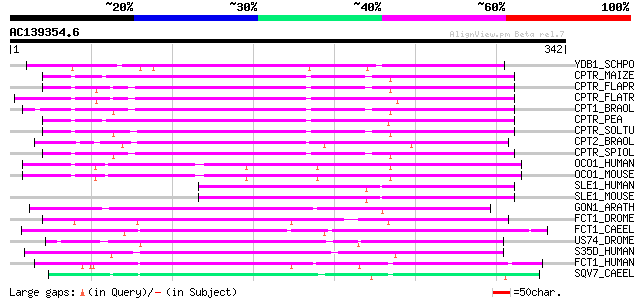
Score E
Sequences producing significant alignments: (bits) Value
YDB1_SCHPO (Q10354) Hypothetical protein C22E12.01 in chromosome I 105 1e-22
CPTR_MAIZE (P49133) Triose phosphate/phosphate translocator, chl... 103 8e-22
CPTR_FLAPR (P49131) Triose phosphate/phosphate translocator, chl... 103 8e-22
CPTR_FLATR (P49132) Triose phosphate/phosphate translocator, chl... 102 1e-21
CPT1_BRAOL (P52177) Triose phosphate/phosphate translocator, chl... 100 9e-21
CPTR_PEA (P21727) Triose phosphate/phosphate translocator, chlor... 99 1e-20
CPTR_SOLTU (P29463) Triose phosphate/phosphate translocator, chl... 99 2e-20
CPT2_BRAOL (P52178) Triose phosphate/phosphate translocator, non... 98 3e-20
CPTR_SPIOL (P11869) Triose phosphate/phosphate translocator, chl... 92 2e-18
OCO1_HUMAN (Q9NQQ7) Solute carrier family 35 member C2 (Ovarian ... 92 2e-18
OCO1_MOUSE (Q8VCX2) Solute carrier family 35 member C2 (Ovarian ... 89 2e-17
SLE1_HUMAN (Q96K37) Solute carrier family 35 member E1 87 8e-17
SLE1_MOUSE (Q8CD26) Solute carrier family 35 member E1 83 9e-16
GON1_ARATH (Q941R4) GDP-mannose transporter 83 9e-16
FCT1_DROME (Q9VHT4) Probable GDP-fucose transporter 82 3e-15
FCT1_CAEEL (Q968A5) GDP-fucose transporter 72 2e-12
US74_DROME (Q95YI5) UDP-sugar transporter UST74c (Fringe connect... 65 2e-10
S35D_HUMAN (Q9NTN3) UDP-glucuronic acid/UDP-N-acetylgalactosamin... 61 5e-09
FCT1_HUMAN (Q96A29) GDP-fucose transporter 1 (Solute carrier fam... 58 3e-08
SQV7_CAEEL (Q18779) UDP-sugar transporter sqv-7 (Squashed vulva ... 55 3e-07
>YDB1_SCHPO (Q10354) Hypothetical protein C22E12.01 in chromosome I
Length = 374
Score = 105 bits (263), Expect = 1e-22
Identities = 85/309 (27%), Positives = 140/309 (44%), Gaps = 20/309 (6%)
Query: 11 VVRSLLAILQWWAFNVTVIIMNKWIFQ--KLDFKFPLSVSCIHFICSAIGAYVVIKVLKL 68
VV +L +L W+ F++ + +MNKWIF K+DF+FPL +S + A + I
Sbjct: 50 VVIIVLIVLAWYFFSLLLSMMNKWIFSESKMDFQFPLFLSSCQMLVQMGFAKLTILAF-- 107
Query: 69 KPLISVDPQDR--WRRIFPMS----FVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVV 122
P + +D W F + V ++I L N SL I +SF +S
Sbjct: 108 -PRYQPNKKDNFSWLEYFYRAGICALVTGLDIGLSNASLETITLSFYTMCRSSILIFVFF 166
Query: 123 LQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEAL 182
+ + FDW + + I G++L TE F + GF + + + + L + L
Sbjct: 167 FSVIFRIEMFDWILLCITLVISAGVVLMVATETQFVLSGFLLVMASSVLSGLRWALTQKL 226
Query: 183 L--HGYKFDSINTVYHMAPFATLIMVFPALLLEGN-GILE---WFSIHPYPWAAMIIIFS 236
L H + + +++ + P L ++ L+ EG +E W P+ ++I
Sbjct: 227 LLDHPWTSNPFTSLFALTPLMFLFLLVAGLIFEGPVRFIESPAWKEFGPF---MSVVILV 283
Query: 237 SGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLV 296
G LAF + S F +I T+ VT +V G LK + ++ S L + + + +N VG ITL
Sbjct: 284 PGTLAFFMVASEFGLIQKTSIVTLSVCGILKEIITIIASTLFYHDILLPINIVGLVITLC 343
Query: 297 GCTFYGYVR 305
G Y Y R
Sbjct: 344 GIGVYNYYR 352
>CPTR_MAIZE (P49133) Triose phosphate/phosphate translocator,
chloroplast precursor (CTPT)
Length = 409
Score = 103 bits (256), Expect = 8e-22
Identities = 75/299 (25%), Positives = 134/299 (44%), Gaps = 18/299 (6%)
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRW 80
W+ NV I+NK I+ F +P VS IH + + Y +I P +
Sbjct: 115 WYFLNVIFNILNKKIYNY--FPYPYFVSLIHLVVGVV--YCLISWSVGLPKRAPINGTLL 170
Query: 81 RRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASL 140
+ +FP++ I + NVS + VSF TIK+ P + + + + +W SL
Sbjct: 171 KLLFPVALCHGIGHITSNVSFAAVAVSFAHTIKALEPFFSAAATQFILGQQVPFSLWLSL 230
Query: 141 VPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPF 200
P+V G+ + S+TELSFN GF A+ ++ + ++I ++ + DS N +++
Sbjct: 231 APVVIGVSMASLTELSFNWTGFINAMISNISFTYRSIYSKKAM--TDMDSTNVYAYISII 288
Query: 201 ATLIMVFPALLLEGNGILEWFSIHPYPWAAMII--------IFSSGVLAFCLNFSIFYVI 252
A ++ + PAL+ EG +++ H + A + +F G+ N +
Sbjct: 289 ALIVCIPPALIFEGPKLMQ----HGFSDAIAKVGLTKFVSDLFLVGLFYHLYNQIATNTL 344
Query: 253 HSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRNMISQQ 311
+T V LK + S ++F N IS +G +I + G Y Y++ I ++
Sbjct: 345 ERVAPLTHAVGNVLKRVFVIGFSIIVFGNKISTQTGIGTSIAIAGVAMYSYIKAKIEEE 403
>CPTR_FLAPR (P49131) Triose phosphate/phosphate translocator,
chloroplast precursor (CTPT)
Length = 408
Score = 103 bits (256), Expect = 8e-22
Identities = 79/302 (26%), Positives = 137/302 (45%), Gaps = 24/302 (7%)
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHF---ICSAIGAYVVIKVLKLKPLISVDPQ 77
W+ NV I+NK I+ F +P VS IH + +G + V + K P+ S
Sbjct: 113 WYFLNVIFNILNKKIYNY--FPYPYFVSAIHLAVGVVYCLGGWAV-GLPKRAPMDS---- 165
Query: 78 DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIW 137
+ + + P++F + V NVS + VSF TIKS P + + +W
Sbjct: 166 NLLKLLIPVAFCHALGHVTSNVSFAAVAVSFTHTIKSLEPFFNAAASQFILGQSIPITLW 225
Query: 138 ASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHM 197
SL P+V G+ + S+TELSFN GF +A+ ++ + ++I ++ + DS N ++
Sbjct: 226 LSLAPVVIGVSMASLTELSFNWLGFISAMISNISFTYRSIYSKKAM--TDMDSTNLYAYI 283
Query: 198 APFATLIMVFPALLLEGNGILEWFSIHPYPWAAMII--------IFSSGVLAFCLNFSIF 249
+ + L + PA++LEG +L+ H + A + +F G+ N
Sbjct: 284 SIISLLFCIPPAIILEGPQLLK----HGFSDAIAKVGMTKFISDLFWVGMFYHLYNQLAI 339
Query: 250 YVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRNMIS 309
+ +T V LK + S ++F N IS A+G +I + G Y ++ I
Sbjct: 340 NTLERVAPLTHAVGNVLKRVFVIGFSIIVFGNKISTQTAIGTSIAIAGVAVYSLIKAKIE 399
Query: 310 QQ 311
++
Sbjct: 400 EE 401
>CPTR_FLATR (P49132) Triose phosphate/phosphate translocator,
chloroplast precursor (CTPT)
Length = 407
Score = 102 bits (254), Expect = 1e-21
Identities = 81/315 (25%), Positives = 139/315 (43%), Gaps = 16/315 (5%)
Query: 4 GLVCQWSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHF---ICSAIGAY 60
G + ++ + + W+ NV I+NK I+ F +P VS IH + +G++
Sbjct: 95 GFLAKYPFLVTGFFFFMWYFLNVIFNILNKKIYNY--FPYPYFVSVIHLAVGVVYCLGSW 152
Query: 61 VVIKVLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATT 120
V + K P+ S + + + P+ F + V NVS + VSF TIK+ P
Sbjct: 153 TV-GLPKRAPVDS----NILKLLIPVGFCHALGHVTSNVSFAAVAVSFTHTIKALEPFFN 207
Query: 121 VVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAE 180
V + +W SL P+V G+ + S+TELSFN GF +A+ ++ + ++I ++
Sbjct: 208 AAASQFVLGQSIPISLWLSLAPVVIGVSMASLTELSFNWLGFISAMISNISFTYRSIYSK 267
Query: 181 ALLHGYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSS--- 237
+ DS N +++ A L + PA+L EG +L+ MI S
Sbjct: 268 KAM--TDMDSTNLYAYISIIALLFCIPPAVLFEGPQLLKHGFNDAIAKVGMIKFISDLFW 325
Query: 238 -GVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLV 296
G+ N + +T V LK + S ++F N IS A+G +I +
Sbjct: 326 VGMFYHLYNQIATNTLERVAPLTHAVGNVLKRVFVIGFSIIVFGNKISTQTAIGTSIAIA 385
Query: 297 GCTFYGYVRNMISQQ 311
G Y ++ I ++
Sbjct: 386 GVAIYSLIKARIEEE 400
>CPT1_BRAOL (P52177) Triose phosphate/phosphate translocator,
chloroplast precursor (CTPT)
Length = 407
Score = 99.8 bits (247), Expect = 9e-21
Identities = 82/313 (26%), Positives = 137/313 (43%), Gaps = 24/313 (7%)
Query: 9 WSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVV--IKVL 66
W V LL L W+ NV I+NK I+ F +P VS IH + V + +
Sbjct: 102 WLVTGILL--LMWYFLNVIFNILNKKIYNY--FPYPYFVSVIHLFVGVVYCLVSWSVGLP 157
Query: 67 KLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWL 126
K P+ S D + + P++ I V NVS + VSF TIK+ P
Sbjct: 158 KRAPVNS----DILKVLIPVAVCHAIGHVTSNVSFAAVAVSFTHTIKALEPFFNASASQF 213
Query: 127 VWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGY 186
+ + +W SL P+V G+ + S+TELSFN GF +A+ ++ + ++I ++ +
Sbjct: 214 LLGQPIPITLWLSLAPVVLGVAMASLTELSFNWLGFISAMISNISFTYRSIFSKKAM--T 271
Query: 187 KFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMII--------IFSSG 238
DS N +++ A + + PA+++EG +L+ H + A + +F G
Sbjct: 272 DMDSTNVYAYISIIALFVCLPPAIIVEGPQLLK----HGFNDAIAKVGMTKFISDLFWVG 327
Query: 239 VLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGC 298
+ N + +T V LK + S +IF N IS +G I + G
Sbjct: 328 MFYHLYNQLATNTLERVAPLTHAVGNVLKRVFVIGFSIVIFGNKISTQTGIGTGIAIAGV 387
Query: 299 TFYGYVRNMISQQ 311
Y ++ I ++
Sbjct: 388 ALYSVIKAKIEEE 400
>CPTR_PEA (P21727) Triose phosphate/phosphate translocator,
chloroplast precursor (CTPT) (p36) (E30)
Length = 402
Score = 99.4 bits (246), Expect = 1e-20
Identities = 71/295 (24%), Positives = 130/295 (44%), Gaps = 10/295 (3%)
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRW 80
W+ NV I+NK I+ F +P VS IH + Y ++ P + +
Sbjct: 107 WYFLNVIFNILNKKIYNY--FPYPYFVSVIHLAVGVV--YCLVSWTVGLPKRAPIDGNLL 162
Query: 81 RRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASL 140
+ + P++ + V NVS + VSF T+K+ P + + +W SL
Sbjct: 163 KLLIPVAVCHALGHVTSNVSFAAVAVSFTHTVKALEPFFNAAASQFILGQSIPITLWLSL 222
Query: 141 VPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPF 200
P+V G+ + S+TELSFN GF +A+ ++ + ++I ++ + DS N +++
Sbjct: 223 APVVIGVSMASLTELSFNWLGFISAMISNISFTYRSIYSKKAM--TDMDSTNIYAYISII 280
Query: 201 ATLIMVFPALLLEGNGILEWFSIHPYPWAAMI----IIFSSGVLAFCLNFSIFYVIHSTT 256
A ++ + PAL++EG +L+ ++ +F G+ N +
Sbjct: 281 ALIVCIPPALIIEGPTLLKTGFNDAIAKVGLVKFVSDLFWVGMFYHLYNQVATNTLERVA 340
Query: 257 AVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRNMISQQ 311
+T V LK + S +IF N IS +G I + G Y +++ I ++
Sbjct: 341 PLTHAVGNVLKRVFVIGFSIIIFGNKISTQTGIGTGIAIAGVALYSFIKAQIEEE 395
>CPTR_SOLTU (P29463) Triose phosphate/phosphate translocator,
chloroplast precursor (CTPT) (E29)
Length = 414
Score = 98.6 bits (244), Expect = 2e-20
Identities = 74/301 (24%), Positives = 133/301 (43%), Gaps = 22/301 (7%)
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVV--IKVLKLKPLISVDPQD 78
W+ NV I+NK I+ F +P VS IH + V + + K P+ S
Sbjct: 119 WYFLNVIFNILNKKIYNY--FPYPYFVSVIHLAVGVVYCLVSWGVGLPKRAPIDST---- 172
Query: 79 RWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWA 138
+ + + P++F + V NVS + VSF T+K+ P + + +W
Sbjct: 173 QLKLLTPVAFCHALGHVTSNVSFAAVRVSFTHTVKALEPFFNAAASQFILGQQIPLALWL 232
Query: 139 SLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMA 198
SL P+V G+ + S+TELSFN GF +A+ ++ + ++I ++ + DS N +++
Sbjct: 233 SLAPVVLGVSMASLTELSFNWLGFTSAMISNISFTYRSIYSKKAM--TDMDSTNVYAYIS 290
Query: 199 PFATLIMVFPALLLEGNGILEWFSIHPYPWAAMII--------IFSSGVLAFCLNFSIFY 250
A + + PA+ +EG +L+ H + A + +F G+ N
Sbjct: 291 IIALIFCLPPAIFIEGPQLLQ----HGFNDAIAKVGLTKFVTDLFWVGMFYHLYNQVATN 346
Query: 251 VIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRNMISQ 310
+ +T V LK + S +IF N IS +G I + G Y +++ + +
Sbjct: 347 TLERVAPLTHAVGNVLKRVFVIGFSIVIFGNKISTQTGIGTCIAIAGVAIYSFIKAKMEE 406
Query: 311 Q 311
+
Sbjct: 407 E 407
>CPT2_BRAOL (P52178) Triose phosphate/phosphate translocator,
non-green plastid, chloroplast precursor (CTPT)
Length = 402
Score = 97.8 bits (242), Expect = 3e-20
Identities = 79/301 (26%), Positives = 143/301 (47%), Gaps = 18/301 (5%)
Query: 16 LAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKL----KPL 71
L W+ FN+ I NK + + L P++V+ + F A+G+ ++ + L +P
Sbjct: 104 LLFAMWYLFNIYFNIYNKQVLKALHA--PMTVTLVQF---AVGSVLITFMWALNLYKRPK 158
Query: 72 ISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKY 131
IS + I P++ V + + N+SL + VSF TIK+ P +VVL + +
Sbjct: 159 ISAA---QLAAILPLAVVHTLGNLFTNMSLGKVSVSFTHTIKAMEPFFSVVLSAMFLGEV 215
Query: 132 FDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSI 191
+ S++PIVGG+ L S+TE+SFN GF +A+ L ++ +L++ ++ K DS+
Sbjct: 216 PTPWVIGSIIPIVGGVALASVTEVSFNWAGFLSAMASNLTNQSRNVLSKKVM-VKKDDSL 274
Query: 192 N--TVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFSSGVLAFCLNF--- 246
+ T++ + +L ++ P I I S + A C +
Sbjct: 275 DNITLFSIITLMSLFLMAPVTFFSEGIKFTPSYIQSAGVNVQQIYTKSLIAALCFHAYQQ 334
Query: 247 SIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRN 306
+ ++ + VT +V +K V ++ S + F+ P+S +NA G I L G Y V+
Sbjct: 335 VSYMILARVSPVTHSVGNCVKRVVVIVSSVIFFKTPVSPVNAFGTGIALAGVFLYSRVKR 394
Query: 307 M 307
+
Sbjct: 395 I 395
>CPTR_SPIOL (P11869) Triose phosphate/phosphate translocator,
chloroplast precursor (CTPT) (P36) (E29)
Length = 404
Score = 92.0 bits (227), Expect = 2e-18
Identities = 73/301 (24%), Positives = 131/301 (43%), Gaps = 22/301 (7%)
Query: 21 WWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVV--IKVLKLKPLISVDPQD 78
W+ NV I+NK I+ F +P VS IH + + + K P+ S
Sbjct: 109 WYFLNVIFNILNKKIYNY--FPYPYFVSVIHLFVGVVYCLASWSVGLPKRAPMDS----K 162
Query: 79 RWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWA 138
+ + P++ I V NVS + VSF TIK+ P V + +W
Sbjct: 163 LLKLLIPVAVCHAIGHVTSNVSFAAVAVSFTHTIKALEPFFNAAASQFVLGQSIPITLWL 222
Query: 139 SLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMA 198
SL P+V G+ + S+TELSFN GF +A+ ++ + +++ ++ + DS N +++
Sbjct: 223 SLAPVVIGVSMASLTELSFNWLGFISAMISNVSFTYRSLYSKKAM--TDMDSTNIYAYIS 280
Query: 199 PFATLIMVFPALLLEGNGILEWFSIHPYPWAAMII--------IFSSGVLAFCLNFSIFY 250
A + + PA+++EG +++ H + A + +F G+ N
Sbjct: 281 IIALFVCLPPAIIVEGPQLMK----HGFNDAIAKVGLTKFISDLFWVGMFYHLYNQLATN 336
Query: 251 VIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYVRNMISQ 310
+ +T V LK + S + F N IS A+G +I + G Y ++ + +
Sbjct: 337 TLERVAPLTHAVGNVLKRVFVIGFSIIAFGNKISTQTAIGTSIAIAGVALYSLIKAKMEE 396
Query: 311 Q 311
+
Sbjct: 397 E 397
>OCO1_HUMAN (Q9NQQ7) Solute carrier family 35 member C2 (Ovarian
cancer overexpressed gene 1 protein) (CGI-15)
Length = 365
Score = 91.7 bits (226), Expect = 2e-18
Identities = 78/324 (24%), Positives = 152/324 (46%), Gaps = 25/324 (7%)
Query: 9 WSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIH----FICSAIGAYVVIK 64
W V +L +L ++ F++ + NKW+ + F FPL ++ +H F+ SA+ +++
Sbjct: 12 WKAVLTLGLVLLYYCFSIGITFYNKWLTKS--FHFPLFMTMLHLAVIFLFSALSR-ALVQ 68
Query: 65 VLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQ 124
+ + + D RR+ P + +++ L N S Y+ VS KS + VL
Sbjct: 69 CSSHRARVVLSWADYLRRVAPTALATALDVGLSNWSFLYVTVSLYTMTKS-----SAVLF 123
Query: 125 WLVWRKYFDWR-IWASLVPIV----GGILLTSITELSFNMFGFCAALFGCLATSTKTILA 179
L++ F + A+LV +V GG+ + + FN+ GF L + L
Sbjct: 124 ILIFSLIFKLEELRAALVLVVLLIAGGLFMFTYKSTQFNVEGFALVLGASFIGGIRWTLT 183
Query: 180 EALLHGYKF---DSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMII--- 233
+ LL + + I+T++H+ P L + + EG + I + +++
Sbjct: 184 QMLLQKAELGLQNPIDTMFHLQPLMFLGLFPLFAVFEGLHLSTSEKIFRFQDTGLLLRVL 243
Query: 234 --IFSSGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGC 291
+F G+LAF L FS F ++ T+++T ++AG K +L++ + + IS +N +G
Sbjct: 244 GSLFLGGILAFGLGFSEFLLVSRTSSLTLSIAGIFKEVCTLLLAAHLLGDQISLLNWLGF 303
Query: 292 AITLVGCTFYGYVRNMISQQPAVP 315
A+ L G + + ++ + S+ P
Sbjct: 304 ALCLSGISLHVALKALHSRGDGGP 327
>OCO1_MOUSE (Q8VCX2) Solute carrier family 35 member C2 (Ovarian
cancer overexpressed gene 1 protein)
Length = 364
Score = 89.0 bits (219), Expect = 2e-17
Identities = 78/324 (24%), Positives = 149/324 (45%), Gaps = 25/324 (7%)
Query: 9 WSVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIH----FICSAIGAYVVIK 64
W +L +L ++ F++ + NKW+ + F FPL ++ +H F+ SA+ +++
Sbjct: 12 WKAALTLGLVLLYYCFSIGITFYNKWLTKS--FHFPLFMTMLHLAVIFLFSALSR-ALVQ 68
Query: 65 VLKLKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQ 124
K + + D RR+ P + +++ L N S YI VS KS + VL
Sbjct: 69 CSSHKARVVLSWTDYLRRVAPTALATALDVGLSNWSFLYITVSLYTMTKS-----SAVLF 123
Query: 125 WLVWRKYFDWR-IWASLVPIV----GGILLTSITELSFNMFGFCAALFGCLATSTKTILA 179
L++ F + A+LV +V GG+ + + FN+ GF L + L
Sbjct: 124 ILIFSLIFKLEELRAALVLVVLLIAGGLFMFTYKSTQFNVEGFALVLGASFIGGIRWTLT 183
Query: 180 EALLHGYKF---DSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMII--- 233
+ LL + I+T++H+ P L + + EG + I + +++
Sbjct: 184 QILLQKADLGLQNPIDTMFHLQPLMFLGLFPLFAIFEGLHLSTSEKIFRFQDTGLLLWVL 243
Query: 234 --IFSSGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGC 291
+ G+LAF L FS F ++ T+++T ++AG K +L++ + + IS +N +G
Sbjct: 244 GSLLLGGILAFGLGFSEFLLVSRTSSLTLSIAGIFKEVCTLLLAAHLLGDQISLLNWLGF 303
Query: 292 AITLVGCTFYGYVRNMISQQPAVP 315
A+ L G + + ++ + S+ P
Sbjct: 304 ALCLSGISLHVALKALHSRGDGGP 327
>SLE1_HUMAN (Q96K37) Solute carrier family 35 member E1
Length = 266
Score = 86.7 bits (213), Expect = 8e-17
Identities = 47/198 (23%), Positives = 104/198 (51%), Gaps = 4/198 (2%)
Query: 117 PATTVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKT 176
P V+L ++ ++ +++ SL+PI+ G+LL ++TELSF+M+G +AL L S +
Sbjct: 2 PIWVVLLSRIIMKEKQSTKVYLSLIPIISGVLLATVTELSFDMWGLVSALAATLCFSLQN 61
Query: 177 ILAEALLHGYKFDSINTVYHMAPFATLIMVFPALLLEGNGIL---EWFSIHPYPWAAMII 233
I ++ +L + + + + A M+ +L++ + L + ++ +PW +++
Sbjct: 62 IFSKKVLRDSRIHHLRLLNILGCHAVFFMIPTWVLVDLSAFLVSSDLTYVYQWPW-TLLL 120
Query: 234 IFSSGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAI 293
+ SG F N F +++ + ++++VA K + + +S ++ RNP++ N +G
Sbjct: 121 LAVSGFCNFAQNVIAFSILNLVSPLSYSVANATKRIMVITVSLIMLRNPVTSTNVLGMMT 180
Query: 294 TLVGCTFYGYVRNMISQQ 311
++G Y + +QQ
Sbjct: 181 AILGVFLYNKTKYDANQQ 198
>SLE1_MOUSE (Q8CD26) Solute carrier family 35 member E1
Length = 265
Score = 83.2 bits (204), Expect = 9e-16
Identities = 47/198 (23%), Positives = 103/198 (51%), Gaps = 4/198 (2%)
Query: 117 PATTVVLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKT 176
P V+L ++ ++ +++ SLVPI+ G+LL ++TELSF+++G +AL L S +
Sbjct: 2 PIWVVLLSRIIMKEKQSTKVYLSLVPIISGVLLATVTELSFDVWGLVSALAATLCFSLQN 61
Query: 177 ILAEALLHGYKFDSINTVYHMAPFATLIMVFPALLLEGNGIL---EWFSIHPYPWAAMII 233
I ++ +L + + + + A M+ +L++ + L + + +PW +++
Sbjct: 62 IFSKKVLRDSRIHHLRLLNILGCHAVFFMIPTWVLVDLSTFLVSSDLAYVSQWPW-TLLL 120
Query: 234 IFSSGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAI 293
+ SG F N F +++ + ++++VA K + + +S ++ RNP++ N +G
Sbjct: 121 LAVSGFCNFAQNVIAFSILNLISPLSYSVANATKRIMVITVSLIMLRNPVTSTNVLGMMT 180
Query: 294 TLVGCTFYGYVRNMISQQ 311
++G Y + +QQ
Sbjct: 181 AILGVFLYNKTKYDANQQ 198
>GON1_ARATH (Q941R4) GDP-mannose transporter
Length = 333
Score = 83.2 bits (204), Expect = 9e-16
Identities = 65/294 (22%), Positives = 139/294 (47%), Gaps = 16/294 (5%)
Query: 13 RSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLI 72
R+LL+ L + + ++I++NK++ +F + + S I ++ L L LI
Sbjct: 32 RALLSGLAYCISSCSMILVNKFVLSSYNFNAGIFLMLYQNFVSVI----IVVGLSLMGLI 87
Query: 73 SVDPQD-RWRRI-FPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRK 130
+ +P R ++ FP++ +F ++ SL+YI V+ + +K+ T T V + ++ K
Sbjct: 88 TTEPLTLRLMKVWFPVNVIFVGMLITSMFSLKYINVAMVTVLKNVTNVITAVGEMYLFNK 147
Query: 131 YFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDS 190
D R+WA+L ++ + IT+LSFN G+ + C T++ ++ + K
Sbjct: 148 QHDNRVWAALFLMIISAVSGGITDLSFNAVGYAWQIANCFLTASYSLTLRKTMDTAK--Q 205
Query: 191 INTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPW--------AAMIIIFSSGVLAF 242
+ ++ F+ +++ L G + +F+ Y + + +++ SG+L
Sbjct: 206 VTQSGNLNEFSMVLLNNTLSLPLGLLLSYFFNEMDYLYQTPLLRLPSFWMVMTLSGLLGL 265
Query: 243 CLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLV 296
++F+ + +H T A T+++ G+L + ++F P S N+ LV
Sbjct: 266 AISFTSMWFLHQTGATTYSLVGSLNKIPLSIAGIVLFNVPTSLQNSASILFGLV 319
>FCT1_DROME (Q9VHT4) Probable GDP-fucose transporter
Length = 337
Score = 81.6 bits (200), Expect = 3e-15
Identities = 69/305 (22%), Positives = 130/305 (42%), Gaps = 26/305 (8%)
Query: 21 WWAFNVTVIIMNKWIFQK--LDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQ- 77
+W ++ + +NK + ++ PL +S + S + +V ++ + P + P+
Sbjct: 25 YWCTSILTVFVNKHLLSSDTVNLGAPLFMSWFQCVVSTVICFVASRLSRKYPSVFTFPEG 84
Query: 78 -----DRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYF 132
D +R+I P+S ++ + I N+SL Y+ V+F +S T +VVL +++ R+
Sbjct: 85 NPLDIDTFRKILPLSVLYTLMIGANNLSLSYVTVAFYYIGRSLTTVFSVVLTYVILRQRT 144
Query: 133 DWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLAT--------STKTILAEALLH 184
++ IV G L E +F + +FG L++ TK L
Sbjct: 145 SFKCLLCCGAIVVGFWLGVDQESLTEVFSWRGTIFGVLSSLALAMFSIQTKKSLGYVNQE 204
Query: 185 GYKFDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMI--IIFSSGVLAF 242
+ N +Y F LI++ NG LE +P+ WA+ + SG+ F
Sbjct: 205 VWLLSYYNNLYSTLLFLPLIII--------NGELESIITYPHLWASWFWAAMTLSGLCGF 256
Query: 243 CLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYG 302
+ F I T+A+T N++G K +I+ + + S + + LV Y
Sbjct: 257 AIGFVTALEIKVTSALTHNISGTAKACAQTVIATQYYHDVRSALWWTSNVVVLVASAAYT 316
Query: 303 YVRNM 307
V+ +
Sbjct: 317 RVKQL 321
>FCT1_CAEEL (Q968A5) GDP-fucose transporter
Length = 363
Score = 72.4 bits (176), Expect = 2e-12
Identities = 66/336 (19%), Positives = 139/336 (40%), Gaps = 18/336 (5%)
Query: 8 QWSVVRSLL-AILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVL 66
+W + ++ A+ +W F++ ++ +NK++ + PL ++ + + + K
Sbjct: 23 KWESYKQVITAVSAYWVFSIGLVFLNKYLLSSVQLDAPLFITWYQCLVTVFLCLFLSKTS 82
Query: 67 KLK-----PLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTV 121
K P + +D + R + P+S VF I N+ L+Y+ VSF +S T V
Sbjct: 83 KAYGLFKFPSMPIDAKIS-REVLPLSVVFVAMISFNNLCLKYVGVSFYYVGRSLTTVFNV 141
Query: 122 VLQWLVWRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEA 181
V +L+ + + I+ G LL E + +FG LA + ++ A
Sbjct: 142 VCTYLILGQKTSGQAIGCCALIIFGFLLGVDQEGVTGTLSYTGVIFGVLA--SLSVALNA 199
Query: 182 LLHGYKFDSIN------TVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIF 235
+ S+ T+Y+ L++ P +L G ++ + I++
Sbjct: 200 IYTRKVLSSVGDCLWRLTMYN--NLNALVLFLPLMLFNGEFGAVFYFDKLFDTTFWILMT 257
Query: 236 SSGVLAFCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITL 295
GV F + + + I +T+ +T N++G K A +++ + + + + + L
Sbjct: 258 LGGVFGFMMGYVTGWQIQATSPLTHNISGTAKAAAQTVMAVVWYSELKTLLWWTSNFVVL 317
Query: 296 VGCTFYGYVRNMISQQPAVPGTPRTPRTPRSKMELL 331
G Y YV+ + + +P + K++LL
Sbjct: 318 FGSGMYTYVQKRVMDKKNSGASPAS-EAKSDKVKLL 352
>US74_DROME (Q95YI5) UDP-sugar transporter UST74c (Fringe connection
protein)
Length = 373
Score = 65.1 bits (157), Expect = 2e-10
Identities = 66/289 (22%), Positives = 129/289 (43%), Gaps = 18/289 (6%)
Query: 23 AFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDR--W 80
+F +TV+ NK + F L +S S VV+ + K L++ P R +
Sbjct: 74 SFMITVV--NKTVLTSYHFPSFLFLSLGQLTASI----VVLGMGKRLKLVNFPPLQRNTF 127
Query: 81 RRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASL 140
+IFP+ +F N++ G + + + ++ F+ T++L+ + + S+
Sbjct: 128 AKIFPLPLIFLGNMMFGLGGTKTLSLPMFAALRRFSILMTMLLELKILGLRPSNAVQVSV 187
Query: 141 VPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPF 200
++GG LL + +LSFNM G+ + T++ + + L + +Y +
Sbjct: 188 YAMIGGALLAASDDLSFNMRGYIYVMITNALTASNGVYVKKKLDTSEIGKYGLMY----Y 243
Query: 201 ATLIMVFPALLLE---GNGILEWFSIHPYPWAAMIIIF-SSGVLAFCLNFSIFYVIHSTT 256
+L M PAL L GN + + + + + ++ F S V+ F L++S +
Sbjct: 244 NSLFMFLPALALNYVTGN-LDQALNFEQWNDSVFVVQFLLSCVMGFILSYSTILCTQFNS 302
Query: 257 AVTFNVAGNLK-VAVAVLISWLIFRNPISYMNAVGCAITLVGCTFYGYV 304
A+T + G LK + V L ++ S++N +G I+++ Y YV
Sbjct: 303 ALTTTIVGCLKNICVTYLGMFIGGDYVFSWLNCIGINISVLASLLYTYV 351
>S35D_HUMAN (Q9NTN3) UDP-glucuronic acid/UDP-N-acetylgalactosamine
transporter (UDP-GlcA/UDP-GalNAc transporter) (Solute
carrier family 35 member D1)
Length = 355
Score = 60.8 bits (146), Expect = 5e-09
Identities = 57/301 (18%), Positives = 127/301 (41%), Gaps = 14/301 (4%)
Query: 10 SVVRSLLAILQWWAFNVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYV--VIKVLK 67
+V LLA + + ++++NK + F L V + + +V ++V+K
Sbjct: 38 TVFLKLLAAGFYGVSSFLIVVVNKSVLTNYRFPSSLCVGLGQMVATVAVLWVGKALRVVK 97
Query: 68 LKPLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLV 127
L P R+ FP+ ++ N + G S + + + ++ F+ T+ + ++
Sbjct: 98 FPDLDRNVP----RKTFPLPLLYFGNQITGLFSTKKLNLPMFTVLRRFSILFTMFAEGVL 153
Query: 128 WRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYK 187
+K F W I ++ ++ G + + ++L+F++ G+ L + T+ + L +
Sbjct: 154 LKKTFSWGIKMTVFAMIIGAFVAASSDLAFDLEGYAFILINDVLTAANGAYVKQKLDSKE 213
Query: 188 FDSINTVYHMAPFATLIMVFPALLLEGNGILEWFSIHPYPWAAMIIIFS---SGVLAFCL 244
+Y+ A L M+ P L + ++ WA + + S V+ F L
Sbjct: 214 LGKYGLLYYNA----LFMILPTLAIAYFTGDAQKAVEFEGWADTLFLLQFTLSCVMGFIL 269
Query: 245 NFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPI-SYMNAVGCAITLVGCTFYGY 303
++ +A+T + G +K + I + + I ++ N +G I++ G Y Y
Sbjct: 270 MYATVLCTQYNSALTTTIVGCIKNILITYIGMVFGGDYIFTWTNFIGLNISIAGSLVYSY 329
Query: 304 V 304
+
Sbjct: 330 I 330
>FCT1_HUMAN (Q96A29) GDP-fucose transporter 1 (Solute carrier family
35 member C2)
Length = 364
Score = 58.2 bits (139), Expect = 3e-08
Identities = 69/327 (21%), Positives = 135/327 (41%), Gaps = 21/327 (6%)
Query: 16 LAILQWWAFNVTVIIMNKWIFQKLDFKF--PLSVS---CI--HFICSAIGAYVVIKVLKL 68
L + +W +++++ +NK++ + P+ V+ C+ +C + A +
Sbjct: 43 LVVSLYWVTSISMVFLNKYLLDSPSLRLDTPIFVTFYQCLVTTLLCKGLSALAACCPGAV 102
Query: 69 K-PLISVDPQDRWRRIFPMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLV 127
P + +D + R + P+S VF I N+ L+Y+ V+F +S T V+L +L+
Sbjct: 103 DFPSLRLDLRVA-RSVLPLSVVFIGMITFNNLCLKYVGVAFYNVGRSLTTVFNVLLSYLL 161
Query: 128 WRKYFDWRIWASLVPIVGGILLTSITELSFNMFGFCAALFGCLAT---STKTILAEALLH 184
++ + + I+GG L E + + +FG LA+ S I +L
Sbjct: 162 LKQTTSFYALLTCGIIIGGFWLGVDQEGAEGTLSWLGTVFGVLASLCVSLNAIYTTKVLP 221
Query: 185 GYKFDSINTVYHMAPFATLIMVFPALLLEG--NGILEWFSI-HPYPWAAMIIIFSSGVLA 241
SI + I+ P LLL G + ++ + + W M + G+
Sbjct: 222 AVD-GSIWRLTFYNNVNACILFLPLLLLLGELQALRDFAQLGSAHFWGMMTL---GGLFG 277
Query: 242 FCLNFSIFYVIHSTTAVTFNVAGNLKVAVAVLISWLIFRNPISYMNAVGCAITLVGCTFY 301
F + + I T+ +T NV+G K +++ L + S++ + L G + Y
Sbjct: 278 FAIGYVTGLQIKFTSPLTHNVSGTAKACAQTVLAVLYYEETKSFLWWTSNMMVLGGSSAY 337
Query: 302 GYVRNMISQQPAVPGTPRTPRTPRSKM 328
+VR + P P + +S M
Sbjct: 338 TWVRGW--EMKKTPEEPSPKDSEKSAM 362
>SQV7_CAEEL (Q18779) UDP-sugar transporter sqv-7 (Squashed vulva
protein 7)
Length = 329
Score = 55.1 bits (131), Expect = 3e-07
Identities = 60/316 (18%), Positives = 127/316 (39%), Gaps = 23/316 (7%)
Query: 25 NVTVIIMNKWIFQKLDFKFPLSVSCIHFICSAIGAYVVIKVLKLKPLISVDPQDRWRRIF 84
+V ++ +NK + F L V + + + + K+ ++ S+D R+I
Sbjct: 23 SVLIVFVNKILLTNYKFPSFLFVGVGQMMATILILFFA-KMFRIVQFPSLDSSIP-RKIM 80
Query: 85 PMSFVFCINIVLGNVSLRYIPVSFMQTIKSFTPATTVVLQWLVWRKYFDWRIWASLVPIV 144
P+ ++ N++ G + I + ++ F+ T++L++ + + S+ ++
Sbjct: 81 PLPLLYFFNLISGLGGTQMINLPMFTVLRRFSILMTMILEFYILNVKASKAVKISVGLMI 140
Query: 145 GGILLTSITELSFNMFGFCAALFGCLATSTKTILAEALLHGYKFDSINTVYHMAPFATLI 204
GG + +I +LSF+ G+ + T+ + + L Y + + L
Sbjct: 141 GGSFIAAIYDLSFDALGYTMIFINNICTAALGVYTKQKLDAKDLGK----YGLMFYNCLF 196
Query: 205 MVFPAL-LLEGNGILEWF-------SIHPYPWAAMIIIFSSGVLAFCLNFSIFYVIHSTT 256
M+ PAL +++ G L+ S+ W ++ S + F LN+S+ H +
Sbjct: 197 MLLPALCVVQYTGDLDRAYSFMLSDSMTSSVWTCFLL---SCICGFVLNYSLVLCTHHNS 253
Query: 257 AVTFNVAGNLKVAVAVLISWLIFRNPI-SYMNAVGCAITLVGCTFYGYV-----RNMISQ 310
A+T G +K + + + + N G +++ G Y YV IS
Sbjct: 254 ALTTTCVGPIKNLFVTYVGMFSSGDYVFQWANFTGINVSVFGSILYTYVTFRSKSTTISY 313
Query: 311 QPAVPGTPRTPRTPRS 326
+P P PR+
Sbjct: 314 KPLPMTMPIDVHKPRN 329
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.331 0.142 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,966,048
Number of Sequences: 164201
Number of extensions: 1567038
Number of successful extensions: 4958
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 24
Number of HSP's successfully gapped in prelim test: 21
Number of HSP's that attempted gapping in prelim test: 4894
Number of HSP's gapped (non-prelim): 51
length of query: 342
length of database: 59,974,054
effective HSP length: 111
effective length of query: 231
effective length of database: 41,747,743
effective search space: 9643728633
effective search space used: 9643728633
T: 11
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 66 (30.0 bits)
Medicago: description of AC139354.6


