

BLAST2 result
BLASTP 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC138453.15 + phase: 0 /pseudo/partial
(195 letters)
Database: sprot
164,201 sequences; 59,974,054 total letters
Searching..................................................done
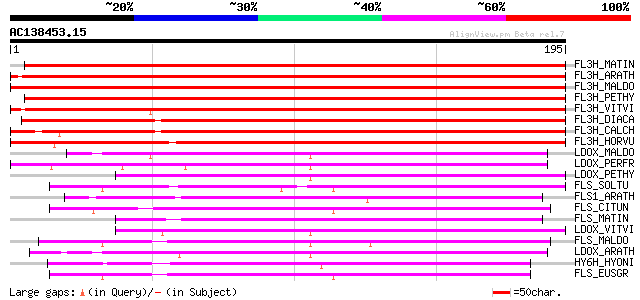
Score E
Sequences producing significant alignments: (bits) Value
FL3H_MATIN (Q05965) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 310 2e-84
FL3H_ARATH (Q9S818) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 309 2e-84
FL3H_MALDO (Q06942) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 307 9e-84
FL3H_PETHY (Q07353) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 307 1e-83
FL3H_VITVI (P41090) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 288 4e-78
FL3H_DIACA (Q05964) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 279 3e-75
FL3H_CALCH (Q05963) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 278 4e-75
FL3H_HORVU (P28038) Naringenin,2-oxoglutarate 3-dioxygenase (EC ... 257 1e-68
LDOX_MALDO (P51091) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 99 8e-21
LDOX_PERFR (O04274) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 92 8e-19
LDOX_PETHY (P51092) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 92 1e-18
FLS_SOLTU (Q41452) Flavonol synthase (EC 1.14.11.-) (FLS) 89 5e-18
FLS1_ARATH (Q96330) Flavonol synthase 1 (EC 1.14.11.-) (FLS 1) 89 6e-18
FLS_CITUN (Q9ZWQ9) Flavonol synthase (EC 1.14.11.-) (FLS) (CitFLS) 87 2e-17
FLS_MATIN (O04395) Flavonol synthase (EC 1.14.11.-) (FLS) (Fragm... 87 2e-17
LDOX_VITVI (P51093) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 86 4e-17
FLS_MALDO (Q9XHG2) Flavonol synthase (EC 1.14.11.-) (FLS) 86 5e-17
LDOX_ARATH (Q96323) Leucoanthocyanidin dioxygenase (EC 1.14.11.1... 86 7e-17
HY6H_HYONI (P24397) Hyoscyamine 6-dioxygenase (EC 1.14.11.11) (H... 86 7e-17
FLS_EUSGR (Q9M547) Flavonol synthase (EC 1.14.11.-) (FLS) 83 4e-16
>FL3H_MATIN (Q05965) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
(Fragment)
Length = 357
Score = 310 bits (793), Expect = 2e-84
Identities = 145/190 (76%), Positives = 166/190 (87%)
Query: 6 TLKYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEAC 65
TL LA E L S F+RD ERPKVAYN FS+EIP+ISLAGIDDVDG R E C +IVEAC
Sbjct: 4 TLTELAGESKLNSKFVRDEDERPKVAYNEFSDEIPVISLAGIDDVDGKRGEICREIVEAC 63
Query: 66 ENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPV 125
ENWGIFQ+VDHGVD L+++MTRLA+ FF LPPEEKLRFD+SGGKKGGFIVSSHLQGE V
Sbjct: 64 ENWGIFQVVDHGVDTSLVADMTRLARDFFALPPEEKLRFDMSGGKKGGFIVSSHLQGEAV 123
Query: 126 RDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKE 185
+DWRE++ YFSYP++ RDYSRWP+KP+GW +VTE+YSEKLM L+CKLLEVLSEAMGLEKE
Sbjct: 124 QDWREIVTYFSYPVRNRDYSRWPDKPQGWAKVTEEYSEKLMGLACKLLEVLSEAMGLEKE 183
Query: 186 ALTKACVDMD 195
+LT ACVDMD
Sbjct: 184 SLTNACVDMD 193
>FL3H_ARATH (Q9S818) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Naringenin 3-dioxygenase) (Flavanone
3-hydroxylase) (FH3) (TRANSPARENT TESTA 6 protein)
Length = 358
Score = 309 bits (792), Expect = 2e-84
Identities = 148/195 (75%), Positives = 170/195 (86%), Gaps = 1/195 (0%)
Query: 1 MAPVKTLKYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNK 60
MAP TL LA E L S F+RD ERPKVAYN FS+EIP+ISLAGIDDVDG R E C +
Sbjct: 1 MAP-GTLTELAGESKLNSKFVRDEDERPKVAYNVFSDEIPVISLAGIDDVDGKRGEICRQ 59
Query: 61 IVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHL 120
IVEACENWGIFQ+VDHGVD L+++MTRLA+ FF LPPE+KLRFD+SGGKKGGFIVSSHL
Sbjct: 60 IVEACENWGIFQVVDHGVDTNLVADMTRLARDFFALPPEDKLRFDMSGGKKGGFIVSSHL 119
Query: 121 QGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAM 180
QGE V+DWRE++ YFSYP++ RDYSRWP+KPEGW +VTE+YSE+LMSL+CKLLEVLSEAM
Sbjct: 120 QGEAVQDWREIVTYFSYPVRNRDYSRWPDKPEGWVKVTEEYSERLMSLACKLLEVLSEAM 179
Query: 181 GLEKEALTKACVDMD 195
GLEKE+LT ACVDMD
Sbjct: 180 GLEKESLTNACVDMD 194
>FL3H_MALDO (Q06942) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 364
Score = 307 bits (787), Expect = 9e-84
Identities = 147/195 (75%), Positives = 169/195 (86%)
Query: 1 MAPVKTLKYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNK 60
MAP TL +A EKTL+ F+RD ERPKVAYN+FSNEIPIISLAGID+V+G R E C K
Sbjct: 1 MAPATTLTSIAHEKTLQQKFVRDEDERPKVAYNDFSNEIPIISLAGIDEVEGRRGEICKK 60
Query: 61 IVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHL 120
IV ACE+WGIFQIVDHGVD +LISEMT LA+ FF LP EEKLRFD+SGGKKGGFIVSSHL
Sbjct: 61 IVAACEDWGIFQIVDHGVDAELISEMTGLAREFFALPSEEKLRFDMSGGKKGGFIVSSHL 120
Query: 121 QGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAM 180
QGE V+DWRE++ YFSYPI+ RDYSRWP+KPE W+EVT++YS++LM L+CKLL VLSEAM
Sbjct: 121 QGEAVQDWREIVTYFSYPIRHRDYSRWPDKPEAWREVTKKYSDELMGLACKLLGVLSEAM 180
Query: 181 GLEKEALTKACVDMD 195
GL+ EALTKACVDMD
Sbjct: 181 GLDTEALTKACVDMD 195
>FL3H_PETHY (Q07353) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
(Fragment)
Length = 369
Score = 307 bits (786), Expect = 1e-83
Identities = 146/190 (76%), Positives = 167/190 (87%)
Query: 6 TLKYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEAC 65
TL LA+EKTL++SFIRD ERPKVAYN FSNEIPIISL GIDD G R E C+KIV+AC
Sbjct: 8 TLTALAEEKTLQTSFIRDEDERPKVAYNQFSNEIPIISLEGIDDETGKRAEICDKIVKAC 67
Query: 66 ENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEPV 125
E+WG+FQ+VDHGVD ++IS+MT AK FF LPPEEKLRFD+SGGKKGGFIVSSHLQGE V
Sbjct: 68 EDWGVFQVVDHGVDAEVISQMTTFAKEFFALPPEEKLRFDMSGGKKGGFIVSSHLQGEVV 127
Query: 126 RDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKE 185
+DWRE++ YFSYP + RDYSRWP+KPEGW VT++YSEKLM L+CKLL+VLSEAMGLEKE
Sbjct: 128 QDWREIVTYFSYPTRARDYSRWPDKPEGWIAVTQKYSEKLMELACKLLDVLSEAMGLEKE 187
Query: 186 ALTKACVDMD 195
ALTKACVDMD
Sbjct: 188 ALTKACVDMD 197
>FL3H_VITVI (P41090) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 364
Score = 288 bits (738), Expect = 4e-78
Identities = 141/196 (71%), Positives = 165/196 (83%), Gaps = 2/196 (1%)
Query: 1 MAPVKTLKYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGID-DVDGLRIETCN 59
MAP TL LA EKTL+SSF+RD ERPKVAYN+FSNEIP+ISL + + E C
Sbjct: 1 MAPT-TLTALAGEKTLQSSFVRDEDERPKVAYNDFSNEIPVISLTKESMKLAAVVDEICR 59
Query: 60 KIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSH 119
KIVEACE+WGIFQ+V+HGVD LISEMTRLA+ FF LPPEE +RFD+SGGKKGGFIVSSH
Sbjct: 60 KIVEACEDWGIFQVVNHGVDSNLISEMTRLAREFFALPPEENVRFDMSGGKKGGFIVSSH 119
Query: 120 LQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEA 179
LQGE V+DWRE++ YFSYP++ RDYSRWP+KPEGW+ VT++YSEKLM L+CKLLEVLSEA
Sbjct: 120 LQGEAVQDWREIVTYFSYPLRTRDYSRWPDKPEGWRSVTQEYSEKLMGLACKLLEVLSEA 179
Query: 180 MGLEKEALTKACVDMD 195
M L+K+ALT ACVDMD
Sbjct: 180 MDLDKDALTNACVDMD 195
>FL3H_DIACA (Q05964) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 365
Score = 279 bits (714), Expect = 3e-75
Identities = 137/191 (71%), Positives = 157/191 (81%), Gaps = 2/191 (1%)
Query: 5 KTLKYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEA 64
KTL L + L S+F+RD ERPKVAYN FSN+IP+ISLAGID R E C KIVEA
Sbjct: 7 KTLTSLEGDDKLNSNFVRDEDERPKVAYNEFSNDIPVISLAGIDGEK--RGEICRKIVEA 64
Query: 65 CENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEP 124
CE+WGIFQ+VDHGV LI++MTRLA+ FF LP EEKLRFD+SGGKKGGFIVSSHLQGE
Sbjct: 65 CEDWGIFQVVDHGVGDDLIADMTRLAREFFALPAEEKLRFDMSGGKKGGFIVSSHLQGEV 124
Query: 125 VRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEK 184
V+DWRE++ YFSYP RDY+RWP+KPEGW +VTE+YS KLM+L+C LL VLSEAMGLE
Sbjct: 125 VQDWREIVTYFSYPTNSRDYTRWPDKPEGWIKVTEEYSNKLMTLACTLLGVLSEAMGLEL 184
Query: 185 EALTKACVDMD 195
EALTKACVDMD
Sbjct: 185 EALTKACVDMD 195
>FL3H_CALCH (Q05963) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 356
Score = 278 bits (712), Expect = 4e-75
Identities = 138/196 (70%), Positives = 164/196 (83%), Gaps = 5/196 (2%)
Query: 1 MAPVKTLKYLAQEKTL-ESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCN 59
MA +LK+ +E +L E+ F+RD ERPKV YN FSNEIP+ISLAGID R E C+
Sbjct: 1 MAAPISLKW--EEHSLHENKFVRDEDERPKVPYNTFSNEIPVISLAGIDGCR--RAEICD 56
Query: 60 KIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSH 119
+IV+ACE+WGIFQ+VDHGVD KL+S+MT LA+ FF LP +EKLRFD++GGKKGGFIVSSH
Sbjct: 57 EIVKACEDWGIFQVVDHGVDTKLLSDMTGLARDFFHLPTQEKLRFDMTGGKKGGFIVSSH 116
Query: 120 LQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEA 179
LQGE V+DWRE++ YFSYPIK RDYSRWP+KP W+ VTE+YS+ LM L+CKLLEVLSEA
Sbjct: 117 LQGEAVQDWREIVTYFSYPIKARDYSRWPDKPNEWRAVTEEYSKVLMGLACKLLEVLSEA 176
Query: 180 MGLEKEALTKACVDMD 195
MGLEKEALTKACVDMD
Sbjct: 177 MGLEKEALTKACVDMD 192
>FL3H_HORVU (P28038) Naringenin,2-oxoglutarate 3-dioxygenase (EC
1.14.11.9) (Flavonone-3-hydroxylase) (F3H) (FHT)
Length = 377
Score = 257 bits (656), Expect = 1e-68
Identities = 125/199 (62%), Positives = 152/199 (75%), Gaps = 6/199 (3%)
Query: 1 MAPVKTLKYLAQEK----TLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIE 56
MAPV +L E TL SF+RD ERPKVA++ FS+ +P+ISL GID +I
Sbjct: 1 MAPVSNETFLPTEAWGEATLRPSFVRDEDERPKVAHDRFSDAVPLISLHGIDGARRAQIR 60
Query: 57 TCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIV 116
+++ ACE+WGIFQ++DHGVD LI++MTRLA+ FF LP E+KLR+D+SGGKKGGFIV
Sbjct: 61 --DRVAAACEDWGIFQVIDHGVDADLIADMTRLAREFFALPAEDKLRYDMSGGKKGGFIV 118
Query: 117 SSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVL 176
SSHLQGE V+DWRE++ YFSYP+K RDY RWP KP GW V E+YSE+LM LSC L+ VL
Sbjct: 119 SSHLQGEAVQDWREIVTYFSYPVKARDYGRWPEKPAGWCAVVERYSERLMGLSCNLMGVL 178
Query: 177 SEAMGLEKEALTKACVDMD 195
SEAMGLE EAL KACVDMD
Sbjct: 179 SEAMGLETEALAKACVDMD 197
>LDOX_MALDO (P51091) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase) (Anthocyanidin synthase)
Length = 357
Score = 98.6 bits (244), Expect = 8e-21
Identities = 55/172 (31%), Positives = 95/172 (54%), Gaps = 6/172 (3%)
Query: 21 IRDVSERPKVAYNNFSNEIPIISLAGID-DVDGLRIETCNKIVEACENWGIFQIVDHGVD 79
I D+ E+ K NN ++P I L I+ D + +R + K+ +A +WG+ +V+HG+
Sbjct: 36 IGDIFEQEK---NNEGPQVPTIDLKEIESDNEKVRAKCREKLKKAAVDWGVMHLVNHGIS 92
Query: 80 QKLISEMTRLAKGFFDLPPEEKLRF--DLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSY 137
+L+ ++ + K FFDLP E+K ++ D + GK G+ +W + + Y
Sbjct: 93 DELMDKVRKAGKAFFDLPIEQKEKYANDQASGKIQGYGSKLANNASGQLEWEDYFFHCVY 152
Query: 138 PIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTK 189
P +RD S WP P + E T +Y+++L L+ K+L+VLS +GL++ L K
Sbjct: 153 PEDKRDLSIWPQTPADYIEATAEYAKQLRELATKVLKVLSLGLGLDEGRLEK 204
>LDOX_PERFR (O04274) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase)
Length = 362
Score = 92.0 bits (227), Expect = 8e-19
Identities = 57/201 (28%), Positives = 98/201 (48%), Gaps = 12/201 (5%)
Query: 1 MAPVKTLKYLAQE--KTLESSFIRDVSERPKVAYNNFSNE-------IPIISLAGIDDVD 51
M P ++ LA+ T+ ++R E + N + E +P I L +D D
Sbjct: 6 MGPSPRVEELARSGLDTIPKDYVRPEEELKSIIGNILAEEKSSEGPQLPTIDLEEMDSRD 65
Query: 52 GLRIETCNK-IVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRF--DLSG 108
+ C++ + +A +WG+ +++HG+ ++LI + K FF+LP EEK + D +
Sbjct: 66 EEGRKKCHEELKKAATDWGVMHLINHGIPEELIDRVKAAGKEFFELPVEEKEAYANDQAA 125
Query: 109 GKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSL 168
G G+ +W + + YP + D S WP KP + T +Y+++L +L
Sbjct: 126 GNVQGYGSKLANNASGQLEWEDYFFHCVYPEHKTDLSIWPTKPPDYIPATSEYAKQLRAL 185
Query: 169 SCKLLEVLSEAMGLEKEALTK 189
+ K+L VLS +GLEK L K
Sbjct: 186 ATKILSVLSIGLGLEKGRLEK 206
>LDOX_PETHY (P51092) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase)
Length = 430
Score = 91.7 bits (226), Expect = 1e-18
Identities = 48/160 (30%), Positives = 84/160 (52%), Gaps = 2/160 (1%)
Query: 38 EIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLP 97
++P I L ID D E C+++ +A WG+ +V+HG+ +LI+ + + FFD P
Sbjct: 51 QVPTIDLKEIDSEDKEIREKCHQLKKAAMEWGVMHLVNHGISDELINRVKVAGETFFDQP 110
Query: 98 PEEKLRF--DLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWK 155
EEK ++ D + G G+ +W + + ++P +RD S WP P +
Sbjct: 111 VEEKEKYANDQANGNVQGYGSKLANSACGQLEWEDYFFHCAFPEDKRDLSIWPKNPTDYT 170
Query: 156 EVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTKACVDMD 195
T +Y++++ +L+ K+L VLS +GLE+ L K M+
Sbjct: 171 PATSEYAKQIRALATKILTVLSIGLGLEEGRLEKEVGGME 210
>FLS_SOLTU (Q41452) Flavonol synthase (EC 1.14.11.-) (FLS)
Length = 349
Score = 89.4 bits (220), Expect = 5e-18
Identities = 53/188 (28%), Positives = 98/188 (51%), Gaps = 13/188 (6%)
Query: 15 TLESSFIRDVSERPKVA-YNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQI 73
T+ S +IR +E+P E+P+I ++ +DD + ++ +IVEA + WGIFQ+
Sbjct: 32 TIPSEYIRSENEQPAATTLQGVVLEVPVIDISNVDDDEEKLVK---EIVEASKEWGIFQV 88
Query: 74 VDHGVDQKLISEMTRLAKGFF-DLPPEEKLRFDLSGGKKG-----GFIVSSHLQGEPVRD 127
++HG+ ++I + ++ K FF ++P EEK +L K G G+ S + E +
Sbjct: 89 INHGIPDEVIENLQKVGKEFFEEVPQEEK---ELIAKKPGAQSLEGYGTSLQKEIEGKKG 145
Query: 128 WREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEAL 187
W + + + +P +Y WP P ++E E+Y++ L ++ + LS +GLE +
Sbjct: 146 WVDHLFHKIWPPSAINYRYWPKNPPSYREANEEYAKWLRKVADGIFRSLSLGLGLEGHEM 205
Query: 188 TKACVDMD 195
+A D
Sbjct: 206 MEAAGSED 213
>FLS1_ARATH (Q96330) Flavonol synthase 1 (EC 1.14.11.-) (FLS 1)
Length = 336
Score = 89.0 bits (219), Expect = 6e-18
Identities = 53/170 (31%), Positives = 87/170 (51%), Gaps = 6/170 (3%)
Query: 20 FIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVD 79
FIR E+P A F P I + + D D + +V+A E WG+FQ+V+HG+
Sbjct: 23 FIRSEKEQP--AITTFRGPTPAIPVVDLSDPDEESVRRA--VVKASEEWGLFQVVNHGIP 78
Query: 80 QKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKGGFIVSSHLQGEP--VRDWREMMIYFSY 137
+LI + + + FF+LP EK K + LQ +P + W + + + +
Sbjct: 79 TELIRRLQDVGRKFFELPSSEKESVAKPEDSKDIEGYGTKLQKDPEGKKAWVDHLFHRIW 138
Query: 138 PIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEAL 187
P +Y WP P ++EV E+Y+ + LS LL +LS+ +GL+++AL
Sbjct: 139 PPSCVNYRFWPKNPPEYREVNEEYAVHVKKLSETLLGILSDGLGLKRDAL 188
>FLS_CITUN (Q9ZWQ9) Flavonol synthase (EC 1.14.11.-) (FLS) (CitFLS)
Length = 335
Score = 87.4 bits (215), Expect = 2e-17
Identities = 53/179 (29%), Positives = 86/179 (47%), Gaps = 8/179 (4%)
Query: 15 TLESSFIRDVSERP-KVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQI 73
T+ + FIR E+P Y+ + EIP I L D ++ I EA WGIFQ+
Sbjct: 18 TIPAEFIRPEKEQPASTTYHGPAPEIPTIDLD-----DPVQDRLVRSIAEASREWGIFQV 72
Query: 74 VDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKG--GFIVSSHLQGEPVRDWREM 131
+HG+ LI ++ + K FF+LP EEK + K G+ + E + W +
Sbjct: 73 TNHGIPSDLICKLQAVGKEFFELPQEEKEVYSRPADAKDVQGYGTKLQKEVEGKKSWVDH 132
Query: 132 MIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTKA 190
+ + +P +Y WP P ++ V E+Y++ + + KL LS +G+E L +A
Sbjct: 133 LFHRVWPPSSINYRFWPKNPPSYRAVNEEYAKYMREVVDKLFTYLSLGLGVEGGVLKEA 191
>FLS_MATIN (O04395) Flavonol synthase (EC 1.14.11.-) (FLS)
(Fragment)
Length = 291
Score = 87.0 bits (214), Expect = 2e-17
Identities = 45/150 (30%), Positives = 83/150 (55%), Gaps = 5/150 (3%)
Query: 38 EIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKGFFDLP 97
++P++ L+ D+ R +V+A E+WG+FQ+V+HG+ +LI + ++ + FF+LP
Sbjct: 1 QVPVVDLSCPDEELVART-----VVKASEDWGVFQVVNHGIPTELIQRLQKVGREFFELP 55
Query: 98 PEEKLRFDLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEV 157
EK G G+ L + + + + ++P +Y WP P ++EV
Sbjct: 56 EAEKRSCAREAGSVEGYGRRIELDIKKRKGIVDQIYLSTWPPSSVNYRYWPKSPPDYREV 115
Query: 158 TEQYSEKLMSLSCKLLEVLSEAMGLEKEAL 187
E+Y+ + +LS K++E LSE +GL +EA+
Sbjct: 116 NEEYARHVKTLSEKIMEWLSEGLGLGREAI 145
>LDOX_VITVI (P51093) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase)
Length = 362
Score = 86.3 bits (212), Expect = 4e-17
Identities = 50/165 (30%), Positives = 82/165 (49%), Gaps = 7/165 (4%)
Query: 38 EIPIISLAGIDDVDG-----LRIETCNKIVEACENWGIFQIVDHGVDQKLISEMTRLAKG 92
++P I L I+ D +R ++ +A WG+ +V+HG+ LI+ + +
Sbjct: 48 QVPTIDLKDIESEDEVVRREIRERCREELKKAAMEWGVMHLVNHGISDDLINRVKVAGET 107
Query: 93 FFDLPPEEKLRF--DLSGGKKGGFIVSSHLQGEPVRDWREMMIYFSYPIKQRDYSRWPNK 150
FF+LP EEK ++ D + GK G+ +W + + +P +RD + WP
Sbjct: 108 FFNLPMEEKEKYANDQASGKIAGYGSKLANNASGQLEWEDYFFHLIFPEDKRDMTIWPKT 167
Query: 151 PEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEKEALTKACVDMD 195
P + T +YS KL SL+ K+L VLS +GLE+ L K M+
Sbjct: 168 PSDYVPATCEYSVKLRSLATKILSVLSLGLGLEEGRLEKEVGGME 212
>FLS_MALDO (Q9XHG2) Flavonol synthase (EC 1.14.11.-) (FLS)
Length = 337
Score = 85.9 bits (211), Expect = 5e-17
Identities = 55/187 (29%), Positives = 96/187 (50%), Gaps = 12/187 (6%)
Query: 11 AQEKTLESSFIRDVSERPKVA-YNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWG 69
+ E T+ + FIR +E+P + + E+PII + D+ + +I EA NWG
Sbjct: 12 SNEGTIPAEFIRSENEQPGITTVHGKVLEVPIIDFSDPDEE-----KLIVQITEASSNWG 66
Query: 70 IFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRF-----DLSGGKKGGFIVSSHLQGEP 124
++QIV+H + ++IS++ + K FF+LP EEK + S G + +G+
Sbjct: 67 MYQIVNHDIPSEVISKLQAVGKEFFELPQEEKEAYAKPPDSASIEGYGTKLFKEISEGDT 126
Query: 125 V-RDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLE 183
+ W + + +P +Y WP P ++E E+Y++ L ++ KL +LS +GLE
Sbjct: 127 TKKGWVDNLFNKIWPPSVVNYQFWPKNPPSYREANEEYAKHLHNVVEKLFRLLSLGLGLE 186
Query: 184 KEALTKA 190
+ L KA
Sbjct: 187 GQELKKA 193
>LDOX_ARATH (Q96323) Leucoanthocyanidin dioxygenase (EC 1.14.11.19)
(LDOX) (Leucocyanidin oxygenase) (Leucoanthocyanidin
hydroxylase) (Anthocyanidin synthase) (ANS)
Length = 356
Score = 85.5 bits (210), Expect = 7e-17
Identities = 53/185 (28%), Positives = 92/185 (49%), Gaps = 8/185 (4%)
Query: 8 KYLAQEKTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETC-NKIVEACE 66
+Y+ ++ LES I DV K ++P I L I+ D E C ++ +A
Sbjct: 21 EYIRPKEELES--INDVFLEEK---KEDGPQVPTIDLKNIESDDEKIRENCIEELKKASL 75
Query: 67 NWGIFQIVDHGVDQKLISEMTRLAKGFFDLPPEEKLRF--DLSGGKKGGFIVSSHLQGEP 124
+WG+ +++HG+ L+ + + + FF L EEK ++ D + GK G+
Sbjct: 76 DWGVMHLINHGIPADLMERVKKAGEEFFSLSVEEKEKYANDQATGKIQGYGSKLANNASG 135
Query: 125 VRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLEK 184
+W + + +YP ++RD S WP P + E T +Y++ L L+ K+ + LS +GLE
Sbjct: 136 QLEWEDYFFHLAYPEEKRDLSIWPKTPSDYIEATSEYAKCLRLLATKVFKALSVGLGLEP 195
Query: 185 EALTK 189
+ L K
Sbjct: 196 DRLEK 200
>HY6H_HYONI (P24397) Hyoscyamine 6-dioxygenase (EC 1.14.11.11)
(Hyoscyamine 6-beta-hydroxylase)
Length = 344
Score = 85.5 bits (210), Expect = 7e-17
Identities = 49/180 (27%), Positives = 92/180 (50%), Gaps = 17/180 (9%)
Query: 14 KTLESSFIRDVSERPKVAYNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQI 73
K++ SFI + +R + N++PII L + +I +AC+++G+FQ+
Sbjct: 11 KSVSESFIAPLQKRAEKDVP-VGNDVPIIDLQQHHHL------LVQQITKACQDFGLFQV 63
Query: 74 VDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSG----------GKKGGFIVSSHLQGE 123
++HG ++L+ E + K FF LP EEK +F G K ++ L E
Sbjct: 64 INHGFPEELMLETMEVCKEFFALPAEEKEKFKPKGEAAKFELPLEQKAKLYVEGEQLSNE 123
Query: 124 PVRDWREMMIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLE 183
W++ + + +P+ Q + WP KP ++EV +YS ++ L+ ++L+ + E +GL+
Sbjct: 124 EFLYWKDTLAHGCHPLDQDLVNSWPEKPAKYREVVAKYSVEVRKLTMRMLDYICEGLGLK 183
>FLS_EUSGR (Q9M547) Flavonol synthase (EC 1.14.11.-) (FLS)
Length = 334
Score = 83.2 bits (204), Expect = 4e-16
Identities = 46/172 (26%), Positives = 89/172 (51%), Gaps = 8/172 (4%)
Query: 15 TLESSFIRDVSERPKVA-YNNFSNEIPIISLAGIDDVDGLRIETCNKIVEACENWGIFQI 73
T+ + +IR +E+P ++ + E+P+I L+ D+ + + EA + WGIFQ+
Sbjct: 18 TIPAEYIRSENEQPVISTVHGVVLEVPVIDLSDSDEK-----KIVGLVSEASKEWGIFQV 72
Query: 74 VDHGVDQKLISEMTRLAKGFFDLPPEEKLRFDLSGGKKG--GFIVSSHLQGEPVRDWREM 131
V+HG+ ++I ++ + K FF+LP EEK G + G+ + + + W +
Sbjct: 73 VNHGIPNEVIRKLQEVGKHFFELPQEEKELIAKPEGSQSIEGYGTRLQKEVDGKKGWVDH 132
Query: 132 MIYFSYPIKQRDYSRWPNKPEGWKEVTEQYSEKLMSLSCKLLEVLSEAMGLE 183
+ + +P +Y WP P ++E E+Y+++L + L + LS + LE
Sbjct: 133 LFHKIWPPSAINYQFWPKNPPAYREANEEYAKRLQLVVDNLFKYLSLGLDLE 184
Database: sprot
Posted date: Nov 25, 2004 10:54 AM
Number of letters in database: 59,974,054
Number of sequences in database: 164,201
Lambda K H
0.318 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,698,720
Number of Sequences: 164201
Number of extensions: 931066
Number of successful extensions: 2376
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 55
Number of HSP's successfully gapped in prelim test: 8
Number of HSP's that attempted gapping in prelim test: 2284
Number of HSP's gapped (non-prelim): 63
length of query: 195
length of database: 59,974,054
effective HSP length: 104
effective length of query: 91
effective length of database: 42,897,150
effective search space: 3903640650
effective search space used: 3903640650
T: 11
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC138453.15


