

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC139354.10 - phase: 0
(381 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
Score E
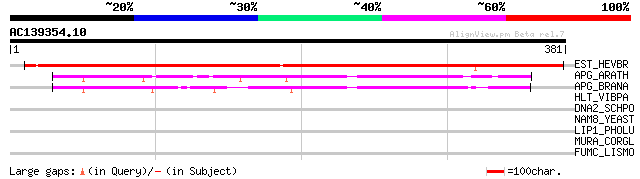
Sequences producing significant alignments: (bits) Value
EST_HEVBR (Q7Y1X1) Esterase precursor (EC 3.1.1.-) (Early nodule... 474 e-133
APG_ARATH (P40602) Anter-specific proline-rich protein APG precu... 129 9e-30
APG_BRANA (P40603) Anter-specific proline-rich protein APG (Prot... 105 1e-22
HLT_VIBPA (Q99289) Thermolabile hemolysin precursor (TL) (Lecith... 35 0.41
DNA2_SCHPO (Q9URU2) DNA replication helicase dna2 32 2.1
NAM8_YEAST (Q00539) NAM8 protein 32 2.7
LIP1_PHOLU (P40601) Lipase 1 precursor (EC 3.1.1.3) (Triacylglyc... 32 3.5
MURA_CORGL (Q8NML5) UDP-N-acetylglucosamine 1-carboxyvinyltransf... 31 4.6
FUMC_LISMO (Q8Y551) Fumarate hydratase class II (EC 4.2.1.2) (Fu... 31 6.0
>EST_HEVBR (Q7Y1X1) Esterase precursor (EC 3.1.1.-) (Early
nodule-specific protein homolog) (Latex allergen Hev b
13)
Length = 391
Score = 474 bits (1219), Expect = e-133
Identities = 229/374 (61%), Positives = 286/374 (76%), Gaps = 6/374 (1%)
Query: 11 PLVTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAAAFRAPPSPYGETYFHRST 70
P++TL L LC+ +A++ CDFPAIF+FG SN DTGG AAAF PYGET+FHRST
Sbjct: 10 PIITLSFL-LCMLSLAYASETCDFPAIFNFGDSNSDTGGKAAAFYPLNPPYGETFFHRST 68
Query: 71 GRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINIPKSILP-NGKLS 129
GR+SDGR+I+DFIA SF LPYLSPYL+SLGSNF HGA+FA+ GSTI +P +I+P +G S
Sbjct: 69 GRYSDGRLIIDFIAESFNLPYLSPYLSSLGSNFKHGADFATAGSTIKLPTTIIPAHGGFS 128
Query: 130 PFSLQIQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNK 189
PF L +QY QF++FI +++ IR+ GG+FA L+P+E YF KALY FDIGQNDLT GF N
Sbjct: 129 PFYLDVQYSQFRQFIPRSQFIRETGGIFAELVPEEYYFEKALYTFDIGQNDLTEGFL-NL 187
Query: 190 TIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKDSYGCA 249
T+++VNATVPD+VN++ N+K IY+LGAR+FWIH TGP GC IL FP A KDS GCA
Sbjct: 188 TVEEVNATVPDLVNSFSANVKKIYDLGARTFWIHNTGPIGCLSFILTYFPWAEKDSAGCA 247
Query: 250 KQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCG 309
K YNEV+Q+FN KLKE +A+LR +L A +VDIY+ KYSLF+ PEK+GFE P + CCG
Sbjct: 248 KAYNEVAQHFNHKLKEIVAQLRKDLPLATFVHVDIYSVKYSLFSEPEKHGFEFPLITCCG 307
Query: 310 YGGEYNIGV--GCGASINI-NGTKIVAGSCKNPSTRIIWDGVHYTEAANEIVFSQILTGV 366
YGG+YN V CG ++ +GTKIV GSC PS R+ WDG HYTEAANE F QI TG
Sbjct: 308 YGGKYNFSVTAPCGDTVTADDGTKIVVGSCACPSVRVNWDGAHYTEAANEYFFDQISTGA 367
Query: 367 FNDPPISLDRACYR 380
F+DPP+ L+ AC++
Sbjct: 368 FSDPPVPLNMACHK 381
>APG_ARATH (P40602) Anter-specific proline-rich protein APG
precursor
Length = 534
Score = 129 bits (325), Expect = 9e-30
Identities = 105/344 (30%), Positives = 158/344 (45%), Gaps = 36/344 (10%)
Query: 30 KNCDFPAIFSFGASNVDTGG---LAAAFRAPPSPYGETY-FHRSTGRFSDGRIILDFIAR 85
+N PA+F FG S DTG L ++ PYG + F +TGRFS+G + D++A+
Sbjct: 198 ENKTIPAVFFFGDSVFDTGNNNNLETKIKSNYRPYGMDFKFRVATGRFSNGMVASDYLAK 257
Query: 86 SFRLP-----YLSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQYIQF 140
+ YL P + ++ G +FASGG+ N S N P Q+ Y F
Sbjct: 258 YMGVKEIVPAYLDPKIQP--NDLLTGVSFASGGAGYNPTTSEAANA--IPMLDQLTY--F 311
Query: 141 KEFISKT-KLIRDQGGVF--ATLIPKEDYFSKALYIFDIGQNDLTIGFFGN---KTIQQV 194
+++I K +L+R + + A L SK + I G NDL I +FG+ + +
Sbjct: 312 QDYIEKVNRLVRQEKSQYKLAGLEKTNQLISKGVAIVVGGSNDLIITYFGSGAQRLKNDI 371
Query: 195 NATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKDSYGCAKQYNE 254
++ I ++ + +Y GAR + GT P GC P +K C ++ N
Sbjct: 372 DSYTTIIADSAASFVLQLYGYGARRIGVIGTPPLGCVP------SQRLKKKKICNEELNY 425
Query: 255 VSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACCGYGGEY 314
SQ FN KL L +L L ++ Y+DIYT + P YGFE CC G
Sbjct: 426 ASQLFNSKLLLILGQLSKTLPNSTFVYMDIYTIISQMLETPAAYGFEETKKPCCKTG--- 482
Query: 315 NIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEIV 358
+ GA + +KI C N S+ + WDGVH T+ A + +
Sbjct: 483 --LLSAGALCKKSTSKI----CPNTSSYLFWDGVHPTQRAYKTI 520
>APG_BRANA (P40603) Anter-specific proline-rich protein APG (Protein
CEX) (Fragment)
Length = 449
Score = 105 bits (263), Expect = 1e-22
Identities = 99/349 (28%), Positives = 149/349 (42%), Gaps = 52/349 (14%)
Query: 30 KNCDFPAIFSFGASNVDTGG---LAAAFRAPPSPYGETY-FHRSTGRFSDGRIILDFIAR 85
+N PA+F FG S DTG L + PYG + +TGRFS+GR+ D+I++
Sbjct: 119 QNKTIPAVFFFGDSIFDTGNNNNLDTKLKCNYRPYGMDFPMGVATGRFSNGRVASDYISK 178
Query: 86 SFRLPYLSPYL--------NSLG-SNFTHGANFASGGSTINIPKSILPNGKLSPFSLQIQ 136
+ + P N L S+ G +FASGG+ +P++ + K++ Q+
Sbjct: 179 YLGVKEIVPAYVDKKLQQNNELQQSDLLTGVSFASGGAGY-LPQTS-ESWKVTTMLDQLT 236
Query: 137 YIQ-----FKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNKTI 191
Y Q K+ + K K + SK I G NDL +FGN
Sbjct: 237 YFQDYKKRMKKLVGKKKT--------------KKIVSKGAAIVVAGSNDLIYTYFGNGAQ 282
Query: 192 Q---QVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKDSYGC 248
V++ + ++ + +Y GAR + GT P GC P +K C
Sbjct: 283 HLKNDVDSFTTMMADSAASFVLQLYGYGARRIGVIGTPPIGCTP------SQRVKKKKIC 336
Query: 249 AKQYNEVSQYFNFKLKEALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACC 308
+ N +Q FN KL L +L L ++ I Y DIY+ + +PE YGFE CC
Sbjct: 337 NEDLNYAAQLFNSKLVIILGQLSKTLPNSTIVYGDIYSIFSKMLESPEDYGFEEIKKPCC 396
Query: 309 GYGGEYNIGVGCGASININGTKIVAGSCKNPSTRIIWDGVHYTEAANEI 357
G GV C + + N S+ + WDG+H ++ A EI
Sbjct: 397 KIGLTKG-GVFC--------KERTLKNMSNASSYLFWDGLHPSQRAYEI 436
>HLT_VIBPA (Q99289) Thermolabile hemolysin precursor (TL)
(Lecithin-dependent haemolysin) (LDH) (Atypical
phospholipase) (Phospholipase A2) (Lysophospholipase)
Length = 418
Score = 34.7 bits (78), Expect = 0.41
Identities = 39/189 (20%), Positives = 71/189 (36%), Gaps = 37/189 (19%)
Query: 186 FGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSAIKDS 245
FG N VP++ +Y E + + + GA++F +L P A K
Sbjct: 245 FGLNDFMNYNRGVPEVKADYAEALIRLTDAGAKNF-------------MLMTLPDATK-- 289
Query: 246 YGCAKQYNEVSQYFNFKLKEALAELRSNLSSAA---------ITYVDIYTPKYSLFTNPE 296
A Q+ +Q K++ + E+ + + A IT D + +L + PE
Sbjct: 290 ---APQFKYSTQEEIDKIRAKVLEMNEFIKAQAMYYKAQGYNITLFDTHALFETLTSAPE 346
Query: 297 KYGFELPFVACCGYGGEYNIGVGCGASININGTKIVAGSCKNPSTR--IIWDGVHYTEAA 354
++GF C + + +S++ T + C + WD H T A
Sbjct: 347 EHGFVNASDPC--------LDINRSSSVDYMYTHALRSECAASGAEKFVFWDVTHPTTAT 398
Query: 355 NEIVFSQIL 363
+ V ++L
Sbjct: 399 HRYVAEKML 407
>DNA2_SCHPO (Q9URU2) DNA replication helicase dna2
Length = 1398
Score = 32.3 bits (72), Expect = 2.1
Identities = 18/45 (40%), Positives = 24/45 (53%), Gaps = 1/45 (2%)
Query: 86 SFRLPY-LSPYLNSLGSNFTHGANFASGGSTINIPKSILPNGKLS 129
+ RL Y ++ +NSL S +G N G TI+ K ILP LS
Sbjct: 1127 TLRLQYRMNEDINSLSSELIYGGNLVCGSKTISQKKLILPKAHLS 1171
>NAM8_YEAST (Q00539) NAM8 protein
Length = 523
Score = 32.0 bits (71), Expect = 2.7
Identities = 32/143 (22%), Positives = 57/143 (39%), Gaps = 17/143 (11%)
Query: 135 IQYIQFKEFISKTKLIRDQGGVFATLIPKEDYFSKALYIFDIGQNDLTIGFFGNK----- 189
+ Y Q + S+ L+R+ + I E S + G +++++G GN+
Sbjct: 1 MSYKQTTYYPSRGNLVRNDSSPYTNTISSETNNSSTSVLSLQGASNVSLGTTGNQLYMGD 60
Query: 190 --------TIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILANFPSA 241
T++Q+ A++ + N N N G+RS GPK +FPS+
Sbjct: 61 LDPTWDKNTVRQIWASLGEANINVRMMWNNTLNNGSRS----SMGPKNNQGYCFVDFPSS 116
Query: 242 IKDSYGCAKQYNEVSQYFNFKLK 264
+ K + + N KLK
Sbjct: 117 THAANALLKNGMLIPNFPNKKLK 139
>LIP1_PHOLU (P40601) Lipase 1 precursor (EC 3.1.1.3)
(Triacylglycerol lipase)
Length = 645
Score = 31.6 bits (70), Expect = 3.5
Identities = 35/129 (27%), Positives = 51/129 (39%), Gaps = 31/129 (24%)
Query: 252 YNEVSQYFNFKLKE---ALAELRSNLSSAAITYVDIYTPKYSLFTNPEKYGFELPFVACC 308
YNE+S + KL + L ++ + + I VD+ + + NP +YGF
Sbjct: 243 YNELSSNAS-KLVDNYNQLEDMALSQENGNIVRVDVNALLHEVIANPLRYGF-------- 293
Query: 309 GYGGEYNIGVGCGASININGTKIVAGSCKNPSTR-------IIWDGVHYTEAANEIVFSQ 361
IG C +N AGSC++ T + D H T A+ IV SQ
Sbjct: 294 ----LNTIGYACAQGVN-------AGSCRSKDTGFDASKPFLFADDFHPTPEAHHIV-SQ 341
Query: 362 ILTGVFNDP 370
V N P
Sbjct: 342 YTVSVLNAP 350
>MURA_CORGL (Q8NML5) UDP-N-acetylglucosamine
1-carboxyvinyltransferase (EC 2.5.1.7) (Enoylpyruvate
transferase) (UDP-N-acetylglucosamine enolpyruvyl
transferase) (EPT)
Length = 418
Score = 31.2 bits (69), Expect = 4.6
Identities = 36/118 (30%), Positives = 50/118 (41%), Gaps = 5/118 (4%)
Query: 13 VTLIVLVLCITPPIFATKNCDFPAIFSFGASNVDTGGLAA-AFRAPPS-PYGETYFHRST 70
VT+ + IT P + N DFPA+ F AS G L A RA S P G+ R
Sbjct: 65 VTIDGSTVTITTPAELSSNADFPAVTQFRASVCVLGPLTARCGRAVVSLPGGDAIGSRPL 124
Query: 71 GRFSDGRIILDFIARSFRLPYLSPYLNSLGSNFTHGANFASGGSTINI-PKSILPNGK 127
G L R ++ +G+N T +F S G+T NI S++ G+
Sbjct: 125 DMHQSGLEKLGATTRISHGAVVAEAEKLVGANIT--LDFPSVGATENILTASVMAEGR 180
>FUMC_LISMO (Q8Y551) Fumarate hydratase class II (EC 4.2.1.2)
(Fumarase C)
Length = 455
Score = 30.8 bits (68), Expect = 6.0
Identities = 31/123 (25%), Positives = 54/123 (43%), Gaps = 11/123 (8%)
Query: 179 NDLTIGFFGNKTIQQVNATVPDIVNNYIENIKNIYNLGARSFWIHGTGPKGCAPVILAN- 237
ND+TI ++ ++N P I+ N++E+IK + + RSF +H C + AN
Sbjct: 336 NDVTINVAASQGNFELNVYKPVIIFNFLESIKLLAD-SMRSFRVH------CLEGLTANE 388
Query: 238 --FPSAIKDSYGCAKQYNEVSQYFN-FKLKEALAELRSNLSSAAITYVDIYTPKYSLFTN 294
+ + DS N Y K+ + + + L AAI + ++ L+ N
Sbjct: 389 KVIETKVNDSLMLVTALNPHIGYEKAAKIAKLAFDENTTLKEAAIKTGFVTEKEFDLWIN 448
Query: 295 PEK 297
P K
Sbjct: 449 PLK 451
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.322 0.141 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,079,768
Number of Sequences: 164201
Number of extensions: 2068713
Number of successful extensions: 4619
Number of sequences better than 10.0: 9
Number of HSP's better than 10.0 without gapping: 4
Number of HSP's successfully gapped in prelim test: 5
Number of HSP's that attempted gapping in prelim test: 4600
Number of HSP's gapped (non-prelim): 11
length of query: 381
length of database: 59,974,054
effective HSP length: 112
effective length of query: 269
effective length of database: 41,583,542
effective search space: 11185972798
effective search space used: 11185972798
T: 11
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC139354.10


